| Full name: olfactory receptor family 1 subfamily M member 1 | Alias Symbol: OR19-6 | ||
| Type: protein-coding gene | Cytoband: 19p13.2 | ||
| Entrez ID: 125963 | HGNC ID: HGNC:8220 | Ensembl Gene: ENSG00000170929 | OMIM ID: |
OR1M1 involved pathways:
| KEGG pathway | Description | View |
|---|---|---|
| hsa04740 | Olfactory transduction |
Expression of OR1M1:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE53622 | OR1M1 | 125963 | 113434 | 0.4779 | 0.0000 | |
| GSE53624 | OR1M1 | 125963 | 113434 | 0.3836 | 0.0004 | |
| GSE97050 | OR1M1 | 125963 | A_33_P3261636 | 0.0003 | 0.9989 |
Upregulated datasets: 0; Downregulated datasets: 0.
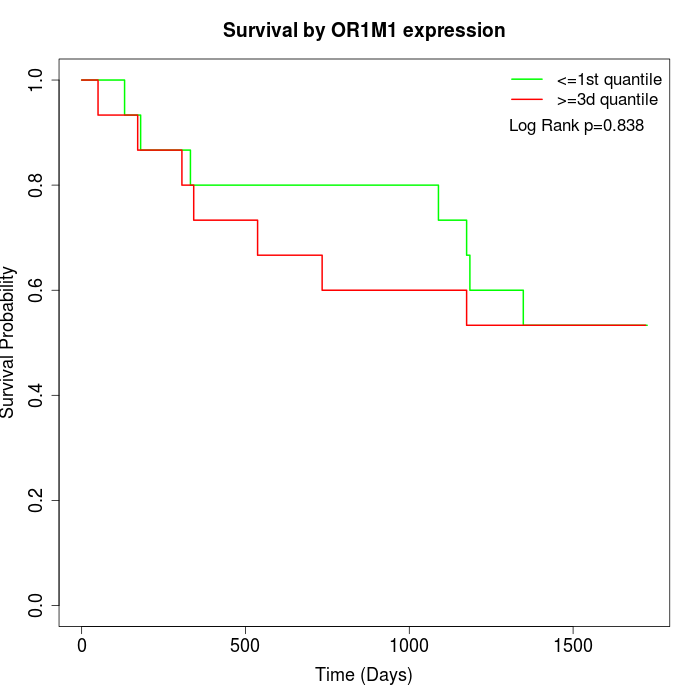
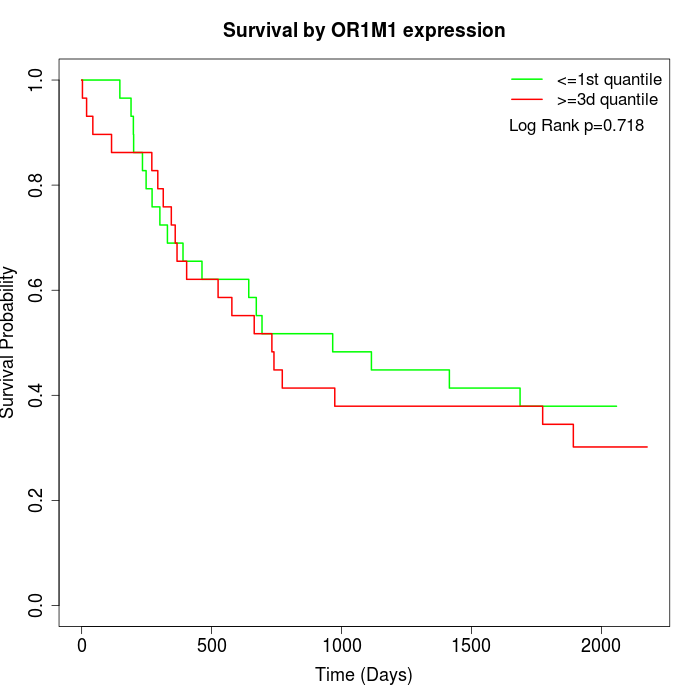
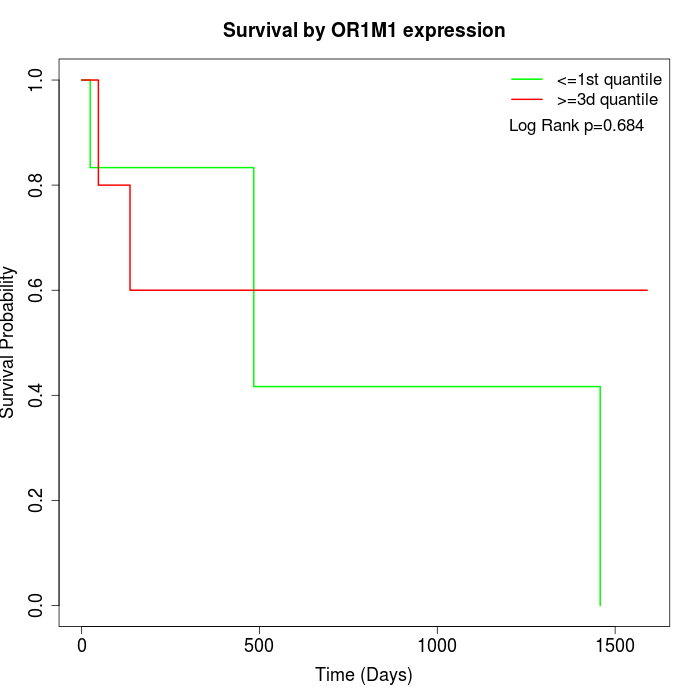
Survival by OR1M1 expression:
Note: Click image to view full size file.
Copy number change of OR1M1:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | OR1M1 | 125963 | 4 | 2 | 24 | |
| GSE20123 | OR1M1 | 125963 | 3 | 1 | 26 | |
| GSE43470 | OR1M1 | 125963 | 3 | 5 | 35 | |
| GSE46452 | OR1M1 | 125963 | 47 | 1 | 11 | |
| GSE47630 | OR1M1 | 125963 | 5 | 7 | 28 | |
| GSE54993 | OR1M1 | 125963 | 16 | 3 | 51 | |
| GSE54994 | OR1M1 | 125963 | 6 | 14 | 33 | |
| GSE60625 | OR1M1 | 125963 | 9 | 0 | 2 | |
| GSE74703 | OR1M1 | 125963 | 3 | 4 | 29 | |
| GSE74704 | OR1M1 | 125963 | 0 | 1 | 19 | |
| TCGA | OR1M1 | 125963 | 10 | 19 | 67 |
Total number of gains: 106; Total number of losses: 57; Total Number of normals: 325.
Somatic mutations of OR1M1:
Generating mutation plots.
Highly correlated genes for OR1M1:
Showing all 8 correlated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| OR1M1 | TMEM179 | 0.745986 | 3 | 0 | 3 |
| OR1M1 | TAS2R30 | 0.738331 | 3 | 0 | 3 |
| OR1M1 | KRT35 | 0.701947 | 3 | 0 | 3 |
| OR1M1 | TAS2R31 | 0.659319 | 3 | 0 | 3 |
| OR1M1 | SYNGAP1 | 0.653536 | 3 | 0 | 3 |
| OR1M1 | GPR52 | 0.631855 | 3 | 0 | 3 |
| OR1M1 | CTRL | 0.608498 | 3 | 0 | 3 |
| OR1M1 | PABPC1L | 0.570364 | 3 | 0 | 3 |
For details and further investigation, click here