| Full name: phospholipase A2 inhibitor and LY6/PLAUR domain containing | Alias Symbol: | ||
| Type: protein-coding gene | Cytoband: 19q13.31 | ||
| Entrez ID: 390940 | HGNC ID: HGNC:44206 | Ensembl Gene: ENSG00000234465 | OMIM ID: |
Expression of PINLYP:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | PINLYP | 390940 | 213556_at | -0.1069 | 0.9011 | |
| GSE20347 | PINLYP | 390940 | 213556_at | 0.1998 | 0.4263 | |
| GSE23400 | PINLYP | 390940 | 213556_at | -0.0312 | 0.6901 | |
| GSE26886 | PINLYP | 390940 | 213556_at | 0.2377 | 0.4591 | |
| GSE29001 | PINLYP | 390940 | 213556_at | -0.1160 | 0.8198 | |
| GSE38129 | PINLYP | 390940 | 213556_at | 0.0983 | 0.6609 | |
| GSE45670 | PINLYP | 390940 | 213556_at | 0.1064 | 0.7335 | |
| GSE53622 | PINLYP | 390940 | 68281 | -0.1480 | 0.4192 | |
| GSE53624 | PINLYP | 390940 | 68281 | -0.0364 | 0.8114 | |
| GSE63941 | PINLYP | 390940 | 213556_at | 0.2113 | 0.8493 | |
| GSE77861 | PINLYP | 390940 | 213556_at | 0.0930 | 0.7934 | |
| SRP064894 | PINLYP | 390940 | RNAseq | 0.0068 | 0.9850 | |
| SRP133303 | PINLYP | 390940 | RNAseq | -0.7457 | 0.0007 | |
| SRP159526 | PINLYP | 390940 | RNAseq | 0.1385 | 0.8272 | |
| SRP219564 | PINLYP | 390940 | RNAseq | -0.9148 | 0.0595 |
Upregulated datasets: 0; Downregulated datasets: 0.
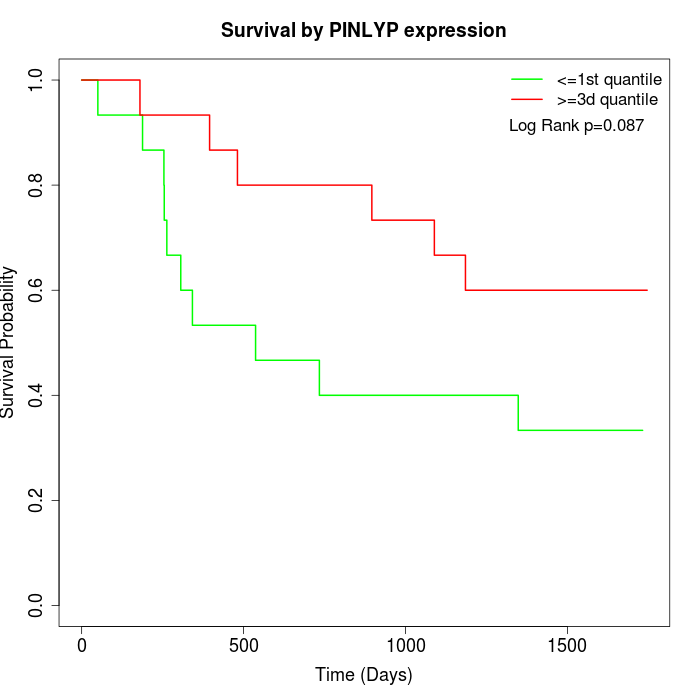
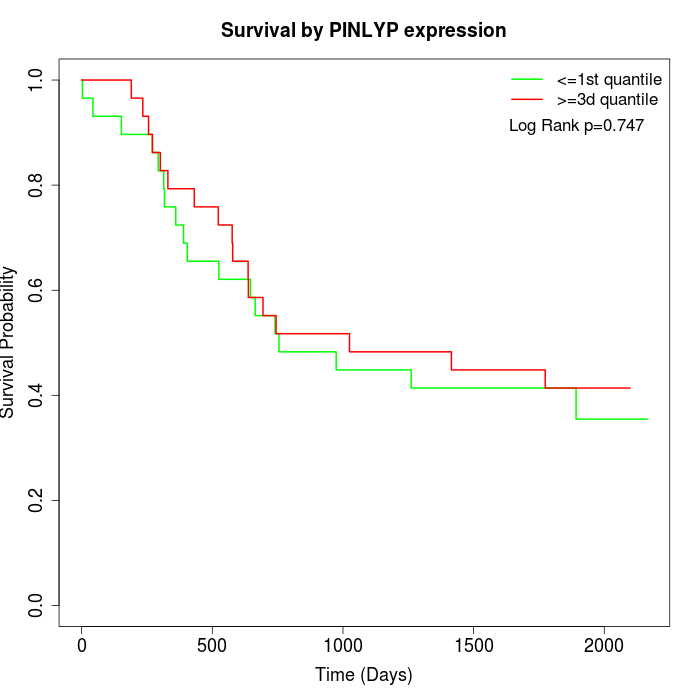
Survival by PINLYP expression:
Note: Click image to view full size file.
Copy number change of PINLYP:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | PINLYP | 390940 | 6 | 5 | 19 | |
| GSE20123 | PINLYP | 390940 | 5 | 4 | 21 | |
| GSE43470 | PINLYP | 390940 | 4 | 11 | 28 | |
| GSE46452 | PINLYP | 390940 | 45 | 1 | 13 | |
| GSE47630 | PINLYP | 390940 | 8 | 6 | 26 | |
| GSE54993 | PINLYP | 390940 | 17 | 4 | 49 | |
| GSE54994 | PINLYP | 390940 | 6 | 12 | 35 | |
| GSE60625 | PINLYP | 390940 | 9 | 0 | 2 | |
| GSE74703 | PINLYP | 390940 | 4 | 7 | 25 | |
| GSE74704 | PINLYP | 390940 | 4 | 2 | 14 | |
| TCGA | PINLYP | 390940 | 14 | 16 | 66 |
Total number of gains: 122; Total number of losses: 68; Total Number of normals: 298.
Somatic mutations of PINLYP:
Generating mutation plots.
Highly correlated genes for PINLYP:
Showing all 7 correlated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| PINLYP | SULT1E1 | 0.604874 | 3 | 0 | 3 |
| PINLYP | UVRAG | 0.601877 | 4 | 0 | 4 |
| PINLYP | PCDH17 | 0.573242 | 3 | 0 | 3 |
| PINLYP | KIAA0319 | 0.564117 | 4 | 0 | 3 |
| PINLYP | KLK5 | 0.559992 | 3 | 0 | 3 |
| PINLYP | CLDN20 | 0.546647 | 5 | 0 | 3 |
| PINLYP | GLI2 | 0.505024 | 5 | 0 | 4 |
For details and further investigation, click here