| Full name: BPI fold containing family A member 2 | Alias Symbol: bA49G10.1|SPLUNC2|PSP | ||
| Type: protein-coding gene | Cytoband: 20q11.21 | ||
| Entrez ID: 140683 | HGNC ID: HGNC:16203 | Ensembl Gene: ENSG00000131050 | OMIM ID: |
| Drug and gene relationship at DGIdb | |||
Expression of BPIFA2:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | BPIFA2 | 140683 | 237099_at | -0.0473 | 0.8709 | |
| GSE26886 | BPIFA2 | 140683 | 237099_at | -0.0482 | 0.7711 | |
| GSE45670 | BPIFA2 | 140683 | 237099_at | -0.0837 | 0.4089 | |
| GSE53622 | BPIFA2 | 140683 | 18513 | -0.1422 | 0.1757 | |
| GSE53624 | BPIFA2 | 140683 | 18513 | -0.2779 | 0.0011 | |
| GSE63941 | BPIFA2 | 140683 | 237099_at | 0.2526 | 0.2164 | |
| GSE77861 | BPIFA2 | 140683 | 237099_at | -0.2492 | 0.0179 |
Upregulated datasets: 0; Downregulated datasets: 0.
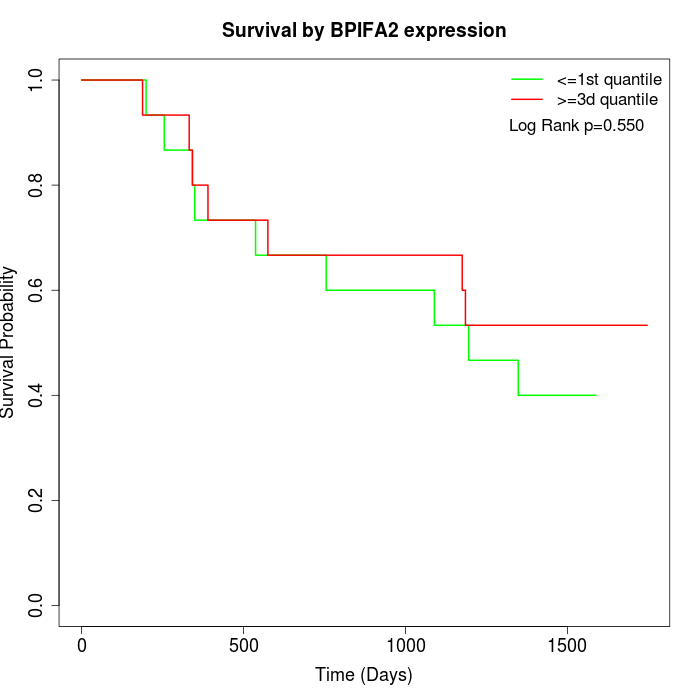
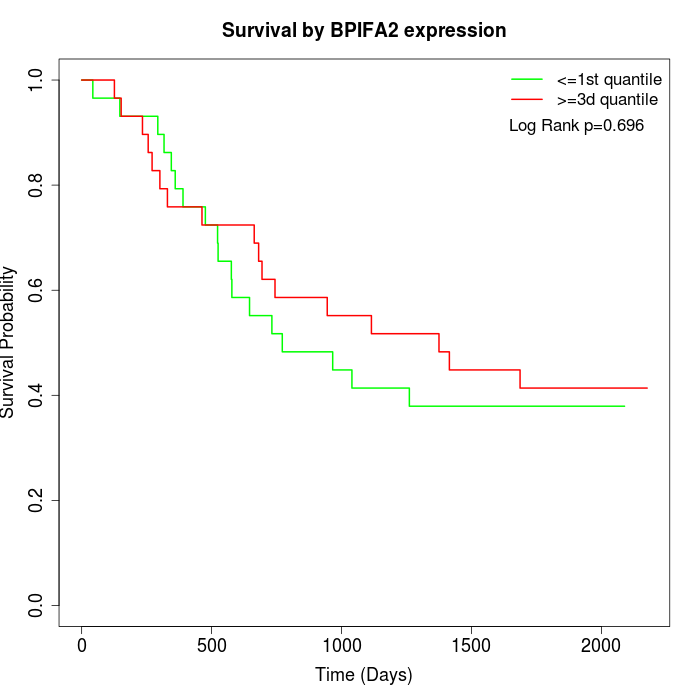
Survival by BPIFA2 expression:
Note: Click image to view full size file.
Copy number change of BPIFA2:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | BPIFA2 | 140683 | 15 | 0 | 15 | |
| GSE20123 | BPIFA2 | 140683 | 15 | 0 | 15 | |
| GSE43470 | BPIFA2 | 140683 | 12 | 0 | 31 | |
| GSE46452 | BPIFA2 | 140683 | 29 | 0 | 30 | |
| GSE47630 | BPIFA2 | 140683 | 24 | 1 | 15 | |
| GSE54993 | BPIFA2 | 140683 | 0 | 19 | 51 | |
| GSE54994 | BPIFA2 | 140683 | 26 | 1 | 26 | |
| GSE60625 | BPIFA2 | 140683 | 0 | 0 | 11 | |
| GSE74703 | BPIFA2 | 140683 | 11 | 0 | 25 | |
| GSE74704 | BPIFA2 | 140683 | 10 | 0 | 10 | |
| TCGA | BPIFA2 | 140683 | 50 | 1 | 45 |
Total number of gains: 192; Total number of losses: 22; Total Number of normals: 274.
Somatic mutations of BPIFA2:
Generating mutation plots.
Highly correlated genes for BPIFA2:
Showing top 20/31 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| BPIFA2 | SPAG11A | 0.715013 | 3 | 0 | 3 |
| BPIFA2 | OR7E24 | 0.697197 | 3 | 0 | 3 |
| BPIFA2 | FATE1 | 0.65859 | 3 | 0 | 3 |
| BPIFA2 | PRRG3 | 0.643025 | 3 | 0 | 3 |
| BPIFA2 | NEGR1-IT1 | 0.635986 | 3 | 0 | 3 |
| BPIFA2 | PRR29 | 0.632624 | 3 | 0 | 3 |
| BPIFA2 | MUC5B | 0.630337 | 3 | 0 | 3 |
| BPIFA2 | PAEP | 0.624396 | 3 | 0 | 3 |
| BPIFA2 | CSNK1G2-AS1 | 0.617893 | 3 | 0 | 3 |
| BPIFA2 | SMIM24 | 0.616961 | 3 | 0 | 3 |
| BPIFA2 | CAPN13 | 0.603399 | 3 | 0 | 3 |
| BPIFA2 | FAM209B | 0.600892 | 4 | 0 | 3 |
| BPIFA2 | CNTNAP5 | 0.593337 | 3 | 0 | 3 |
| BPIFA2 | TEKT2 | 0.589819 | 3 | 0 | 3 |
| BPIFA2 | NPHP3-AS1 | 0.577959 | 4 | 0 | 3 |
| BPIFA2 | PLK5 | 0.574942 | 4 | 0 | 3 |
| BPIFA2 | PPEF1 | 0.565418 | 4 | 0 | 3 |
| BPIFA2 | FAM53B-AS1 | 0.564403 | 3 | 0 | 3 |
| BPIFA2 | SLC30A3 | 0.553179 | 3 | 0 | 3 |
| BPIFA2 | C20orf202 | 0.552671 | 4 | 0 | 3 |
For details and further investigation, click here