| Full name: laminin subunit alpha 2 | Alias Symbol: | ||
| Type: protein-coding gene | Cytoband: 6q22.33 | ||
| Entrez ID: 3908 | HGNC ID: HGNC:6482 | Ensembl Gene: ENSG00000196569 | OMIM ID: 156225 |
| Related drugs: OCRIPLASMIN... [more] | |||
LAMA2 involved pathways:
| KEGG pathway | Description | View |
|---|---|---|
| hsa04151 | PI3K-Akt signaling pathway | |
| hsa04510 | Focal adhesion | |
| hsa05200 | Pathways in cancer | |
| hsa05222 | Small cell lung cancer |
Expression of LAMA2:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | LAMA2 | 3908 | 216840_s_at | -1.0360 | 0.1680 | |
| GSE20347 | LAMA2 | 3908 | 216840_s_at | 0.1299 | 0.3595 | |
| GSE23400 | LAMA2 | 3908 | 213519_s_at | -0.3181 | 0.0014 | |
| GSE26886 | LAMA2 | 3908 | 213519_s_at | 0.3919 | 0.0998 | |
| GSE29001 | LAMA2 | 3908 | 216840_s_at | -0.0497 | 0.9020 | |
| GSE38129 | LAMA2 | 3908 | 213519_s_at | -0.3127 | 0.3077 | |
| GSE45670 | LAMA2 | 3908 | 213519_s_at | -1.7610 | 0.0000 | |
| GSE53622 | LAMA2 | 3908 | 42001 | -0.5610 | 0.0001 | |
| GSE53624 | LAMA2 | 3908 | 42001 | -0.4252 | 0.0033 | |
| GSE63941 | LAMA2 | 3908 | 213519_s_at | -4.9136 | 0.0000 | |
| GSE77861 | LAMA2 | 3908 | 213519_s_at | 0.1260 | 0.2430 | |
| GSE97050 | LAMA2 | 3908 | A_23_P70719 | -0.5111 | 0.2805 | |
| SRP007169 | LAMA2 | 3908 | RNAseq | 1.0595 | 0.0103 | |
| SRP008496 | LAMA2 | 3908 | RNAseq | 1.8138 | 0.0000 | |
| SRP064894 | LAMA2 | 3908 | RNAseq | -0.1945 | 0.6631 | |
| SRP133303 | LAMA2 | 3908 | RNAseq | -0.7450 | 0.0081 | |
| SRP159526 | LAMA2 | 3908 | RNAseq | -0.0584 | 0.8793 | |
| SRP193095 | LAMA2 | 3908 | RNAseq | 1.1250 | 0.0008 | |
| SRP219564 | LAMA2 | 3908 | RNAseq | -0.5268 | 0.4178 | |
| TCGA | LAMA2 | 3908 | RNAseq | -0.8459 | 0.0000 |
Upregulated datasets: 3; Downregulated datasets: 2.
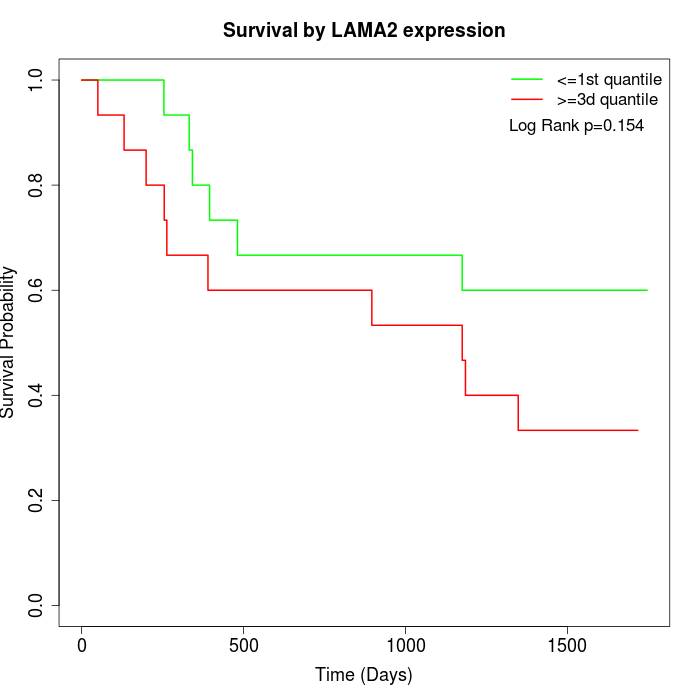
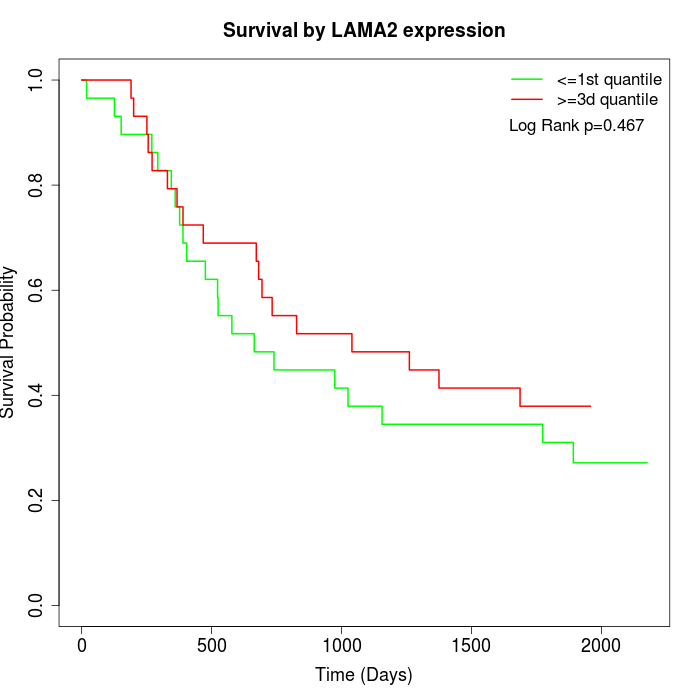
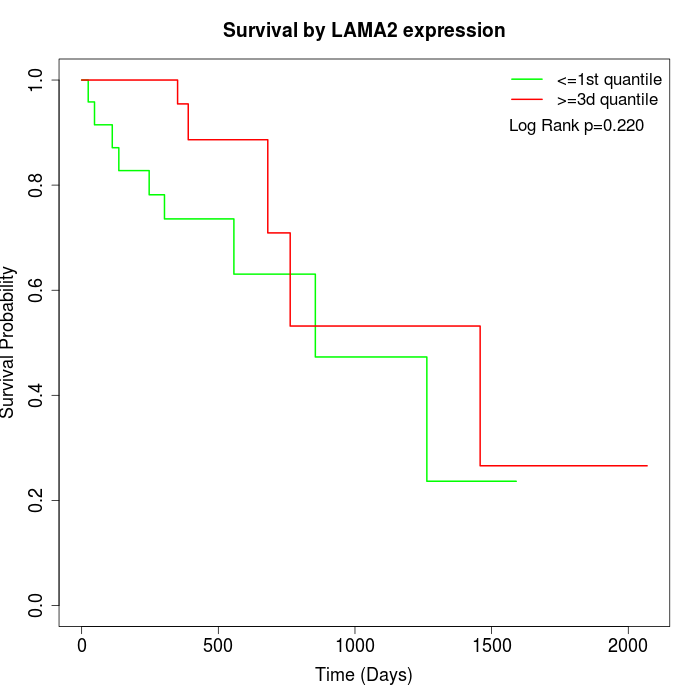
Survival by LAMA2 expression:
Note: Click image to view full size file.
Copy number change of LAMA2:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | LAMA2 | 3908 | 1 | 4 | 25 | |
| GSE20123 | LAMA2 | 3908 | 2 | 3 | 25 | |
| GSE43470 | LAMA2 | 3908 | 5 | 0 | 38 | |
| GSE46452 | LAMA2 | 3908 | 2 | 11 | 46 | |
| GSE47630 | LAMA2 | 3908 | 9 | 4 | 27 | |
| GSE54993 | LAMA2 | 3908 | 3 | 3 | 64 | |
| GSE54994 | LAMA2 | 3908 | 9 | 7 | 37 | |
| GSE60625 | LAMA2 | 3908 | 0 | 1 | 10 | |
| GSE74703 | LAMA2 | 3908 | 4 | 0 | 32 | |
| GSE74704 | LAMA2 | 3908 | 0 | 1 | 19 | |
| TCGA | LAMA2 | 3908 | 11 | 18 | 67 |
Total number of gains: 46; Total number of losses: 52; Total Number of normals: 390.
Somatic mutations of LAMA2:
Generating mutation plots.
Highly correlated genes for LAMA2:
Showing top 20/844 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| LAMA2 | CNTN4 | 0.895556 | 3 | 0 | 3 |
| LAMA2 | LRRN4CL | 0.847716 | 5 | 0 | 5 |
| LAMA2 | CNRIP1 | 0.844803 | 5 | 0 | 5 |
| LAMA2 | PEG3 | 0.830752 | 5 | 0 | 5 |
| LAMA2 | MAGI2-AS3 | 0.830108 | 5 | 0 | 5 |
| LAMA2 | AGTR1 | 0.82704 | 7 | 0 | 7 |
| LAMA2 | BMPR1B | 0.793911 | 3 | 0 | 3 |
| LAMA2 | MYOCD | 0.79222 | 4 | 0 | 4 |
| LAMA2 | DNM3OS | 0.783947 | 3 | 0 | 3 |
| LAMA2 | TEK | 0.783901 | 5 | 0 | 5 |
| LAMA2 | PABPC5 | 0.783286 | 5 | 0 | 5 |
| LAMA2 | NAP1L3 | 0.782032 | 9 | 0 | 9 |
| LAMA2 | ABCA8 | 0.781246 | 7 | 0 | 7 |
| LAMA2 | MRVI1 | 0.779936 | 5 | 0 | 5 |
| LAMA2 | CCDC80 | 0.778632 | 5 | 0 | 5 |
| LAMA2 | ZCCHC24 | 0.777281 | 8 | 0 | 8 |
| LAMA2 | ZBTB47 | 0.776141 | 4 | 0 | 4 |
| LAMA2 | FLNC | 0.774306 | 6 | 0 | 6 |
| LAMA2 | CD34 | 0.77265 | 5 | 0 | 5 |
| LAMA2 | TMEM100 | 0.771185 | 8 | 0 | 8 |
For details and further investigation, click here