| Full name: long intergenic non-protein coding RNA 486 | Alias Symbol: | ||
| Type: non-coding RNA | Cytoband: 2p22.3 | ||
| Entrez ID: 285045 | HGNC ID: HGNC:42946 | Ensembl Gene: ENSG00000230876 | OMIM ID: |
Expression of LINC00486:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | LINC00486 | 285045 | 1563082_at | -0.0585 | 0.8512 | |
| GSE26886 | LINC00486 | 285045 | 1563082_at | 0.3118 | 0.0455 | |
| GSE45670 | LINC00486 | 285045 | 1563082_at | 0.2248 | 0.0052 | |
| GSE53622 | LINC00486 | 285045 | 22283 | 0.0009 | 0.9928 | |
| GSE53624 | LINC00486 | 285045 | 22283 | 0.0974 | 0.4407 | |
| GSE63941 | LINC00486 | 285045 | 1563082_at | -0.0224 | 0.8769 | |
| GSE77861 | LINC00486 | 285045 | 1563082_at | 0.0656 | 0.6157 |
Upregulated datasets: 0; Downregulated datasets: 0.
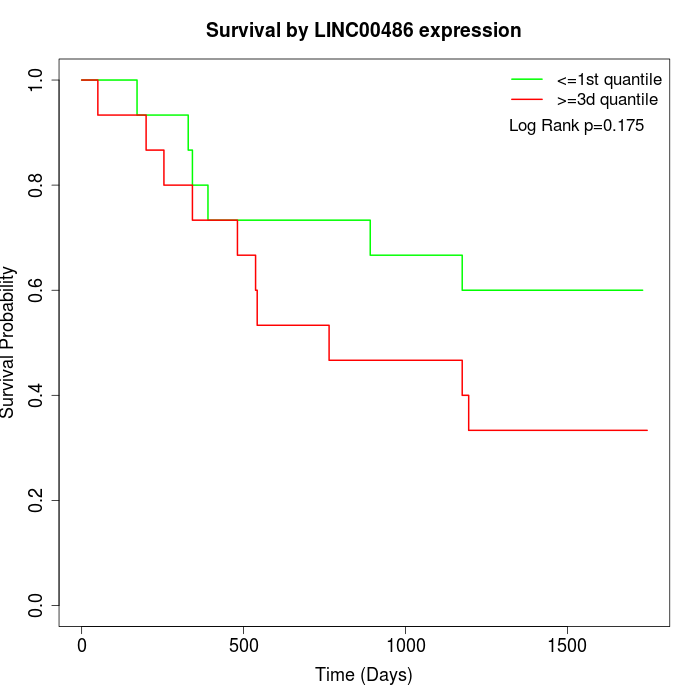
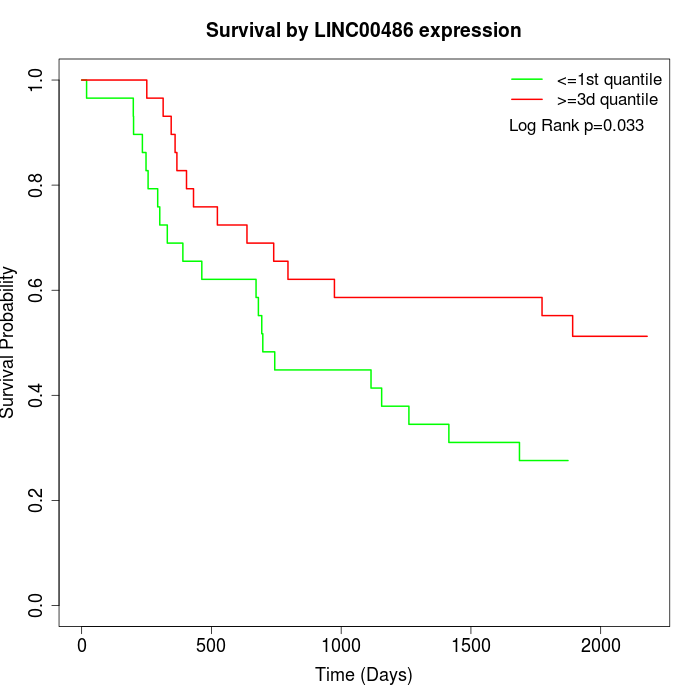
Survival by LINC00486 expression:
Note: Click image to view full size file.
Copy number change of LINC00486:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | LINC00486 | 285045 | 10 | 1 | 19 | |
| GSE20123 | LINC00486 | 285045 | 9 | 1 | 20 | |
| GSE43470 | LINC00486 | 285045 | 2 | 1 | 40 | |
| GSE46452 | LINC00486 | 285045 | 2 | 4 | 53 | |
| GSE47630 | LINC00486 | 285045 | 8 | 0 | 32 | |
| GSE54993 | LINC00486 | 285045 | 0 | 5 | 65 | |
| GSE54994 | LINC00486 | 285045 | 10 | 0 | 43 | |
| GSE60625 | LINC00486 | 285045 | 0 | 3 | 8 | |
| GSE74703 | LINC00486 | 285045 | 2 | 0 | 34 | |
| GSE74704 | LINC00486 | 285045 | 9 | 1 | 10 | |
| TCGA | LINC00486 | 285045 | 36 | 3 | 57 |
Total number of gains: 88; Total number of losses: 19; Total Number of normals: 381.
Somatic mutations of LINC00486:
Generating mutation plots.
Highly correlated genes for LINC00486:
Showing top 20/152 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| LINC00486 | KAAG1 | 0.767346 | 3 | 0 | 3 |
| LINC00486 | TMEM105 | 0.733142 | 3 | 0 | 3 |
| LINC00486 | NUTM1 | 0.730789 | 3 | 0 | 3 |
| LINC00486 | TNNI1 | 0.722282 | 3 | 0 | 3 |
| LINC00486 | AGRP | 0.717008 | 3 | 0 | 3 |
| LINC00486 | CDH22 | 0.709703 | 4 | 0 | 4 |
| LINC00486 | CECR9 | 0.705693 | 3 | 0 | 3 |
| LINC00486 | ENTPD2 | 0.699607 | 3 | 0 | 3 |
| LINC00486 | C16orf86 | 0.69851 | 3 | 0 | 3 |
| LINC00486 | C8orf74 | 0.695011 | 3 | 0 | 3 |
| LINC00486 | SUN5 | 0.694576 | 3 | 0 | 3 |
| LINC00486 | SLC22A25 | 0.689572 | 3 | 0 | 3 |
| LINC00486 | ASB12 | 0.688257 | 3 | 0 | 3 |
| LINC00486 | TSPAN11 | 0.687724 | 3 | 0 | 3 |
| LINC00486 | MYBPC3 | 0.683193 | 3 | 0 | 3 |
| LINC00486 | CCDC154 | 0.679766 | 3 | 0 | 3 |
| LINC00486 | KRTAP4-8 | 0.679102 | 3 | 0 | 3 |
| LINC00486 | LINC00161 | 0.679087 | 5 | 0 | 4 |
| LINC00486 | TMEM63B | 0.678095 | 4 | 0 | 4 |
| LINC00486 | FZD9 | 0.67799 | 4 | 0 | 4 |
For details and further investigation, click here