| Full name: lipocalin 12 | Alias Symbol: MGC48935 | ||
| Type: protein-coding gene | Cytoband: 9q34.3 | ||
| Entrez ID: 286256 | HGNC ID: HGNC:28733 | Ensembl Gene: ENSG00000184925 | OMIM ID: 612905 |
Expression of LCN12:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | LCN12 | 286256 | 230717_at | -0.0420 | 0.8727 | |
| GSE26886 | LCN12 | 286256 | 230717_at | -0.1808 | 0.0734 | |
| GSE45670 | LCN12 | 286256 | 230717_at | -0.0014 | 0.9888 | |
| GSE63941 | LCN12 | 286256 | 230717_at | 0.1441 | 0.3043 | |
| GSE77861 | LCN12 | 286256 | 230717_at | -0.1317 | 0.4889 | |
| GSE97050 | LCN12 | 286256 | A_23_P32115 | -0.8272 | 0.3324 | |
| SRP219564 | LCN12 | 286256 | RNAseq | 1.0752 | 0.2514 | |
| TCGA | LCN12 | 286256 | RNAseq | -1.1338 | 0.0304 |
Upregulated datasets: 0; Downregulated datasets: 1.
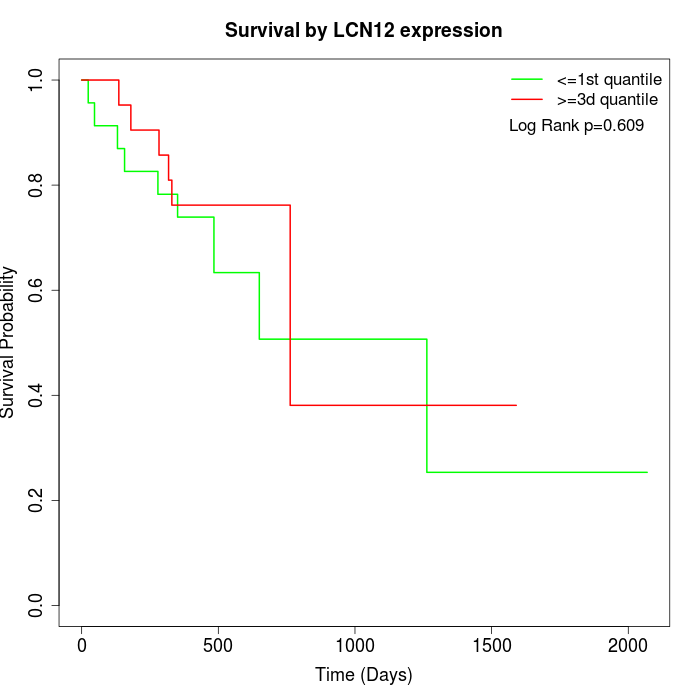
Survival by LCN12 expression:
Note: Click image to view full size file.
Copy number change of LCN12:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | LCN12 | 286256 | 5 | 7 | 18 | |
| GSE20123 | LCN12 | 286256 | 5 | 7 | 18 | |
| GSE43470 | LCN12 | 286256 | 3 | 7 | 33 | |
| GSE46452 | LCN12 | 286256 | 6 | 13 | 40 | |
| GSE47630 | LCN12 | 286256 | 6 | 15 | 19 | |
| GSE54993 | LCN12 | 286256 | 3 | 3 | 64 | |
| GSE54994 | LCN12 | 286256 | 12 | 8 | 33 | |
| GSE60625 | LCN12 | 286256 | 0 | 0 | 11 | |
| GSE74703 | LCN12 | 286256 | 3 | 5 | 28 | |
| GSE74704 | LCN12 | 286256 | 3 | 5 | 12 | |
| TCGA | LCN12 | 286256 | 29 | 23 | 44 |
Total number of gains: 75; Total number of losses: 93; Total Number of normals: 320.
Somatic mutations of LCN12:
Generating mutation plots.
Highly correlated genes for LCN12:
Showing top 20/64 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| LCN12 | INHBA-AS1 | 0.68922 | 3 | 0 | 3 |
| LCN12 | BIRC7 | 0.646077 | 3 | 0 | 3 |
| LCN12 | KCNJ4 | 0.638561 | 3 | 0 | 3 |
| LCN12 | TTYH1 | 0.632875 | 3 | 0 | 3 |
| LCN12 | DEFA4 | 0.628482 | 4 | 0 | 3 |
| LCN12 | REXO1 | 0.615718 | 4 | 0 | 3 |
| LCN12 | LINC00845 | 0.612188 | 3 | 0 | 3 |
| LCN12 | CNGB1 | 0.608469 | 4 | 0 | 3 |
| LCN12 | LINC01134 | 0.598139 | 3 | 0 | 3 |
| LCN12 | IQCJ-SCHIP1-AS1 | 0.596931 | 3 | 0 | 3 |
| LCN12 | VPS9D1 | 0.590093 | 3 | 0 | 3 |
| LCN12 | PATE1 | 0.585059 | 4 | 0 | 3 |
| LCN12 | KIF12 | 0.584354 | 4 | 0 | 3 |
| LCN12 | CACNB1 | 0.583616 | 5 | 0 | 4 |
| LCN12 | GBX2 | 0.581236 | 3 | 0 | 3 |
| LCN12 | LINC00410 | 0.573734 | 4 | 0 | 3 |
| LCN12 | RPS6KA2-AS1 | 0.571072 | 4 | 0 | 3 |
| LCN12 | GSX2 | 0.568492 | 5 | 0 | 4 |
| LCN12 | NTN3 | 0.56741 | 3 | 0 | 3 |
| LCN12 | FUT7 | 0.559618 | 5 | 0 | 3 |
For details and further investigation, click here