| Full name: calcium voltage-gated channel auxiliary subunit beta 1 | Alias Symbol: | ||
| Type: protein-coding gene | Cytoband: 17q12 | ||
| Entrez ID: 782 | HGNC ID: HGNC:1401 | Ensembl Gene: ENSG00000067191 | OMIM ID: 114207 |
| Related drugs: ATAGABALIN, BEPRIDIL HYDROCHLORIDE, GABAPENTIN, GABAPENTIN ENACARBIL, IBUTILIDE, IMAGABALIN, PREGABALIN, VERAPAMIL... [more] | |||
CACNB1 involved pathways:
| KEGG pathway | Description | View |
|---|---|---|
| hsa04010 | MAPK signaling pathway | |
| hsa04261 | Adrenergic signaling in cardiomyocytes | |
| hsa04921 | Oxytocin signaling pathway |
Expression of CACNB1:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | CACNB1 | 782 | 210185_at | 0.0430 | 0.8957 | |
| GSE20347 | CACNB1 | 782 | 210185_at | -0.0189 | 0.9011 | |
| GSE23400 | CACNB1 | 782 | 210185_at | -0.1399 | 0.0050 | |
| GSE26886 | CACNB1 | 782 | 210185_at | -0.1536 | 0.3597 | |
| GSE29001 | CACNB1 | 782 | 210185_at | -0.2687 | 0.2610 | |
| GSE38129 | CACNB1 | 782 | 210185_at | -0.1321 | 0.1991 | |
| GSE45670 | CACNB1 | 782 | 210185_at | 0.0335 | 0.7717 | |
| GSE53622 | CACNB1 | 782 | 75712 | -0.3115 | 0.0000 | |
| GSE53624 | CACNB1 | 782 | 75712 | -0.1813 | 0.0000 | |
| GSE63941 | CACNB1 | 782 | 210185_at | -0.0258 | 0.9030 | |
| GSE77861 | CACNB1 | 782 | 210185_at | -0.0884 | 0.4889 | |
| GSE97050 | CACNB1 | 782 | A_24_P91165 | -0.2052 | 0.5933 | |
| SRP064894 | CACNB1 | 782 | RNAseq | 0.6580 | 0.0000 | |
| SRP133303 | CACNB1 | 782 | RNAseq | 0.4971 | 0.0292 | |
| SRP159526 | CACNB1 | 782 | RNAseq | 0.7561 | 0.0545 | |
| SRP193095 | CACNB1 | 782 | RNAseq | 1.1486 | 0.0000 | |
| SRP219564 | CACNB1 | 782 | RNAseq | 0.1357 | 0.6674 | |
| TCGA | CACNB1 | 782 | RNAseq | 0.1914 | 0.0860 |
Upregulated datasets: 1; Downregulated datasets: 0.
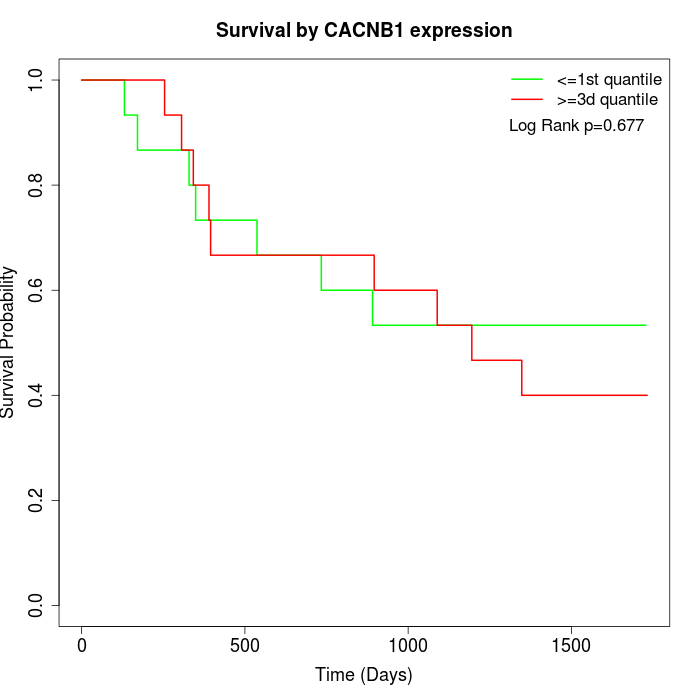
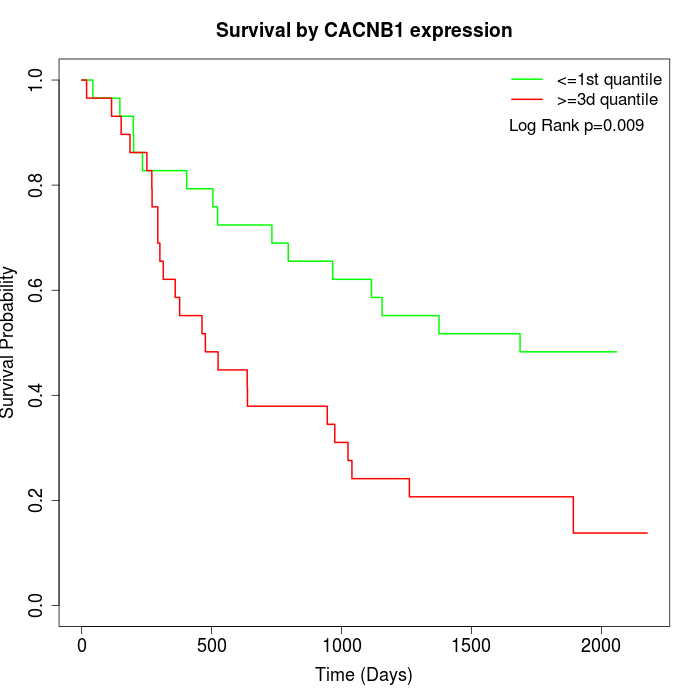
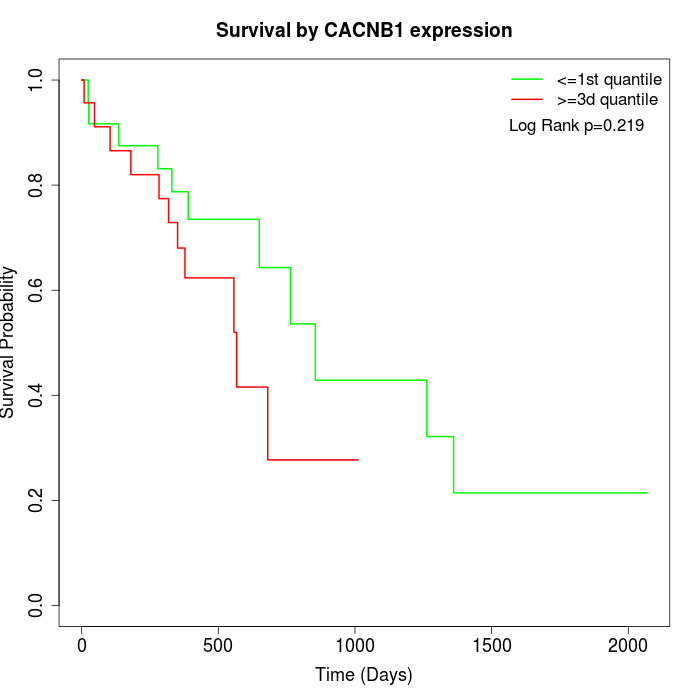
Survival by CACNB1 expression:
Note: Click image to view full size file.
Copy number change of CACNB1:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | CACNB1 | 782 | 7 | 1 | 22 | |
| GSE20123 | CACNB1 | 782 | 7 | 1 | 22 | |
| GSE43470 | CACNB1 | 782 | 1 | 1 | 41 | |
| GSE46452 | CACNB1 | 782 | 34 | 0 | 25 | |
| GSE47630 | CACNB1 | 782 | 8 | 1 | 31 | |
| GSE54993 | CACNB1 | 782 | 3 | 4 | 63 | |
| GSE54994 | CACNB1 | 782 | 8 | 5 | 40 | |
| GSE60625 | CACNB1 | 782 | 4 | 0 | 7 | |
| GSE74703 | CACNB1 | 782 | 1 | 0 | 35 | |
| GSE74704 | CACNB1 | 782 | 4 | 1 | 15 | |
| TCGA | CACNB1 | 782 | 20 | 8 | 68 |
Total number of gains: 97; Total number of losses: 22; Total Number of normals: 369.
Somatic mutations of CACNB1:
Generating mutation plots.
Highly correlated genes for CACNB1:
Showing top 20/947 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| CACNB1 | HIPK3 | 0.796329 | 3 | 0 | 3 |
| CACNB1 | PLXNA4 | 0.754154 | 3 | 0 | 3 |
| CACNB1 | CAMK2B | 0.749496 | 5 | 0 | 5 |
| CACNB1 | C8A | 0.74185 | 6 | 0 | 6 |
| CACNB1 | ZNF358 | 0.725281 | 3 | 0 | 3 |
| CACNB1 | CIITA | 0.71753 | 4 | 0 | 4 |
| CACNB1 | RHAG | 0.717404 | 3 | 0 | 3 |
| CACNB1 | BCAN | 0.711997 | 7 | 0 | 6 |
| CACNB1 | IFNA10 | 0.71079 | 5 | 0 | 5 |
| CACNB1 | NPFF | 0.710461 | 6 | 0 | 6 |
| CACNB1 | KRTAP5-AS1 | 0.708256 | 4 | 0 | 3 |
| CACNB1 | PROX1 | 0.707522 | 4 | 0 | 3 |
| CACNB1 | ZNF460 | 0.706848 | 6 | 0 | 6 |
| CACNB1 | GALNT8 | 0.704987 | 4 | 0 | 4 |
| CACNB1 | GDF5 | 0.694714 | 7 | 0 | 5 |
| CACNB1 | EPN1 | 0.694435 | 5 | 0 | 5 |
| CACNB1 | KCNQ3 | 0.694389 | 6 | 0 | 6 |
| CACNB1 | IFITM10 | 0.694028 | 3 | 0 | 3 |
| CACNB1 | GATA1 | 0.693097 | 6 | 0 | 6 |
| CACNB1 | FNDC8 | 0.689622 | 5 | 0 | 5 |
For details and further investigation, click here