| Full name: long intergenic non-protein coding RNA 226 | Alias Symbol: | ||
| Type: non-coding RNA | Cytoband: 14q32.33 | ||
| Entrez ID: 338004 | HGNC ID: HGNC:20168 | Ensembl Gene: ENSG00000276210 | OMIM ID: |
Expression of LINC00226:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | LINC00226 | 338004 | 238380_s_at | 0.0650 | 0.7696 | |
| GSE26886 | LINC00226 | 338004 | 238380_s_at | -0.1852 | 0.0839 | |
| GSE45670 | LINC00226 | 338004 | 238387_s_at | 0.0491 | 0.5545 | |
| GSE53622 | LINC00226 | 338004 | 107937 | 0.0274 | 0.8423 | |
| GSE53624 | LINC00226 | 338004 | 107937 | 0.4426 | 0.0029 | |
| GSE63941 | LINC00226 | 338004 | 238380_s_at | -0.0593 | 0.6353 | |
| GSE77861 | LINC00226 | 338004 | 238380_s_at | -0.1069 | 0.4539 |
Upregulated datasets: 0; Downregulated datasets: 0.
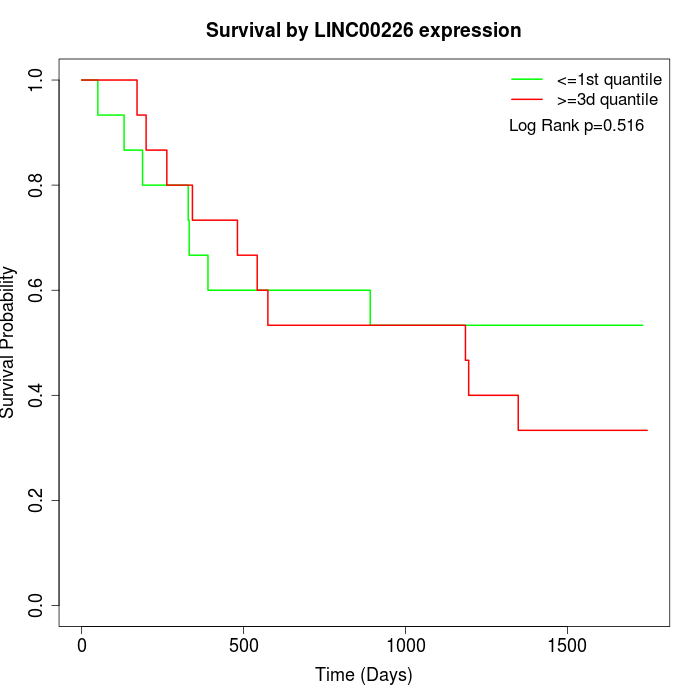
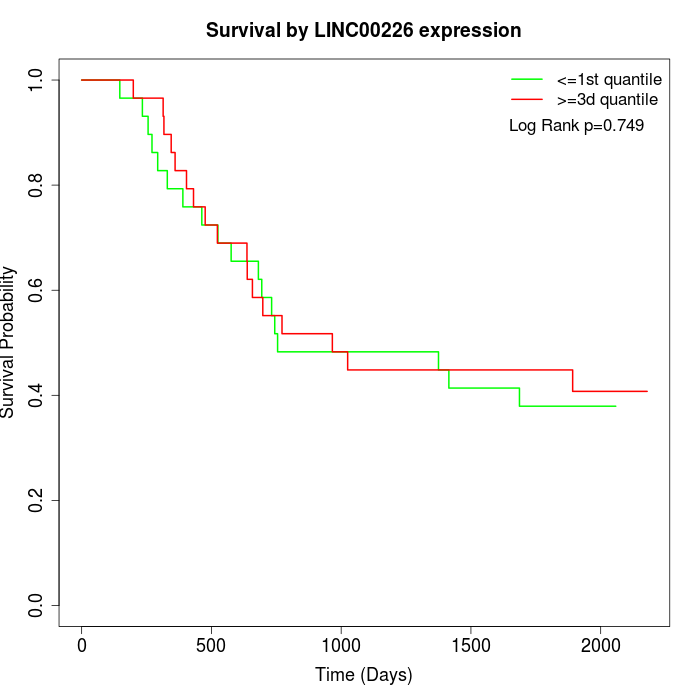
Survival by LINC00226 expression:
Note: Click image to view full size file.
Copy number change of LINC00226:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | LINC00226 | 338004 | 10 | 3 | 17 | |
| GSE20123 | LINC00226 | 338004 | 10 | 2 | 18 | |
| GSE43470 | LINC00226 | 338004 | 6 | 3 | 34 | |
| GSE46452 | LINC00226 | 338004 | 18 | 4 | 37 | |
| GSE47630 | LINC00226 | 338004 | 10 | 10 | 20 | |
| GSE54993 | LINC00226 | 338004 | 4 | 11 | 55 | |
| GSE54994 | LINC00226 | 338004 | 21 | 4 | 28 | |
| GSE60625 | LINC00226 | 338004 | 0 | 11 | 0 | |
| GSE74703 | LINC00226 | 338004 | 4 | 3 | 29 | |
| GSE74704 | LINC00226 | 338004 | 5 | 2 | 13 | |
| TCGA | LINC00226 | 338004 | 31 | 18 | 47 |
Total number of gains: 119; Total number of losses: 71; Total Number of normals: 298.
Somatic mutations of LINC00226:
Generating mutation plots.
Highly correlated genes for LINC00226:
Showing top 20/24 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| LINC00226 | C20orf203 | 0.685033 | 3 | 0 | 3 |
| LINC00226 | GNAO1 | 0.672997 | 3 | 0 | 3 |
| LINC00226 | GKN2 | 0.650387 | 3 | 0 | 3 |
| LINC00226 | CD80 | 0.621207 | 3 | 0 | 3 |
| LINC00226 | MAP3K19 | 0.614928 | 4 | 0 | 3 |
| LINC00226 | CACNA1E | 0.612845 | 3 | 0 | 3 |
| LINC00226 | LINC01119 | 0.609553 | 3 | 0 | 3 |
| LINC00226 | FGA | 0.608182 | 4 | 0 | 3 |
| LINC00226 | CELF2-AS1 | 0.602752 | 3 | 0 | 3 |
| LINC00226 | PCGEM1 | 0.592173 | 3 | 0 | 3 |
| LINC00226 | LHX3 | 0.587789 | 4 | 0 | 3 |
| LINC00226 | LINC01222 | 0.580418 | 4 | 0 | 3 |
| LINC00226 | MORF4L2-AS1 | 0.577821 | 3 | 0 | 3 |
| LINC00226 | LINC01312 | 0.574492 | 3 | 0 | 3 |
| LINC00226 | LINC00943 | 0.57419 | 4 | 0 | 3 |
| LINC00226 | CCDC172 | 0.559201 | 3 | 0 | 3 |
| LINC00226 | LINC01115 | 0.5562 | 4 | 0 | 3 |
| LINC00226 | KCNIP2-AS1 | 0.533931 | 4 | 0 | 3 |
| LINC00226 | TM4SF4 | 0.533914 | 4 | 0 | 3 |
| LINC00226 | SHOX | 0.533117 | 3 | 0 | 3 |
For details and further investigation, click here