| Full name: mediator complex subunit 25 | Alias Symbol: ARC92|ACID1|TCBAP0758|DKFZp434K0512 | ||
| Type: protein-coding gene | Cytoband: 19q13.3 | ||
| Entrez ID: 81857 | HGNC ID: HGNC:28845 | Ensembl Gene: ENSG00000104973 | OMIM ID: 610197 |
Screen Evidence:
| |||
Expression of MED25:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | MED25 | 81857 | 208110_x_at | 0.0295 | 0.9489 | |
| GSE20347 | MED25 | 81857 | 208110_x_at | -0.2739 | 0.0035 | |
| GSE23400 | MED25 | 81857 | 208110_x_at | -0.0514 | 0.2983 | |
| GSE26886 | MED25 | 81857 | 208110_x_at | 0.4059 | 0.0117 | |
| GSE29001 | MED25 | 81857 | 208110_x_at | -0.2139 | 0.2159 | |
| GSE38129 | MED25 | 81857 | 208110_x_at | -0.1675 | 0.0462 | |
| GSE45670 | MED25 | 81857 | 208110_x_at | -0.0047 | 0.9689 | |
| GSE53622 | MED25 | 81857 | 97088 | -0.1785 | 0.0209 | |
| GSE53624 | MED25 | 81857 | 97088 | 0.0465 | 0.5187 | |
| GSE63941 | MED25 | 81857 | 208110_x_at | 0.1872 | 0.3613 | |
| GSE77861 | MED25 | 81857 | 208110_x_at | -0.0899 | 0.5063 | |
| GSE97050 | MED25 | 81857 | A_33_P3322553 | 0.0386 | 0.8802 | |
| SRP007169 | MED25 | 81857 | RNAseq | -1.7223 | 0.0000 | |
| SRP008496 | MED25 | 81857 | RNAseq | -2.0058 | 0.0000 | |
| SRP064894 | MED25 | 81857 | RNAseq | -0.2027 | 0.3655 | |
| SRP133303 | MED25 | 81857 | RNAseq | -0.2553 | 0.2523 | |
| SRP159526 | MED25 | 81857 | RNAseq | -0.2429 | 0.2789 | |
| SRP193095 | MED25 | 81857 | RNAseq | -0.1221 | 0.4635 | |
| SRP219564 | MED25 | 81857 | RNAseq | -0.0152 | 0.9672 | |
| TCGA | MED25 | 81857 | RNAseq | -0.0968 | 0.0285 |
Upregulated datasets: 0; Downregulated datasets: 2.
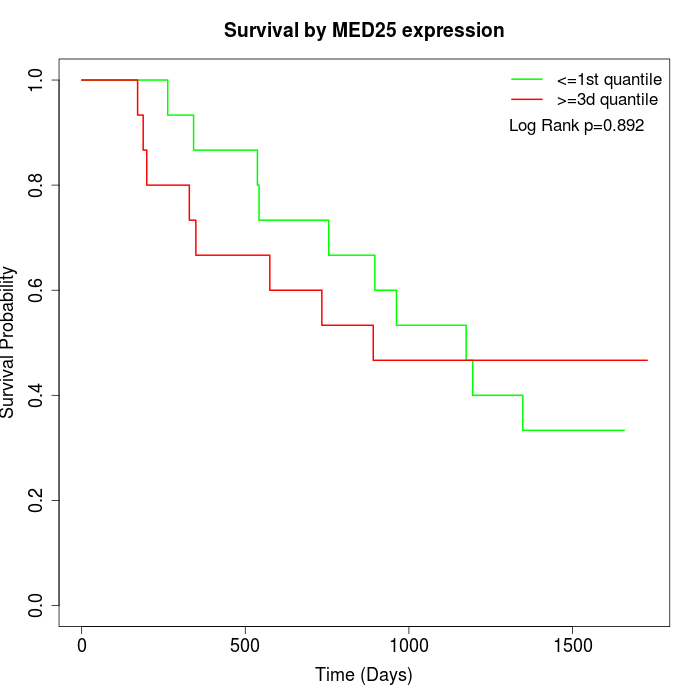
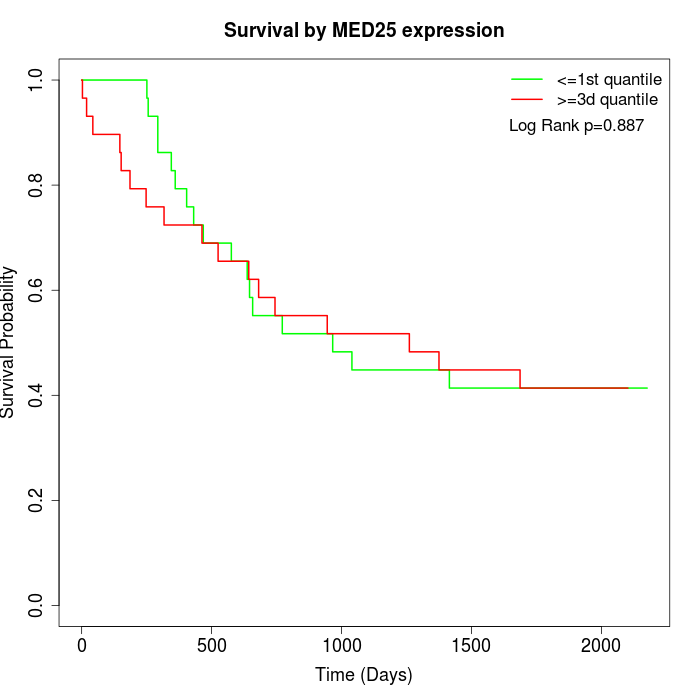
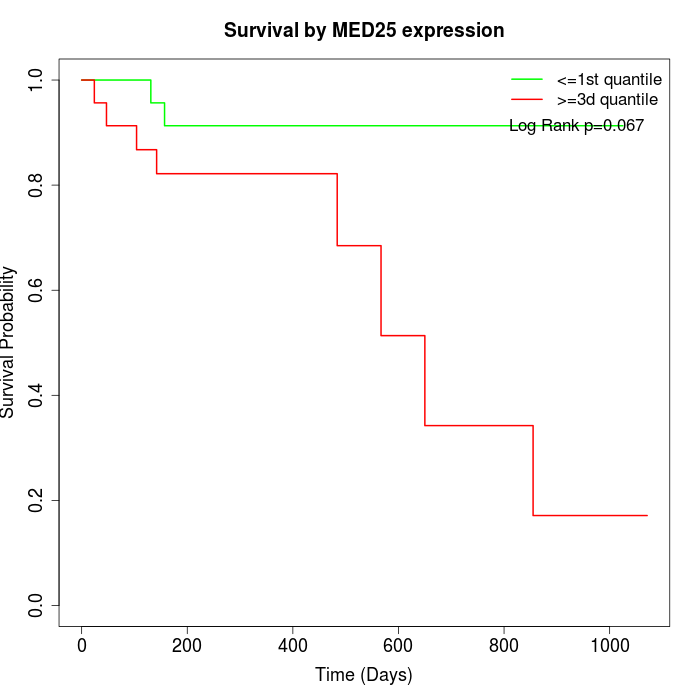
Survival by MED25 expression:
Note: Click image to view full size file.
Copy number change of MED25:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | MED25 | 81857 | 4 | 4 | 22 | |
| GSE20123 | MED25 | 81857 | 4 | 3 | 23 | |
| GSE43470 | MED25 | 81857 | 5 | 11 | 27 | |
| GSE46452 | MED25 | 81857 | 45 | 1 | 13 | |
| GSE47630 | MED25 | 81857 | 10 | 6 | 24 | |
| GSE54993 | MED25 | 81857 | 17 | 4 | 49 | |
| GSE54994 | MED25 | 81857 | 4 | 14 | 35 | |
| GSE60625 | MED25 | 81857 | 9 | 0 | 2 | |
| GSE74703 | MED25 | 81857 | 5 | 7 | 24 | |
| GSE74704 | MED25 | 81857 | 4 | 1 | 15 | |
| TCGA | MED25 | 81857 | 14 | 18 | 64 |
Total number of gains: 121; Total number of losses: 69; Total Number of normals: 298.
Somatic mutations of MED25:
Generating mutation plots.
Highly correlated genes for MED25:
Showing top 20/567 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| MED25 | TAS2R4 | 0.788328 | 3 | 0 | 3 |
| MED25 | CYP27A1 | 0.773357 | 3 | 0 | 3 |
| MED25 | C19orf54 | 0.770327 | 3 | 0 | 3 |
| MED25 | ADCK1 | 0.7574 | 3 | 0 | 3 |
| MED25 | KCNK2 | 0.749636 | 3 | 0 | 3 |
| MED25 | GAS2L1 | 0.742216 | 3 | 0 | 3 |
| MED25 | FAM104B | 0.741026 | 3 | 0 | 3 |
| MED25 | TMEM129 | 0.738531 | 3 | 0 | 3 |
| MED25 | ZNF358 | 0.728806 | 4 | 0 | 3 |
| MED25 | ZNF76 | 0.727833 | 3 | 0 | 3 |
| MED25 | ZNRD1 | 0.726781 | 3 | 0 | 3 |
| MED25 | FAM131B | 0.719412 | 3 | 0 | 3 |
| MED25 | ARAF | 0.716699 | 3 | 0 | 3 |
| MED25 | ZMIZ2 | 0.714944 | 3 | 0 | 3 |
| MED25 | GRIN2D | 0.713305 | 4 | 0 | 4 |
| MED25 | SLC9A1 | 0.712573 | 3 | 0 | 3 |
| MED25 | NOL12 | 0.708712 | 5 | 0 | 4 |
| MED25 | COG1 | 0.708556 | 3 | 0 | 3 |
| MED25 | OTUD7B | 0.707642 | 3 | 0 | 3 |
| MED25 | ZNF771 | 0.707316 | 4 | 0 | 4 |
For details and further investigation, click here