| Full name: testis expressed 22 | Alias Symbol: | ||
| Type: protein-coding gene | Cytoband: 14q32.33 | ||
| Entrez ID: 647310 | HGNC ID: HGNC:40026 | Ensembl Gene: ENSG00000226174 | OMIM ID: |
Expression of TEX22:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | TEX22 | 647310 | 1558581_at | 0.0018 | 0.9960 | |
| GSE26886 | TEX22 | 647310 | 1558581_at | -0.0152 | 0.9101 | |
| GSE45670 | TEX22 | 647310 | 1558581_at | 0.0851 | 0.3345 | |
| GSE53622 | TEX22 | 647310 | 137858 | 0.2775 | 0.0000 | |
| GSE53624 | TEX22 | 647310 | 137858 | 0.5670 | 0.0000 | |
| GSE63941 | TEX22 | 647310 | 1558581_at | -0.3670 | 0.0079 | |
| GSE77861 | TEX22 | 647310 | 1558581_at | -0.0247 | 0.8471 | |
| GSE97050 | TEX22 | 647310 | A_33_P3261695 | -0.0018 | 0.9946 | |
| SRP133303 | TEX22 | 647310 | RNAseq | 0.4342 | 0.2574 | |
| SRP219564 | TEX22 | 647310 | RNAseq | 0.8003 | 0.1224 |
Upregulated datasets: 0; Downregulated datasets: 0.
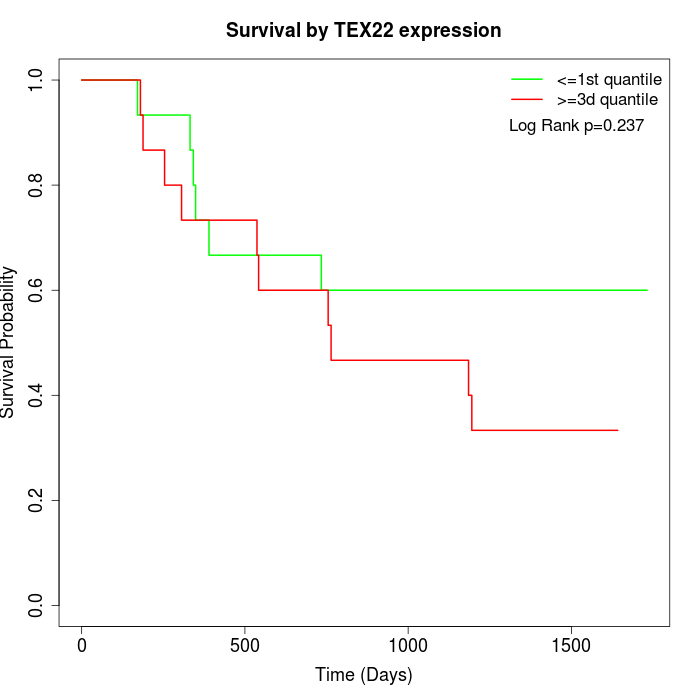
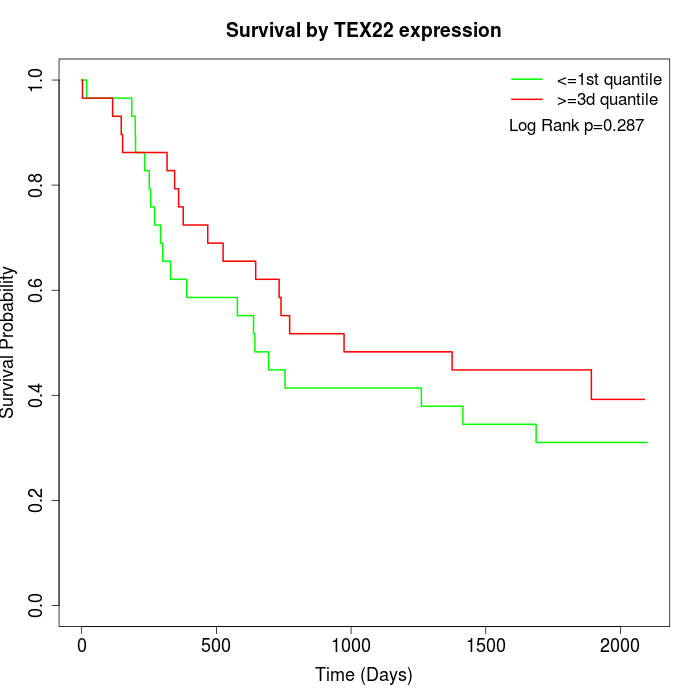
Survival by TEX22 expression:
Note: Click image to view full size file.
Copy number change of TEX22:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | TEX22 | 647310 | 11 | 3 | 16 | |
| GSE20123 | TEX22 | 647310 | 10 | 2 | 18 | |
| GSE43470 | TEX22 | 647310 | 5 | 3 | 35 | |
| GSE46452 | TEX22 | 647310 | 18 | 4 | 37 | |
| GSE47630 | TEX22 | 647310 | 10 | 10 | 20 | |
| GSE54993 | TEX22 | 647310 | 4 | 11 | 55 | |
| GSE54994 | TEX22 | 647310 | 21 | 4 | 28 | |
| GSE60625 | TEX22 | 647310 | 0 | 2 | 9 | |
| GSE74703 | TEX22 | 647310 | 4 | 3 | 29 | |
| GSE74704 | TEX22 | 647310 | 5 | 2 | 13 | |
| TCGA | TEX22 | 647310 | 31 | 18 | 47 |
Total number of gains: 119; Total number of losses: 62; Total Number of normals: 307.
Somatic mutations of TEX22:
Generating mutation plots.
Highly correlated genes for TEX22:
Showing top 20/164 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| TEX22 | MRGPRG | 0.794234 | 3 | 0 | 3 |
| TEX22 | MS4A18 | 0.783079 | 3 | 0 | 3 |
| TEX22 | C1QL2 | 0.767255 | 3 | 0 | 3 |
| TEX22 | OR5A1 | 0.764182 | 3 | 0 | 3 |
| TEX22 | OR2T1 | 0.753688 | 3 | 0 | 3 |
| TEX22 | OR10W1 | 0.750961 | 3 | 0 | 3 |
| TEX22 | BLACE | 0.748493 | 3 | 0 | 3 |
| TEX22 | DCDC1 | 0.746318 | 3 | 0 | 3 |
| TEX22 | OR2A5 | 0.745842 | 3 | 0 | 3 |
| TEX22 | PRSS38 | 0.745223 | 3 | 0 | 3 |
| TEX22 | CIB3 | 0.744103 | 4 | 0 | 4 |
| TEX22 | INSC | 0.742437 | 3 | 0 | 3 |
| TEX22 | OR2D3 | 0.739587 | 3 | 0 | 3 |
| TEX22 | OR2B11 | 0.736161 | 3 | 0 | 3 |
| TEX22 | TRPM8 | 0.734339 | 3 | 0 | 3 |
| TEX22 | ROM1 | 0.730158 | 3 | 0 | 3 |
| TEX22 | LHB | 0.72972 | 3 | 0 | 3 |
| TEX22 | PSMB11 | 0.728395 | 3 | 0 | 3 |
| TEX22 | PABPN1L | 0.726882 | 3 | 0 | 3 |
| TEX22 | SLC29A4 | 0.724291 | 3 | 0 | 3 |
For details and further investigation, click here