| Full name: phosphatase and tensin homolog | Alias Symbol: MMAC1|TEP1|PTEN1 | ||
| Type: protein-coding gene | Cytoband: 10q23.31 | ||
| Entrez ID: 5728 | HGNC ID: HGNC:9588 | Ensembl Gene: ENSG00000171862 | OMIM ID: 601728 |
| Related drugs: 2-METHOXYESTRADIOL, ABIRATERONE, AEW-541, ALPELISIB, APITOLISIB, ARQ-092, AT-13148, AZD-5363, AZD-6482, AZD-8055... [more] | |||
Screen Evidence:
| |||
PTEN involved pathways:
Expression of PTEN:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | PTEN | 5728 | 225363_at | -0.3734 | 0.1474 | |
| GSE20347 | PTEN | 5728 | 204053_x_at | -0.5274 | 0.0002 | |
| GSE23400 | PTEN | 5728 | 204053_x_at | -0.2329 | 0.0045 | |
| GSE26886 | PTEN | 5728 | 225363_at | -0.7312 | 0.0002 | |
| GSE29001 | PTEN | 5728 | 204053_x_at | -0.7042 | 0.0718 | |
| GSE38129 | PTEN | 5728 | 204053_x_at | -0.4472 | 0.0010 | |
| GSE45670 | PTEN | 5728 | 225363_at | -0.3434 | 0.0156 | |
| GSE53622 | PTEN | 5728 | 24595 | -0.4517 | 0.0000 | |
| GSE53624 | PTEN | 5728 | 24595 | -0.5071 | 0.0000 | |
| GSE63941 | PTEN | 5728 | 204053_x_at | 0.2547 | 0.5485 | |
| GSE77861 | PTEN | 5728 | 225363_at | -0.3141 | 0.0883 | |
| GSE97050 | PTEN | 5728 | A_23_P98085 | -0.1707 | 0.4920 | |
| SRP007169 | PTEN | 5728 | RNAseq | -1.4445 | 0.0000 | |
| SRP008496 | PTEN | 5728 | RNAseq | -0.8702 | 0.0002 | |
| SRP064894 | PTEN | 5728 | RNAseq | -0.2975 | 0.1103 | |
| SRP133303 | PTEN | 5728 | RNAseq | 0.0639 | 0.6506 | |
| SRP159526 | PTEN | 5728 | RNAseq | -0.5664 | 0.0313 | |
| SRP193095 | PTEN | 5728 | RNAseq | -0.4900 | 0.0000 | |
| SRP219564 | PTEN | 5728 | RNAseq | -0.5202 | 0.0649 | |
| TCGA | PTEN | 5728 | RNAseq | -0.1230 | 0.0056 |
Upregulated datasets: 0; Downregulated datasets: 1.
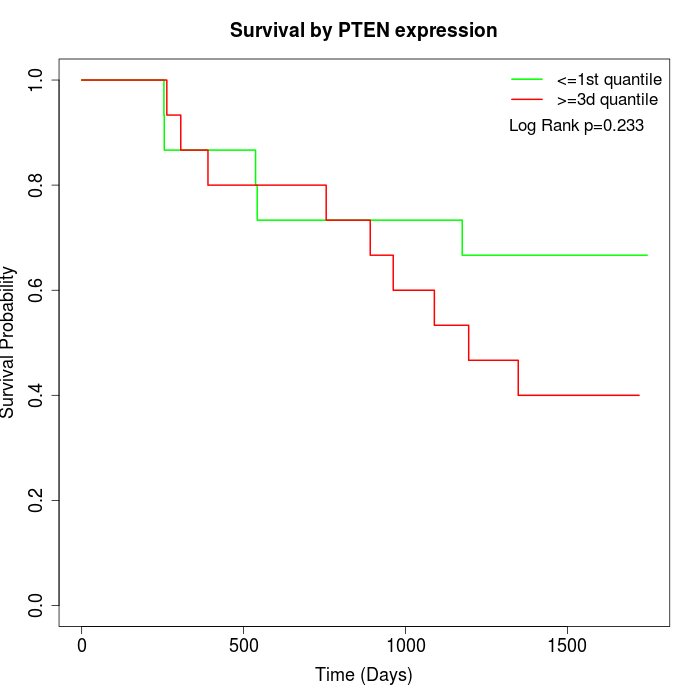
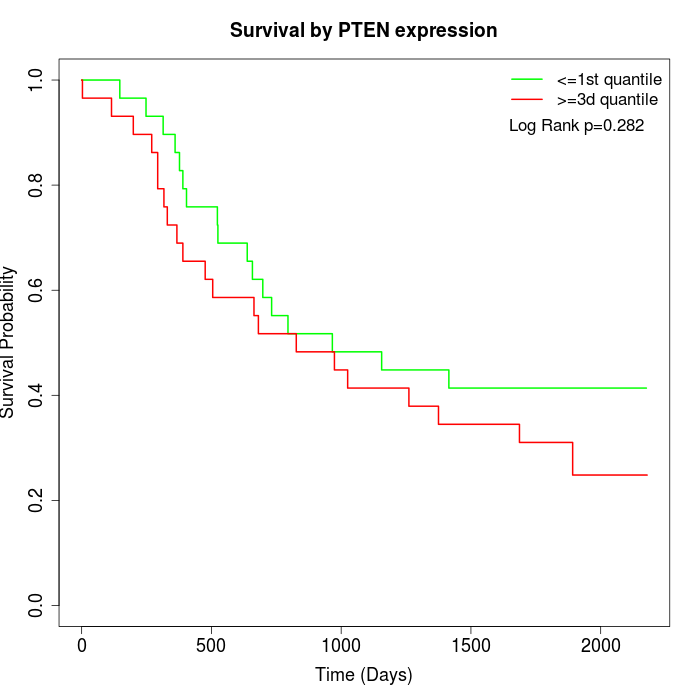
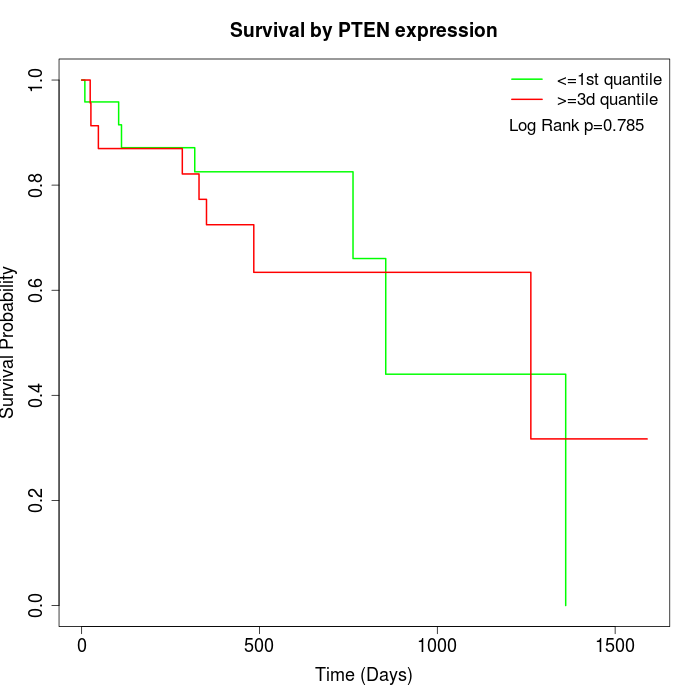
Survival by PTEN expression:
Note: Click image to view full size file.
Copy number change of PTEN:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | PTEN | 5728 | 2 | 7 | 21 | |
| GSE20123 | PTEN | 5728 | 2 | 6 | 22 | |
| GSE43470 | PTEN | 5728 | 0 | 9 | 34 | |
| GSE46452 | PTEN | 5728 | 0 | 11 | 48 | |
| GSE47630 | PTEN | 5728 | 2 | 14 | 24 | |
| GSE54993 | PTEN | 5728 | 7 | 0 | 63 | |
| GSE54994 | PTEN | 5728 | 1 | 11 | 41 | |
| GSE60625 | PTEN | 5728 | 0 | 0 | 11 | |
| GSE74703 | PTEN | 5728 | 0 | 7 | 29 | |
| GSE74704 | PTEN | 5728 | 1 | 4 | 15 | |
| TCGA | PTEN | 5728 | 6 | 30 | 60 |
Total number of gains: 21; Total number of losses: 99; Total Number of normals: 368.
Somatic mutations of PTEN:
Generating mutation plots.
Highly correlated genes for PTEN:
Showing top 20/967 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| PTEN | ZMPSTE24 | 0.739555 | 3 | 0 | 3 |
| PTEN | ZNF416 | 0.7241 | 3 | 0 | 3 |
| PTEN | AXL | 0.718017 | 3 | 0 | 3 |
| PTEN | GRHPR | 0.716537 | 3 | 0 | 3 |
| PTEN | FAM160B1 | 0.71261 | 3 | 0 | 3 |
| PTEN | ATP6V0D1 | 0.696525 | 4 | 0 | 4 |
| PTEN | KLHDC8B | 0.694906 | 4 | 0 | 4 |
| PTEN | GNPDA2 | 0.692526 | 4 | 0 | 4 |
| PTEN | BLOC1S2 | 0.690018 | 4 | 0 | 3 |
| PTEN | SAMD9L | 0.689992 | 3 | 0 | 3 |
| PTEN | MIER1 | 0.68825 | 6 | 0 | 6 |
| PTEN | RNF144B | 0.683428 | 3 | 0 | 3 |
| PTEN | PTPRB | 0.678193 | 4 | 0 | 3 |
| PTEN | RCAN2 | 0.677799 | 5 | 0 | 4 |
| PTEN | ERI2 | 0.675524 | 3 | 0 | 3 |
| PTEN | DNAJB9 | 0.674428 | 4 | 0 | 3 |
| PTEN | ZNF436 | 0.672191 | 4 | 0 | 4 |
| PTEN | DNAJC24 | 0.670028 | 3 | 0 | 3 |
| PTEN | EGR1 | 0.670025 | 3 | 0 | 3 |
| PTEN | CTSO | 0.669796 | 4 | 0 | 4 |
For details and further investigation, click here