| Full name: ADAM metallopeptidase domain 2 | Alias Symbol: PH-30b|PH30|CT15 | ||
| Type: protein-coding gene | Cytoband: 8p11.22 | ||
| Entrez ID: 2515 | HGNC ID: HGNC:198 | Ensembl Gene: ENSG00000104755 | OMIM ID: 601533 |
Screen Evidence:
| |||
Expression of ADAM2:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | ADAM2 | 2515 | 207664_at | 0.0689 | 0.6959 | |
| GSE20347 | ADAM2 | 2515 | 207664_at | -0.0187 | 0.7187 | |
| GSE23400 | ADAM2 | 2515 | 207664_at | -0.0200 | 0.2252 | |
| GSE26886 | ADAM2 | 2515 | 207664_at | 0.0568 | 0.5593 | |
| GSE29001 | ADAM2 | 2515 | 207664_at | -0.0873 | 0.3550 | |
| GSE38129 | ADAM2 | 2515 | 207664_at | -0.0096 | 0.8504 | |
| GSE45670 | ADAM2 | 2515 | 207664_at | 0.0898 | 0.1834 | |
| GSE53622 | ADAM2 | 2515 | 11589 | -0.1524 | 0.2182 | |
| GSE53624 | ADAM2 | 2515 | 11589 | -0.0454 | 0.6230 | |
| GSE63941 | ADAM2 | 2515 | 207664_at | -0.0520 | 0.7191 | |
| GSE77861 | ADAM2 | 2515 | 207664_at | -0.0183 | 0.8345 |
Upregulated datasets: 0; Downregulated datasets: 0.
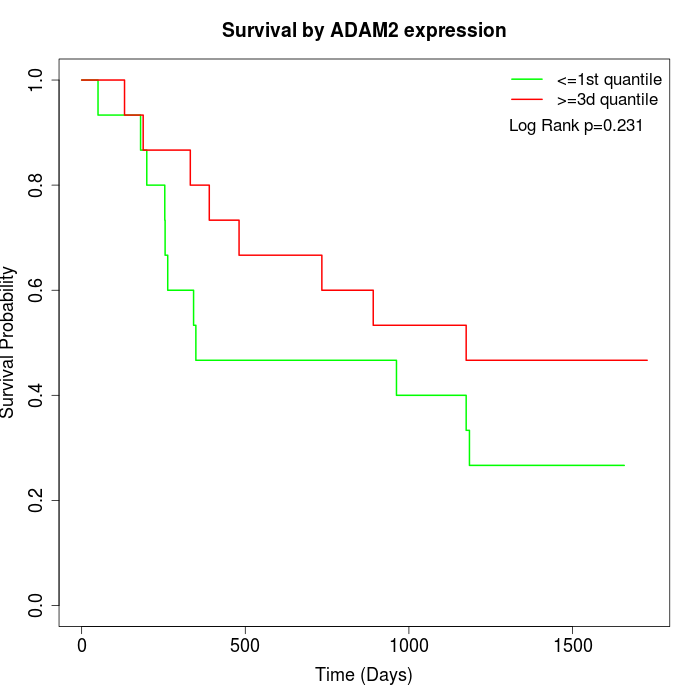
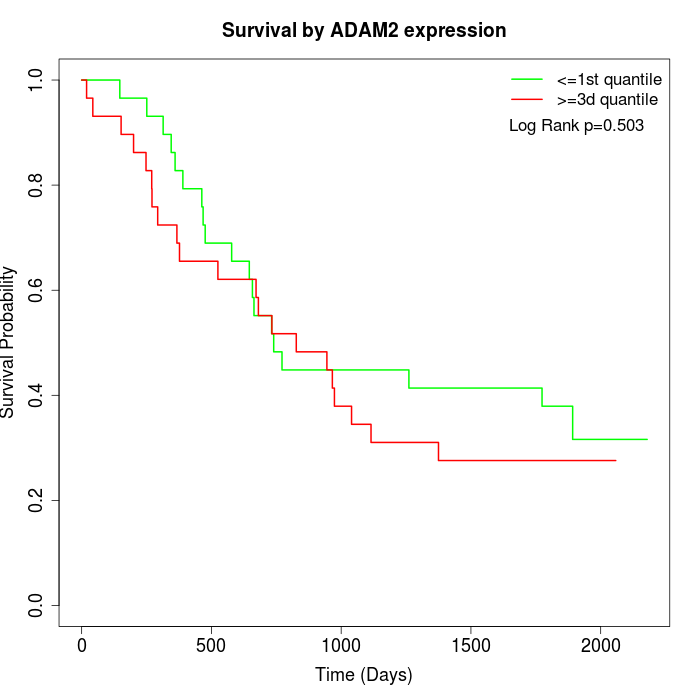
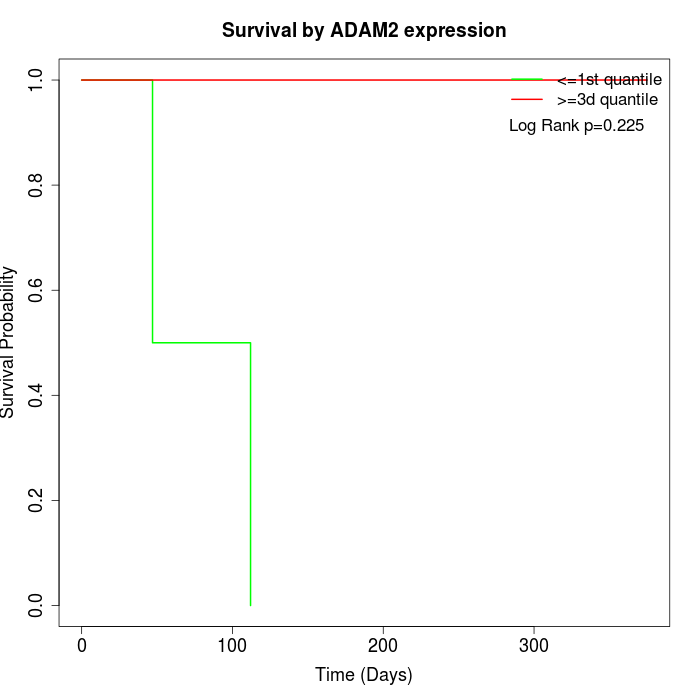
Survival by ADAM2 expression:
Note: Click image to view full size file.
Copy number change of ADAM2:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | ADAM2 | 2515 | 10 | 4 | 16 | |
| GSE20123 | ADAM2 | 2515 | 10 | 4 | 16 | |
| GSE43470 | ADAM2 | 2515 | 8 | 7 | 28 | |
| GSE46452 | ADAM2 | 2515 | 20 | 5 | 34 | |
| GSE47630 | ADAM2 | 2515 | 22 | 1 | 17 | |
| GSE54993 | ADAM2 | 2515 | 2 | 16 | 52 | |
| GSE54994 | ADAM2 | 2515 | 18 | 8 | 27 | |
| GSE60625 | ADAM2 | 2515 | 3 | 0 | 8 | |
| GSE74703 | ADAM2 | 2515 | 8 | 6 | 22 | |
| GSE74704 | ADAM2 | 2515 | 9 | 1 | 10 | |
| TCGA | ADAM2 | 2515 | 36 | 21 | 39 |
Total number of gains: 146; Total number of losses: 73; Total Number of normals: 269.
Somatic mutations of ADAM2:
Generating mutation plots.
Highly correlated genes for ADAM2:
Showing top 20/103 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| ADAM2 | FABP7 | 0.721012 | 3 | 0 | 3 |
| ADAM2 | MSI1 | 0.699686 | 4 | 0 | 4 |
| ADAM2 | ATXN2L | 0.666285 | 4 | 0 | 3 |
| ADAM2 | DSCR4 | 0.642174 | 3 | 0 | 3 |
| ADAM2 | DRD4 | 0.63414 | 4 | 0 | 3 |
| ADAM2 | PRKAR1B | 0.624673 | 3 | 0 | 3 |
| ADAM2 | SERPINA2 | 0.622403 | 3 | 0 | 3 |
| ADAM2 | SEMA3B | 0.616331 | 3 | 0 | 3 |
| ADAM2 | FAM163A | 0.613993 | 4 | 0 | 3 |
| ADAM2 | JAK3 | 0.60377 | 4 | 0 | 3 |
| ADAM2 | AGAP2 | 0.585464 | 4 | 0 | 3 |
| ADAM2 | RAG2 | 0.582682 | 4 | 0 | 4 |
| ADAM2 | MYOG | 0.582573 | 5 | 0 | 4 |
| ADAM2 | DNM3 | 0.581481 | 3 | 0 | 3 |
| ADAM2 | ART1 | 0.579999 | 5 | 0 | 3 |
| ADAM2 | HDAC10 | 0.579763 | 3 | 0 | 3 |
| ADAM2 | HPCAL4 | 0.579085 | 4 | 0 | 3 |
| ADAM2 | TMEM214 | 0.577314 | 4 | 0 | 3 |
| ADAM2 | CHRNB4 | 0.57681 | 3 | 0 | 3 |
| ADAM2 | GABRG3 | 0.576049 | 3 | 0 | 3 |
For details and further investigation, click here