| Full name: all-trans retinoic acid induced differentiation factor | Alias Symbol: HSPC013|p18|APR3 | ||
| Type: protein-coding gene | Cytoband: 2p23.3 | ||
| Entrez ID: 51374 | HGNC ID: HGNC:24090 | Ensembl Gene: ENSG00000138085 | OMIM ID: |
Expression of ATRAID:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | ATRAID | 51374 | 219329_s_at | 0.1496 | 0.7375 | |
| GSE20347 | ATRAID | 51374 | 219329_s_at | 0.4358 | 0.0318 | |
| GSE23400 | ATRAID | 51374 | 219329_s_at | 0.5716 | 0.0000 | |
| GSE26886 | ATRAID | 51374 | 219329_s_at | 0.4150 | 0.0192 | |
| GSE29001 | ATRAID | 51374 | 219329_s_at | 0.4383 | 0.2479 | |
| GSE38129 | ATRAID | 51374 | 219329_s_at | 0.4261 | 0.0061 | |
| GSE45670 | ATRAID | 51374 | 219329_s_at | 0.1396 | 0.4608 | |
| GSE53622 | ATRAID | 51374 | 115414 | 0.4432 | 0.0000 | |
| GSE53624 | ATRAID | 51374 | 53371 | 0.6394 | 0.0000 | |
| GSE63941 | ATRAID | 51374 | 219329_s_at | -0.0070 | 0.9883 | |
| GSE77861 | ATRAID | 51374 | 219329_s_at | 0.2140 | 0.3616 | |
| SRP007169 | ATRAID | 51374 | RNAseq | -0.1116 | 0.7008 | |
| SRP008496 | ATRAID | 51374 | RNAseq | 0.2684 | 0.1787 | |
| SRP064894 | ATRAID | 51374 | RNAseq | 0.4642 | 0.0714 | |
| SRP133303 | ATRAID | 51374 | RNAseq | 0.4332 | 0.0012 | |
| SRP159526 | ATRAID | 51374 | RNAseq | 0.6804 | 0.0180 | |
| SRP193095 | ATRAID | 51374 | RNAseq | 0.1874 | 0.1647 | |
| SRP219564 | ATRAID | 51374 | RNAseq | 0.2661 | 0.4416 |
Upregulated datasets: 0; Downregulated datasets: 0.
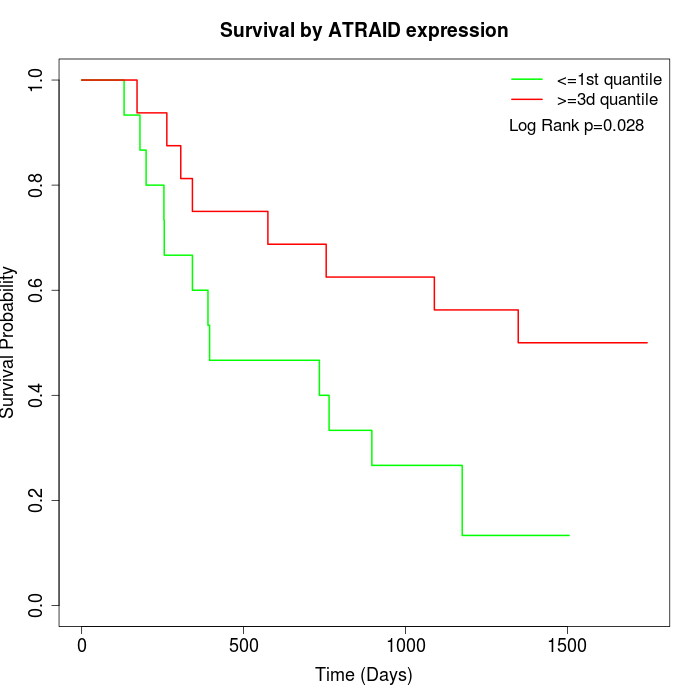
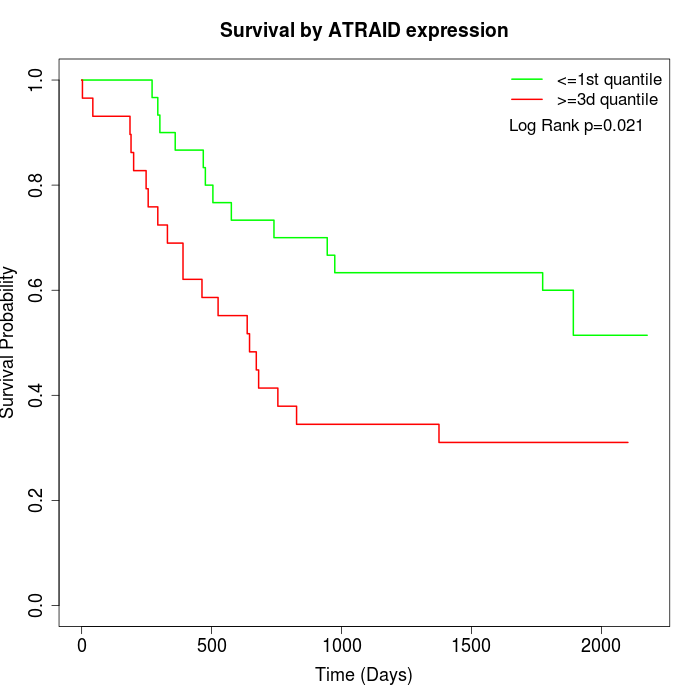
Survival by ATRAID expression:
Note: Click image to view full size file.
Copy number change of ATRAID:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | ATRAID | 51374 | 11 | 2 | 17 | |
| GSE20123 | ATRAID | 51374 | 11 | 2 | 17 | |
| GSE43470 | ATRAID | 51374 | 3 | 1 | 39 | |
| GSE46452 | ATRAID | 51374 | 3 | 4 | 52 | |
| GSE47630 | ATRAID | 51374 | 8 | 0 | 32 | |
| GSE54993 | ATRAID | 51374 | 0 | 6 | 64 | |
| GSE54994 | ATRAID | 51374 | 10 | 0 | 43 | |
| GSE60625 | ATRAID | 51374 | 0 | 3 | 8 | |
| GSE74703 | ATRAID | 51374 | 3 | 0 | 33 | |
| GSE74704 | ATRAID | 51374 | 9 | 1 | 10 | |
| TCGA | ATRAID | 51374 | 37 | 2 | 57 |
Total number of gains: 95; Total number of losses: 21; Total Number of normals: 372.
Somatic mutations of ATRAID:
Generating mutation plots.
Highly correlated genes for ATRAID:
Showing top 20/800 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| ATRAID | KRTCAP3 | 0.707887 | 4 | 0 | 4 |
| ATRAID | ZC3H12C | 0.682834 | 3 | 0 | 3 |
| ATRAID | RCN2 | 0.675716 | 7 | 0 | 7 |
| ATRAID | ZDHHC4 | 0.672778 | 9 | 0 | 7 |
| ATRAID | SSR2 | 0.66464 | 7 | 0 | 7 |
| ATRAID | TMEM60 | 0.661225 | 4 | 0 | 3 |
| ATRAID | MPV17 | 0.658184 | 10 | 0 | 9 |
| ATRAID | TRMT61B | 0.65734 | 8 | 0 | 8 |
| ATRAID | PREB | 0.650104 | 7 | 0 | 5 |
| ATRAID | DGUOK | 0.648089 | 11 | 0 | 9 |
| ATRAID | LAMTOR4 | 0.64482 | 5 | 0 | 4 |
| ATRAID | KIAA2013 | 0.644035 | 4 | 0 | 4 |
| ATRAID | HLTF | 0.643332 | 8 | 0 | 8 |
| ATRAID | ARSJ | 0.642428 | 3 | 0 | 3 |
| ATRAID | COX7A2L | 0.641772 | 10 | 0 | 8 |
| ATRAID | DYNC2LI1 | 0.640314 | 5 | 0 | 4 |
| ATRAID | DENND4A | 0.637954 | 3 | 0 | 3 |
| ATRAID | LSM2 | 0.63795 | 7 | 0 | 6 |
| ATRAID | TTC30B | 0.635969 | 4 | 0 | 3 |
| ATRAID | DHX40 | 0.633155 | 4 | 0 | 3 |
For details and further investigation, click here