| Full name: CACNA1G antisense RNA 1 | Alias Symbol: CAS1 | ||
| Type: non-coding RNA | Cytoband: 17q21.33 | ||
| Entrez ID: 253962 | HGNC ID: HGNC:27377 | Ensembl Gene: ENSG00000250107 | OMIM ID: |
Expression of CACNA1G-AS1:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | CACNA1G-AS1 | 253962 | 1559061_at | 0.0316 | 0.9082 | |
| GSE26886 | CACNA1G-AS1 | 253962 | 1559061_at | -0.0049 | 0.9777 | |
| GSE45670 | CACNA1G-AS1 | 253962 | 1559061_at | -0.0658 | 0.4783 | |
| GSE53622 | CACNA1G-AS1 | 253962 | 143944 | -0.1799 | 0.0094 | |
| GSE53624 | CACNA1G-AS1 | 253962 | 143944 | -0.2507 | 0.0019 | |
| GSE63941 | CACNA1G-AS1 | 253962 | 1559061_at | 0.2327 | 0.1277 | |
| GSE77861 | CACNA1G-AS1 | 253962 | 1559061_at | -0.1988 | 0.0883 |
Upregulated datasets: 0; Downregulated datasets: 0.
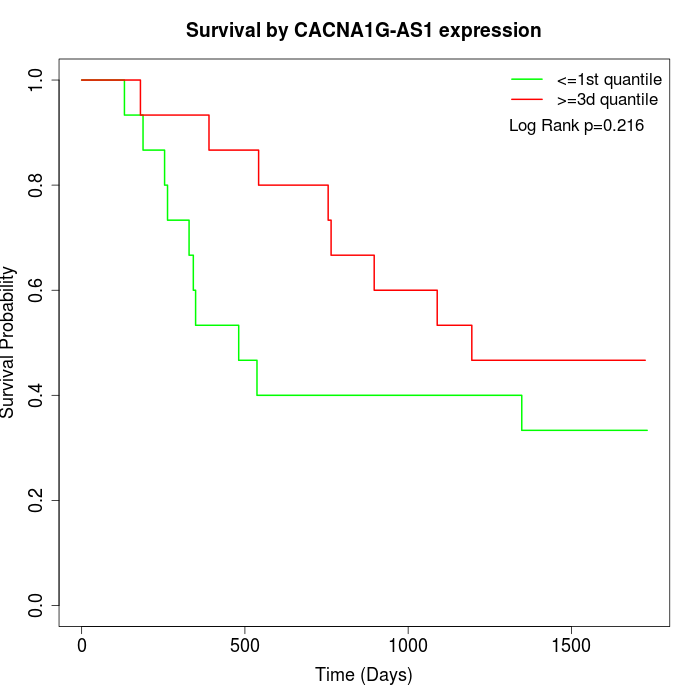
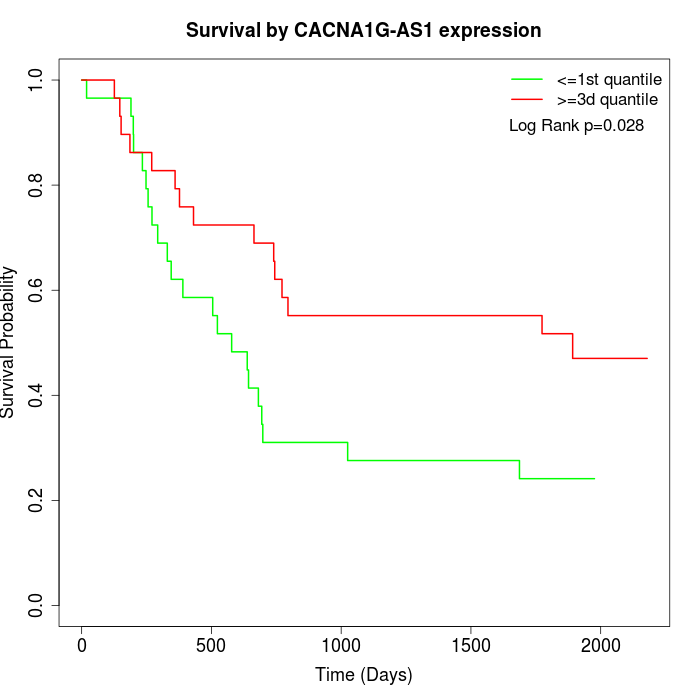
Survival by CACNA1G-AS1 expression:
Note: Click image to view full size file.
Copy number change of CACNA1G-AS1:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | CACNA1G-AS1 | 253962 | 5 | 2 | 23 | |
| GSE20123 | CACNA1G-AS1 | 253962 | 5 | 2 | 23 | |
| GSE43470 | CACNA1G-AS1 | 253962 | 3 | 2 | 38 | |
| GSE46452 | CACNA1G-AS1 | 253962 | 32 | 0 | 27 | |
| GSE47630 | CACNA1G-AS1 | 253962 | 9 | 0 | 31 | |
| GSE54993 | CACNA1G-AS1 | 253962 | 2 | 4 | 64 | |
| GSE54994 | CACNA1G-AS1 | 253962 | 9 | 6 | 38 | |
| GSE60625 | CACNA1G-AS1 | 253962 | 4 | 0 | 7 | |
| GSE74703 | CACNA1G-AS1 | 253962 | 3 | 2 | 31 | |
| GSE74704 | CACNA1G-AS1 | 253962 | 4 | 1 | 15 |
Total number of gains: 76; Total number of losses: 19; Total Number of normals: 297.
Somatic mutations of CACNA1G-AS1:
Generating mutation plots.
Highly correlated genes for CACNA1G-AS1:
Showing top 20/109 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| CACNA1G-AS1 | SLC24A4 | 0.702477 | 3 | 0 | 3 |
| CACNA1G-AS1 | RHO | 0.692235 | 3 | 0 | 3 |
| CACNA1G-AS1 | TXNDC2 | 0.684055 | 3 | 0 | 3 |
| CACNA1G-AS1 | DCD | 0.682591 | 4 | 0 | 4 |
| CACNA1G-AS1 | LINC00421 | 0.666862 | 3 | 0 | 3 |
| CACNA1G-AS1 | VGLL2 | 0.665318 | 3 | 0 | 3 |
| CACNA1G-AS1 | NPPB | 0.659861 | 3 | 0 | 3 |
| CACNA1G-AS1 | POM121L12 | 0.656048 | 3 | 0 | 3 |
| CACNA1G-AS1 | CHRNA6 | 0.650911 | 4 | 0 | 4 |
| CACNA1G-AS1 | GRIK5 | 0.647337 | 3 | 0 | 3 |
| CACNA1G-AS1 | DLGAP3 | 0.646704 | 3 | 0 | 3 |
| CACNA1G-AS1 | NKX6-3 | 0.645774 | 3 | 0 | 3 |
| CACNA1G-AS1 | BTNL8 | 0.644321 | 4 | 0 | 3 |
| CACNA1G-AS1 | IL36B | 0.642724 | 3 | 0 | 3 |
| CACNA1G-AS1 | MATN1 | 0.640243 | 3 | 0 | 3 |
| CACNA1G-AS1 | C2orf72 | 0.639853 | 5 | 0 | 4 |
| CACNA1G-AS1 | SERPINA10 | 0.63981 | 3 | 0 | 3 |
| CACNA1G-AS1 | FAM71A | 0.629692 | 4 | 0 | 3 |
| CACNA1G-AS1 | FAM170B-AS1 | 0.625567 | 4 | 0 | 3 |
| CACNA1G-AS1 | CLEC1B | 0.623928 | 4 | 0 | 3 |
For details and further investigation, click here