| Full name: cyclin H | Alias Symbol: p34|p37|CycH | ||
| Type: protein-coding gene | Cytoband: 5q14.3 | ||
| Entrez ID: 902 | HGNC ID: HGNC:1594 | Ensembl Gene: ENSG00000134480 | OMIM ID: 601953 |
Screen Evidence:
| |||
CCNH involved pathways:
| KEGG pathway | Description | View |
|---|---|---|
| hsa04110 | Cell cycle |
Expression of CCNH:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | CCNH | 902 | 204093_at | -0.4705 | 0.5160 | |
| GSE20347 | CCNH | 902 | 204093_at | -0.3153 | 0.0150 | |
| GSE23400 | CCNH | 902 | 204093_at | 0.0252 | 0.7113 | |
| GSE26886 | CCNH | 902 | 204093_at | 0.4296 | 0.0079 | |
| GSE29001 | CCNH | 902 | 204093_at | -0.0454 | 0.8712 | |
| GSE38129 | CCNH | 902 | 204093_at | -0.1460 | 0.2721 | |
| GSE45670 | CCNH | 902 | 204093_at | -0.0623 | 0.7029 | |
| GSE53622 | CCNH | 902 | 19200 | -0.3966 | 0.0000 | |
| GSE53624 | CCNH | 902 | 19200 | -0.3699 | 0.0000 | |
| GSE63941 | CCNH | 902 | 204093_at | -0.4184 | 0.2777 | |
| GSE77861 | CCNH | 902 | 204093_at | -0.3244 | 0.2873 | |
| GSE97050 | CCNH | 902 | A_23_P30338 | -0.0803 | 0.7360 | |
| SRP007169 | CCNH | 902 | RNAseq | 0.0660 | 0.8858 | |
| SRP008496 | CCNH | 902 | RNAseq | -0.2818 | 0.3273 | |
| SRP064894 | CCNH | 902 | RNAseq | -0.1939 | 0.1906 | |
| SRP133303 | CCNH | 902 | RNAseq | -0.2390 | 0.0967 | |
| SRP159526 | CCNH | 902 | RNAseq | -0.5393 | 0.0559 | |
| SRP193095 | CCNH | 902 | RNAseq | -0.3916 | 0.0004 | |
| SRP219564 | CCNH | 902 | RNAseq | -0.2128 | 0.5497 | |
| TCGA | CCNH | 902 | RNAseq | -0.1142 | 0.0408 |
Upregulated datasets: 0; Downregulated datasets: 0.
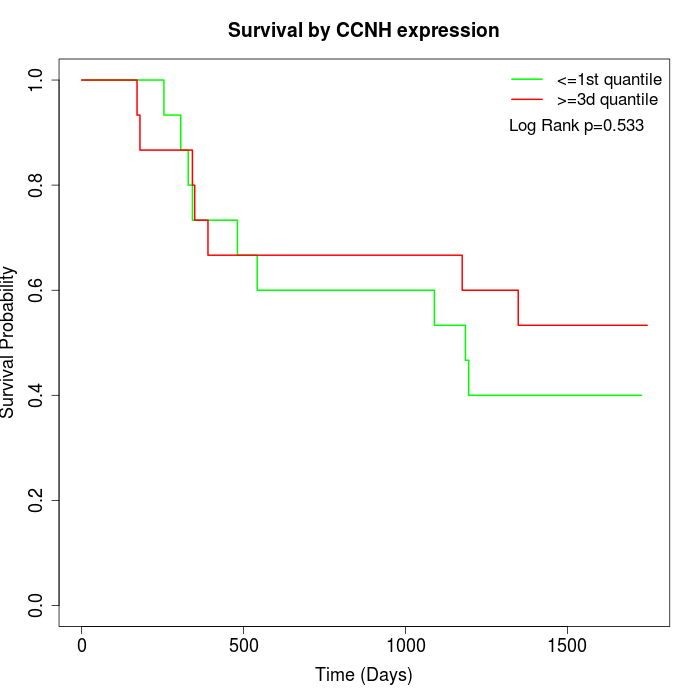
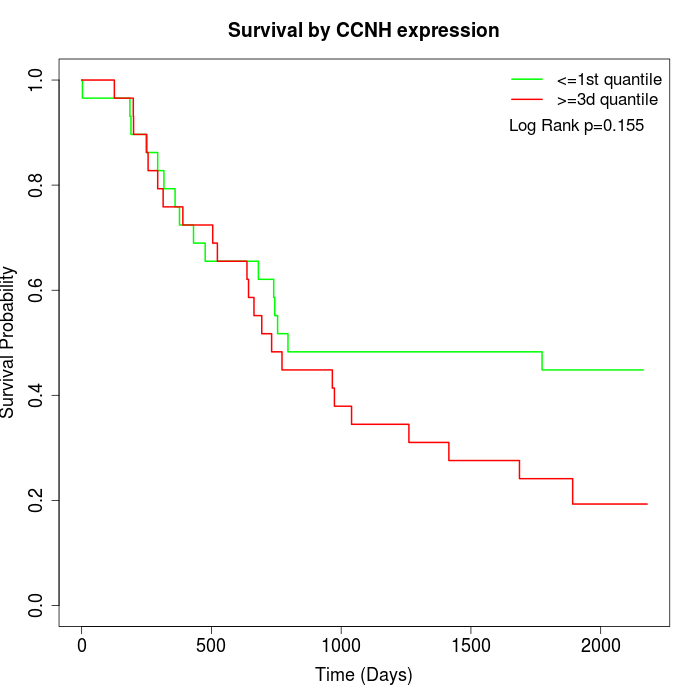
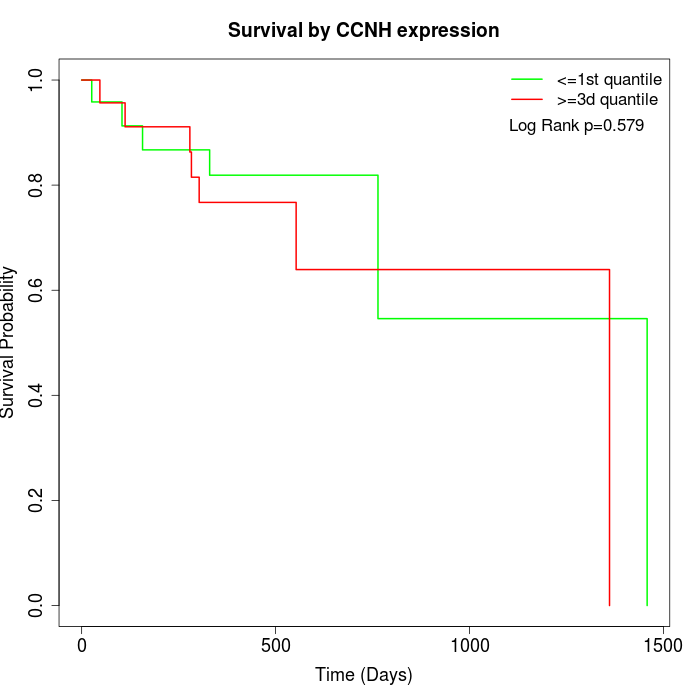
Survival by CCNH expression:
Note: Click image to view full size file.
Copy number change of CCNH:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | CCNH | 902 | 1 | 14 | 15 | |
| GSE20123 | CCNH | 902 | 1 | 14 | 15 | |
| GSE43470 | CCNH | 902 | 2 | 10 | 31 | |
| GSE46452 | CCNH | 902 | 0 | 28 | 31 | |
| GSE47630 | CCNH | 902 | 1 | 20 | 19 | |
| GSE54993 | CCNH | 902 | 8 | 1 | 61 | |
| GSE54994 | CCNH | 902 | 1 | 18 | 34 | |
| GSE60625 | CCNH | 902 | 0 | 0 | 11 | |
| GSE74703 | CCNH | 902 | 1 | 8 | 27 | |
| GSE74704 | CCNH | 902 | 1 | 7 | 12 | |
| TCGA | CCNH | 902 | 2 | 50 | 44 |
Total number of gains: 18; Total number of losses: 170; Total Number of normals: 300.
Somatic mutations of CCNH:
Generating mutation plots.
Highly correlated genes for CCNH:
Showing top 20/439 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| CCNH | PFDN1 | 0.789278 | 3 | 0 | 3 |
| CCNH | RAB18 | 0.771282 | 3 | 0 | 3 |
| CCNH | FMR1 | 0.763652 | 3 | 0 | 3 |
| CCNH | ARSK | 0.762481 | 3 | 0 | 3 |
| CCNH | CD47 | 0.76147 | 3 | 0 | 3 |
| CCNH | TTC8 | 0.751854 | 3 | 0 | 3 |
| CCNH | GCSH | 0.745987 | 3 | 0 | 3 |
| CCNH | FAM126A | 0.739973 | 3 | 0 | 3 |
| CCNH | EGLN1 | 0.73789 | 3 | 0 | 3 |
| CCNH | CCNDBP1 | 0.72929 | 3 | 0 | 3 |
| CCNH | ADAMTS18 | 0.728685 | 3 | 0 | 3 |
| CCNH | ZBTB26 | 0.725221 | 3 | 0 | 3 |
| CCNH | SGTB | 0.725203 | 4 | 0 | 4 |
| CCNH | RNF139 | 0.707359 | 3 | 0 | 3 |
| CCNH | NR3C1 | 0.706197 | 4 | 0 | 3 |
| CCNH | TUBB2A | 0.69966 | 5 | 0 | 4 |
| CCNH | FSD1L | 0.696334 | 3 | 0 | 3 |
| CCNH | UBXN10 | 0.686075 | 3 | 0 | 3 |
| CCNH | LYVE1 | 0.684319 | 4 | 0 | 3 |
| CCNH | CCDC51 | 0.68095 | 3 | 0 | 3 |
For details and further investigation, click here