| Full name: endothelin receptor type B | Alias Symbol: ETB | ||
| Type: protein-coding gene | Cytoband: 13q22.3 | ||
| Entrez ID: 1910 | HGNC ID: HGNC:3180 | Ensembl Gene: ENSG00000136160 | OMIM ID: 131244 |
| Related drugs: A-192621, AMBRISENTAN, ATRASENTAN, BOSENTAN, CHEMBL3188091, CHEMBL37807, CHEMBL72410, DARUSENTAN, ENRASENTAN, GALANTAMINE... [more] | |||
EDNRB involved pathways:
| KEGG pathway | Description | View |
|---|---|---|
| hsa04020 | Calcium signaling pathway | |
| hsa04022 | cGMP-PKG signaling pathway | |
| hsa04916 | Melanogenesis |
Expression of EDNRB:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | EDNRB | 1910 | 204271_s_at | -0.6953 | 0.3205 | |
| GSE20347 | EDNRB | 1910 | 204271_s_at | 0.0683 | 0.5437 | |
| GSE23400 | EDNRB | 1910 | 204271_s_at | -0.0316 | 0.4800 | |
| GSE26886 | EDNRB | 1910 | 204271_s_at | -0.0606 | 0.8109 | |
| GSE29001 | EDNRB | 1910 | 204271_s_at | 0.1223 | 0.6842 | |
| GSE38129 | EDNRB | 1910 | 204271_s_at | -0.5013 | 0.1532 | |
| GSE45670 | EDNRB | 1910 | 204271_s_at | -1.1817 | 0.0008 | |
| GSE53622 | EDNRB | 1910 | 34048 | -0.9612 | 0.0000 | |
| GSE53624 | EDNRB | 1910 | 34048 | -0.6674 | 0.0000 | |
| GSE63941 | EDNRB | 1910 | 204271_s_at | -7.2503 | 0.0000 | |
| GSE77861 | EDNRB | 1910 | 204271_s_at | 0.0331 | 0.6996 | |
| GSE97050 | EDNRB | 1910 | A_24_P330263 | -0.5179 | 0.3273 | |
| SRP007169 | EDNRB | 1910 | RNAseq | 0.7110 | 0.2409 | |
| SRP064894 | EDNRB | 1910 | RNAseq | 0.6966 | 0.0985 | |
| SRP133303 | EDNRB | 1910 | RNAseq | -0.8101 | 0.0000 | |
| SRP159526 | EDNRB | 1910 | RNAseq | 0.5518 | 0.3913 | |
| SRP193095 | EDNRB | 1910 | RNAseq | -0.3527 | 0.3675 | |
| SRP219564 | EDNRB | 1910 | RNAseq | -0.1986 | 0.7784 | |
| TCGA | EDNRB | 1910 | RNAseq | -0.7994 | 0.0000 |
Upregulated datasets: 0; Downregulated datasets: 2.
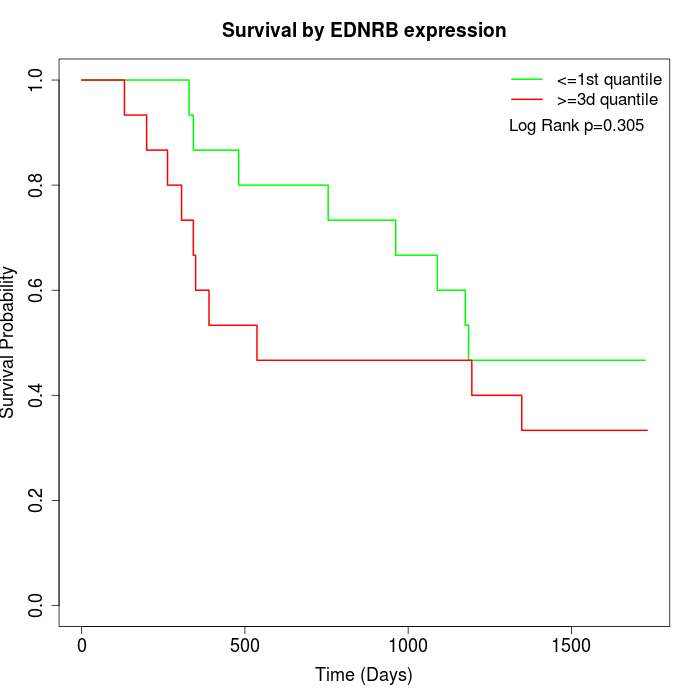
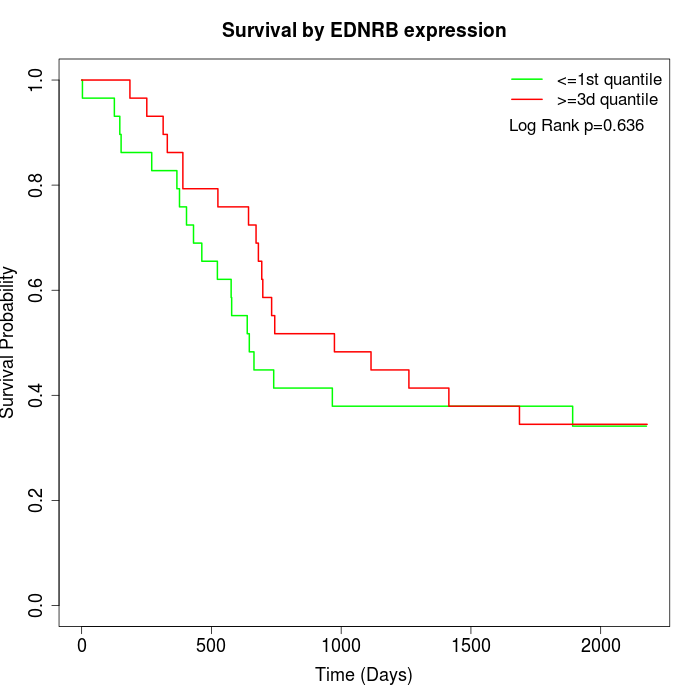
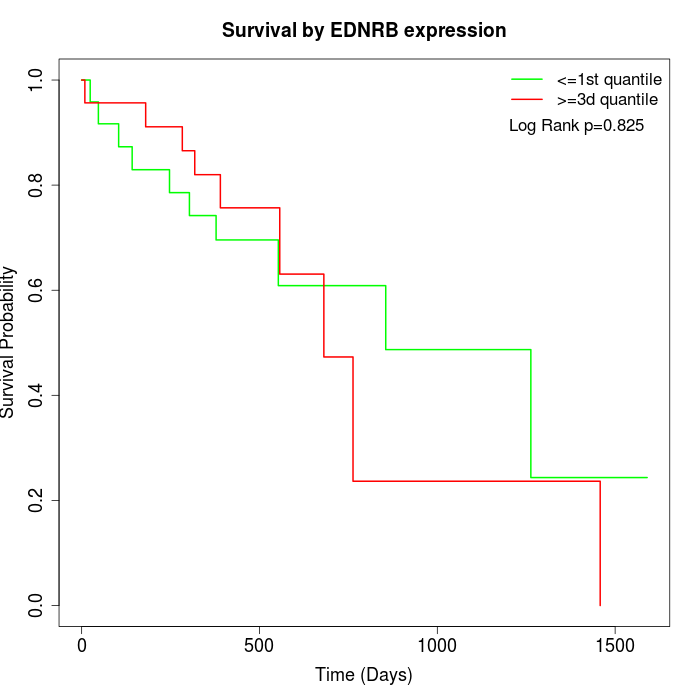
Survival by EDNRB expression:
Note: Click image to view full size file.
Copy number change of EDNRB:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | EDNRB | 1910 | 4 | 11 | 15 | |
| GSE20123 | EDNRB | 1910 | 4 | 10 | 16 | |
| GSE43470 | EDNRB | 1910 | 4 | 11 | 28 | |
| GSE46452 | EDNRB | 1910 | 0 | 32 | 27 | |
| GSE47630 | EDNRB | 1910 | 3 | 27 | 10 | |
| GSE54993 | EDNRB | 1910 | 11 | 3 | 56 | |
| GSE54994 | EDNRB | 1910 | 3 | 14 | 36 | |
| GSE60625 | EDNRB | 1910 | 0 | 3 | 8 | |
| GSE74703 | EDNRB | 1910 | 3 | 9 | 24 | |
| GSE74704 | EDNRB | 1910 | 2 | 9 | 9 | |
| TCGA | EDNRB | 1910 | 12 | 33 | 51 |
Total number of gains: 46; Total number of losses: 162; Total Number of normals: 280.
Somatic mutations of EDNRB:
Generating mutation plots.
Highly correlated genes for EDNRB:
Showing top 20/1201 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| EDNRB | C11orf96 | 0.835984 | 4 | 0 | 4 |
| EDNRB | SLC39A13 | 0.808948 | 3 | 0 | 3 |
| EDNRB | FIBIN | 0.785657 | 5 | 0 | 5 |
| EDNRB | TNFSF12 | 0.778885 | 3 | 0 | 3 |
| EDNRB | RSPO3 | 0.774864 | 6 | 0 | 6 |
| EDNRB | MIER1 | 0.774166 | 3 | 0 | 3 |
| EDNRB | UBXN10 | 0.77294 | 3 | 0 | 3 |
| EDNRB | KCNT2 | 0.765966 | 6 | 0 | 6 |
| EDNRB | PEG3 | 0.765951 | 6 | 0 | 6 |
| EDNRB | LINC01279 | 0.76098 | 3 | 0 | 3 |
| EDNRB | SMPD1 | 0.760332 | 4 | 0 | 4 |
| EDNRB | UBE2Q2 | 0.757308 | 3 | 0 | 3 |
| EDNRB | SLC35D2 | 0.754204 | 3 | 0 | 3 |
| EDNRB | FENDRR | 0.751806 | 5 | 0 | 5 |
| EDNRB | ING1 | 0.750423 | 3 | 0 | 3 |
| EDNRB | KLF2 | 0.748476 | 8 | 0 | 8 |
| EDNRB | BHMT2 | 0.747627 | 10 | 0 | 10 |
| EDNRB | EMCN | 0.746893 | 5 | 0 | 4 |
| EDNRB | RSPH9 | 0.744986 | 3 | 0 | 3 |
| EDNRB | MAGI2-AS3 | 0.743375 | 6 | 0 | 5 |
For details and further investigation, click here