| Full name: EYA transcriptional coactivator and phosphatase 4 | Alias Symbol: | ||
| Type: protein-coding gene | Cytoband: 6q23.2 | ||
| Entrez ID: 2070 | HGNC ID: HGNC:3522 | Ensembl Gene: ENSG00000112319 | OMIM ID: 603550 |
Expression of EYA4:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | EYA4 | 2070 | 238877_at | 1.6510 | 0.1165 | |
| GSE20347 | EYA4 | 2070 | 207327_at | 0.3681 | 0.0267 | |
| GSE23400 | EYA4 | 2070 | 207327_at | 0.0669 | 0.1527 | |
| GSE26886 | EYA4 | 2070 | 207327_at | 0.1569 | 0.1445 | |
| GSE29001 | EYA4 | 2070 | 207327_at | 0.4926 | 0.0805 | |
| GSE38129 | EYA4 | 2070 | 207327_at | 0.0996 | 0.4979 | |
| GSE45670 | EYA4 | 2070 | 207327_at | 0.1021 | 0.7105 | |
| GSE53622 | EYA4 | 2070 | 7067 | 0.4575 | 0.0213 | |
| GSE53624 | EYA4 | 2070 | 16687 | 1.3658 | 0.0000 | |
| GSE63941 | EYA4 | 2070 | 238877_at | -4.1468 | 0.0039 | |
| GSE77861 | EYA4 | 2070 | 238877_at | 0.7719 | 0.1173 | |
| SRP133303 | EYA4 | 2070 | RNAseq | 1.2188 | 0.0020 | |
| SRP159526 | EYA4 | 2070 | RNAseq | 1.5405 | 0.0001 | |
| TCGA | EYA4 | 2070 | RNAseq | 0.0946 | 0.8038 |
Upregulated datasets: 3; Downregulated datasets: 1.
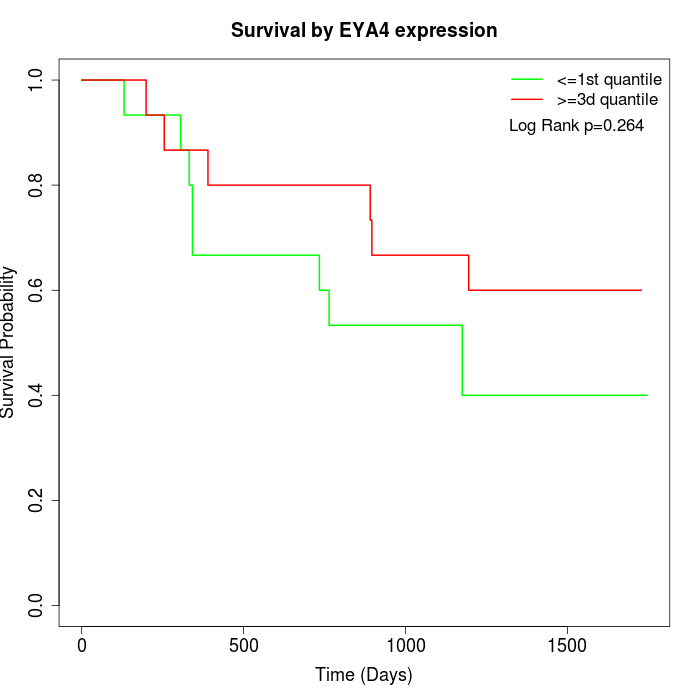
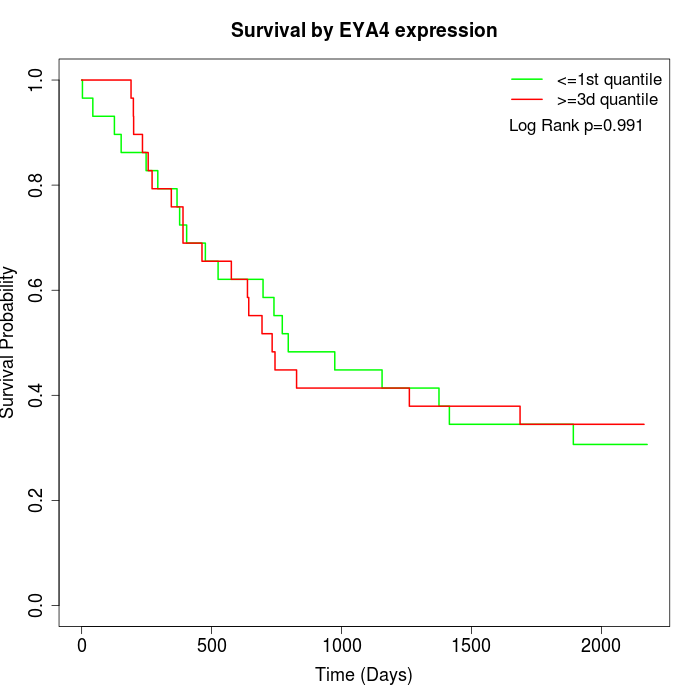
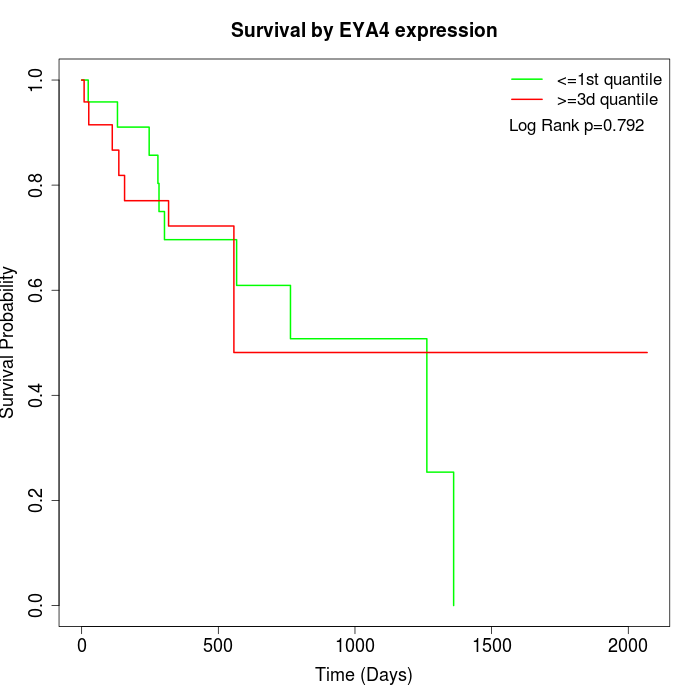
Survival by EYA4 expression:
Note: Click image to view full size file.
Copy number change of EYA4:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | EYA4 | 2070 | 3 | 4 | 23 | |
| GSE20123 | EYA4 | 2070 | 3 | 3 | 24 | |
| GSE43470 | EYA4 | 2070 | 5 | 0 | 38 | |
| GSE46452 | EYA4 | 2070 | 3 | 10 | 46 | |
| GSE47630 | EYA4 | 2070 | 9 | 4 | 27 | |
| GSE54993 | EYA4 | 2070 | 3 | 2 | 65 | |
| GSE54994 | EYA4 | 2070 | 9 | 7 | 37 | |
| GSE60625 | EYA4 | 2070 | 0 | 1 | 10 | |
| GSE74703 | EYA4 | 2070 | 4 | 0 | 32 | |
| GSE74704 | EYA4 | 2070 | 1 | 1 | 18 | |
| TCGA | EYA4 | 2070 | 11 | 18 | 67 |
Total number of gains: 51; Total number of losses: 50; Total Number of normals: 387.
Somatic mutations of EYA4:
Generating mutation plots.
Highly correlated genes for EYA4:
Showing top 20/161 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| EYA4 | SLC23A2 | 0.684394 | 3 | 0 | 3 |
| EYA4 | RBP1 | 0.676614 | 3 | 0 | 3 |
| EYA4 | PIGG | 0.649837 | 3 | 0 | 3 |
| EYA4 | IL24 | 0.647411 | 4 | 0 | 4 |
| EYA4 | SUGCT | 0.644836 | 4 | 0 | 3 |
| EYA4 | TBCE | 0.642057 | 3 | 0 | 3 |
| EYA4 | MRGBP | 0.640112 | 4 | 0 | 3 |
| EYA4 | FOXN2 | 0.638692 | 4 | 0 | 3 |
| EYA4 | TIMM17A | 0.637756 | 3 | 0 | 3 |
| EYA4 | SGTB | 0.629567 | 3 | 0 | 3 |
| EYA4 | MTERF3 | 0.625726 | 3 | 0 | 3 |
| EYA4 | CYP24A1 | 0.623393 | 4 | 0 | 4 |
| EYA4 | ERVMER34-1 | 0.622836 | 4 | 0 | 3 |
| EYA4 | NAV1 | 0.621068 | 3 | 0 | 3 |
| EYA4 | GLS | 0.617243 | 6 | 0 | 4 |
| EYA4 | LINC01252 | 0.616056 | 4 | 0 | 3 |
| EYA4 | PCGF1 | 0.60964 | 3 | 0 | 3 |
| EYA4 | TNFRSF10B | 0.609606 | 5 | 0 | 3 |
| EYA4 | GUCY1A2 | 0.607818 | 4 | 0 | 4 |
| EYA4 | AFAP1 | 0.606724 | 4 | 0 | 3 |
For details and further investigation, click here