| Full name: lysine methyltransferase 2C | Alias Symbol: KIAA1506|HALR | ||
| Type: protein-coding gene | Cytoband: 7q36.1 | ||
| Entrez ID: 58508 | HGNC ID: HGNC:13726 | Ensembl Gene: ENSG00000055609 | OMIM ID: 606833 |
Screen Evidence:
| |||
Expression of KMT2C:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | KMT2C | 58508 | 222415_at | -0.3924 | 0.2131 | |
| GSE26886 | KMT2C | 58508 | 222415_at | 0.5764 | 0.0283 | |
| GSE45670 | KMT2C | 58508 | 222415_at | -0.2625 | 0.0338 | |
| GSE53622 | KMT2C | 58508 | 4523 | -0.3312 | 0.0000 | |
| GSE53624 | KMT2C | 58508 | 35231 | -0.4868 | 0.0000 | |
| GSE63941 | KMT2C | 58508 | 222415_at | -0.4816 | 0.3861 | |
| GSE77861 | KMT2C | 58508 | 222415_at | -0.1165 | 0.8003 | |
| SRP007169 | KMT2C | 58508 | RNAseq | 0.4671 | 0.2410 | |
| SRP008496 | KMT2C | 58508 | RNAseq | 0.3971 | 0.0947 | |
| SRP064894 | KMT2C | 58508 | RNAseq | -0.7197 | 0.0089 | |
| SRP133303 | KMT2C | 58508 | RNAseq | -0.2795 | 0.0014 | |
| SRP159526 | KMT2C | 58508 | RNAseq | -0.7260 | 0.0006 | |
| SRP193095 | KMT2C | 58508 | RNAseq | -0.3054 | 0.0049 | |
| SRP219564 | KMT2C | 58508 | RNAseq | -0.7309 | 0.0566 |
Upregulated datasets: 0; Downregulated datasets: 0.
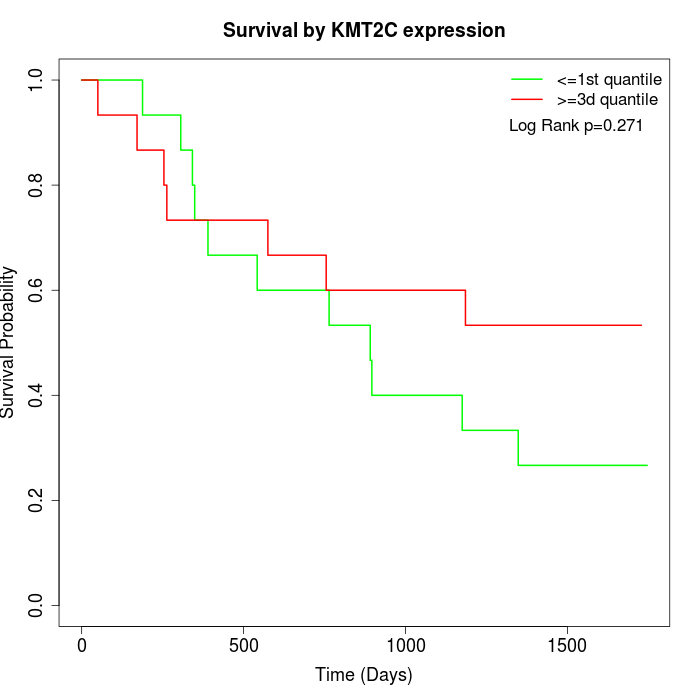
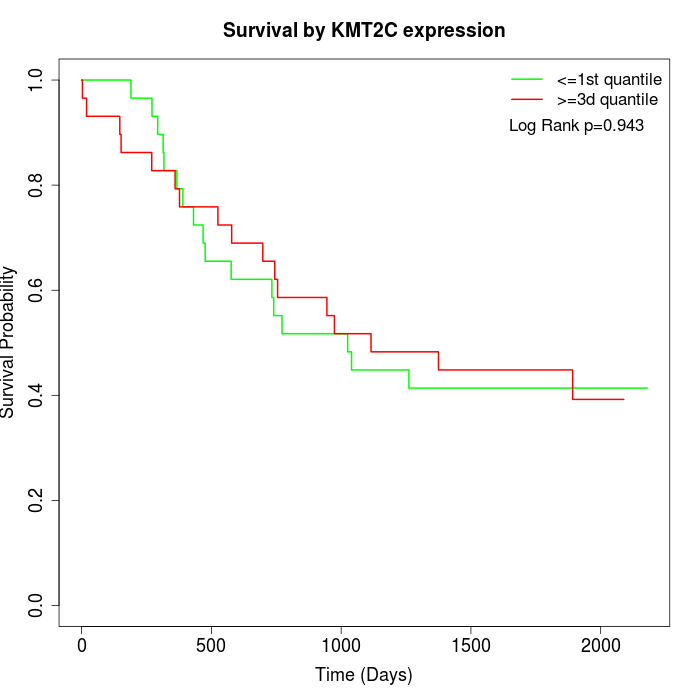
Survival by KMT2C expression:
Note: Click image to view full size file.
Copy number change of KMT2C:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | KMT2C | 58508 | 2 | 4 | 24 | |
| GSE20123 | KMT2C | 58508 | 2 | 4 | 24 | |
| GSE43470 | KMT2C | 58508 | 2 | 5 | 36 | |
| GSE46452 | KMT2C | 58508 | 7 | 2 | 50 | |
| GSE47630 | KMT2C | 58508 | 6 | 9 | 25 | |
| GSE54993 | KMT2C | 58508 | 3 | 3 | 64 | |
| GSE54994 | KMT2C | 58508 | 6 | 11 | 36 | |
| GSE60625 | KMT2C | 58508 | 0 | 0 | 11 | |
| GSE74703 | KMT2C | 58508 | 2 | 4 | 30 | |
| GSE74704 | KMT2C | 58508 | 1 | 4 | 15 | |
| TCGA | KMT2C | 58508 | 25 | 30 | 41 |
Total number of gains: 56; Total number of losses: 76; Total Number of normals: 356.
Somatic mutations of KMT2C:
Generating mutation plots.
Highly correlated genes for KMT2C:
Showing top 20/64 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| KMT2C | CREBBP | 0.728573 | 3 | 0 | 3 |
| KMT2C | BRAF | 0.690827 | 5 | 0 | 5 |
| KMT2C | ZNF277 | 0.63128 | 3 | 0 | 3 |
| KMT2C | CREBZF | 0.631262 | 3 | 0 | 3 |
| KMT2C | MTERF2 | 0.627134 | 3 | 0 | 3 |
| KMT2C | LANCL1 | 0.626092 | 3 | 0 | 3 |
| KMT2C | LUC7L2 | 0.623945 | 5 | 0 | 5 |
| KMT2C | KPNA4 | 0.623304 | 3 | 0 | 3 |
| KMT2C | GSTK1 | 0.620576 | 4 | 0 | 3 |
| KMT2C | ARHGAP35 | 0.609519 | 4 | 0 | 4 |
| KMT2C | CNOT4 | 0.607736 | 4 | 0 | 3 |
| KMT2C | ZADH2 | 0.604281 | 4 | 0 | 3 |
| KMT2C | WASL | 0.603642 | 4 | 0 | 3 |
| KMT2C | GOLGB1 | 0.601147 | 4 | 0 | 3 |
| KMT2C | ZNF717 | 0.598955 | 4 | 0 | 3 |
| KMT2C | CTR9 | 0.597292 | 3 | 0 | 3 |
| KMT2C | EPHX2 | 0.593135 | 4 | 0 | 3 |
| KMT2C | TIMM10B | 0.592008 | 3 | 0 | 3 |
| KMT2C | ASH1L | 0.591055 | 5 | 0 | 3 |
| KMT2C | CCDC146 | 0.586302 | 3 | 0 | 3 |
For details and further investigation, click here