| Full name: Rho GTPase activating protein 35 | Alias Symbol: GRF-1|p190ARhoGAP|P190A|KIAA1722|p190RhoGAP | ||
| Type: protein-coding gene | Cytoband: 19q13.32 | ||
| Entrez ID: 2909 | HGNC ID: HGNC:4591 | Ensembl Gene: ENSG00000160007 | OMIM ID: 605277 |
Screen Evidence:
| |||
ARHGAP35 involved pathways:
| KEGG pathway | Description | View |
|---|---|---|
| hsa04510 | Focal adhesion | |
| hsa04611 | Platelet activation | |
| hsa04670 | Leukocyte transendothelial migration | |
| hsa04810 | Regulation of actin cytoskeleton |
Expression of ARHGAP35:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | ARHGAP35 | 2909 | 229394_s_at | 0.0890 | 0.8605 | |
| GSE20347 | ARHGAP35 | 2909 | 202044_at | -0.0057 | 0.9489 | |
| GSE23400 | ARHGAP35 | 2909 | 202044_at | -0.0138 | 0.7505 | |
| GSE26886 | ARHGAP35 | 2909 | 229394_s_at | 0.1614 | 0.4303 | |
| GSE29001 | ARHGAP35 | 2909 | 202045_s_at | 0.0464 | 0.8880 | |
| GSE38129 | ARHGAP35 | 2909 | 202044_at | -0.0672 | 0.3032 | |
| GSE45670 | ARHGAP35 | 2909 | 229394_s_at | -0.1206 | 0.3759 | |
| GSE53622 | ARHGAP35 | 2909 | 8140 | 0.0682 | 0.2908 | |
| GSE53624 | ARHGAP35 | 2909 | 8140 | 0.1781 | 0.0019 | |
| GSE63941 | ARHGAP35 | 2909 | 229394_s_at | -0.3413 | 0.4585 | |
| GSE77861 | ARHGAP35 | 2909 | 229394_s_at | 0.0607 | 0.8421 | |
| GSE97050 | ARHGAP35 | 2909 | A_24_P36299 | 0.0539 | 0.9029 | |
| SRP007169 | ARHGAP35 | 2909 | RNAseq | 0.6490 | 0.0313 | |
| SRP008496 | ARHGAP35 | 2909 | RNAseq | 0.5346 | 0.0120 | |
| SRP064894 | ARHGAP35 | 2909 | RNAseq | -0.2110 | 0.1036 | |
| SRP133303 | ARHGAP35 | 2909 | RNAseq | 0.0083 | 0.9593 | |
| SRP159526 | ARHGAP35 | 2909 | RNAseq | 0.3474 | 0.1722 | |
| SRP193095 | ARHGAP35 | 2909 | RNAseq | 0.0957 | 0.2867 | |
| SRP219564 | ARHGAP35 | 2909 | RNAseq | -0.3420 | 0.1900 |
Upregulated datasets: 0; Downregulated datasets: 0.
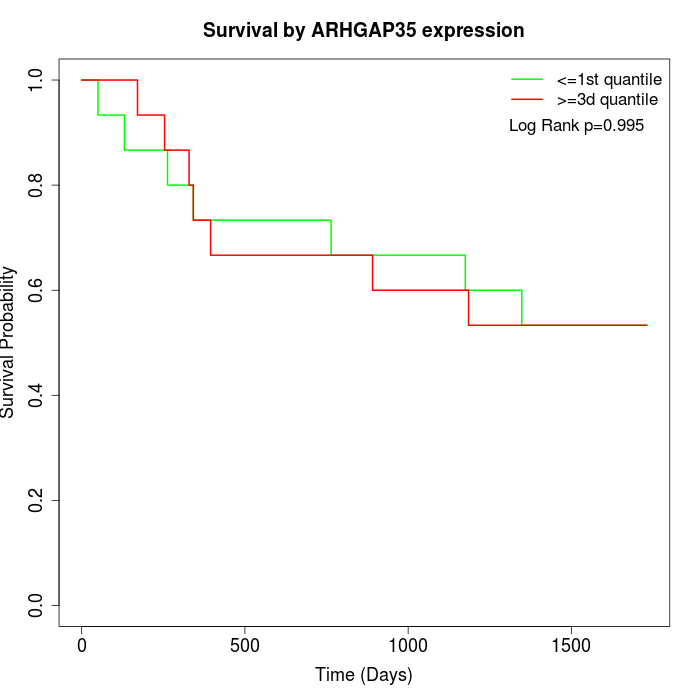
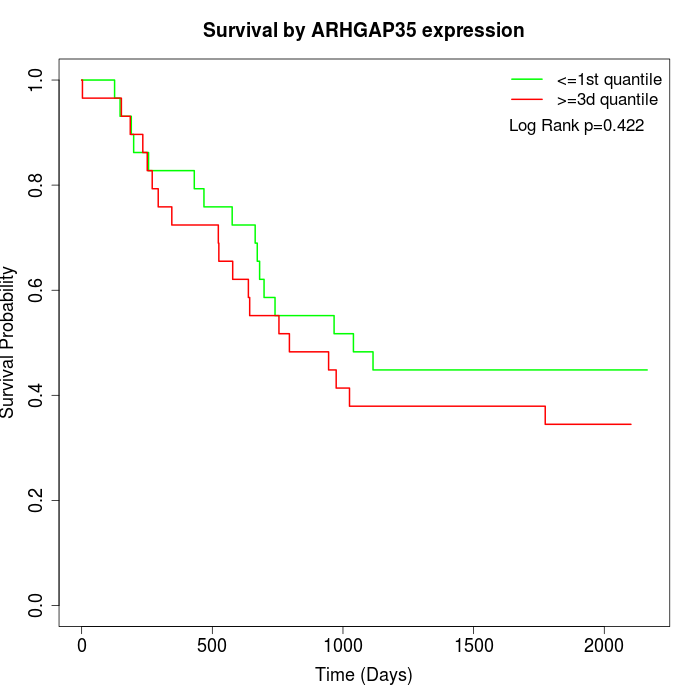
Survival by ARHGAP35 expression:
Note: Click image to view full size file.
Copy number change of ARHGAP35:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | ARHGAP35 | 2909 | 4 | 4 | 22 | |
| GSE20123 | ARHGAP35 | 2909 | 4 | 3 | 23 | |
| GSE43470 | ARHGAP35 | 2909 | 3 | 11 | 29 | |
| GSE46452 | ARHGAP35 | 2909 | 45 | 1 | 13 | |
| GSE47630 | ARHGAP35 | 2909 | 8 | 7 | 25 | |
| GSE54993 | ARHGAP35 | 2909 | 17 | 4 | 49 | |
| GSE54994 | ARHGAP35 | 2909 | 5 | 14 | 34 | |
| GSE60625 | ARHGAP35 | 2909 | 9 | 0 | 2 | |
| GSE74703 | ARHGAP35 | 2909 | 3 | 7 | 26 | |
| GSE74704 | ARHGAP35 | 2909 | 4 | 1 | 15 | |
| TCGA | ARHGAP35 | 2909 | 15 | 17 | 64 |
Total number of gains: 117; Total number of losses: 69; Total Number of normals: 302.
Somatic mutations of ARHGAP35:
Generating mutation plots.
Highly correlated genes for ARHGAP35:
Showing top 20/223 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| ARHGAP35 | GTPBP1 | 0.794762 | 3 | 0 | 3 |
| ARHGAP35 | RNASEH2B | 0.7583 | 3 | 0 | 3 |
| ARHGAP35 | ZNF155 | 0.750105 | 3 | 0 | 3 |
| ARHGAP35 | ERGIC1 | 0.746034 | 3 | 0 | 3 |
| ARHGAP35 | ZNF518B | 0.744533 | 3 | 0 | 3 |
| ARHGAP35 | TRIM8 | 0.738229 | 3 | 0 | 3 |
| ARHGAP35 | SYNE2 | 0.738064 | 3 | 0 | 3 |
| ARHGAP35 | NUDT19 | 0.737686 | 3 | 0 | 3 |
| ARHGAP35 | ZC3HAV1 | 0.730992 | 3 | 0 | 3 |
| ARHGAP35 | ZNF146 | 0.723266 | 3 | 0 | 3 |
| ARHGAP35 | JTB | 0.719766 | 3 | 0 | 3 |
| ARHGAP35 | GPR107 | 0.716428 | 4 | 0 | 4 |
| ARHGAP35 | RNF4 | 0.716239 | 3 | 0 | 3 |
| ARHGAP35 | ZNF121 | 0.715515 | 3 | 0 | 3 |
| ARHGAP35 | CYTH2 | 0.711944 | 4 | 0 | 3 |
| ARHGAP35 | RRAGC | 0.711845 | 3 | 0 | 3 |
| ARHGAP35 | PAIP1 | 0.709285 | 3 | 0 | 3 |
| ARHGAP35 | ARID2 | 0.702572 | 3 | 0 | 3 |
| ARHGAP35 | GTPBP6 | 0.700927 | 3 | 0 | 3 |
| ARHGAP35 | MYH3 | 0.698482 | 3 | 0 | 3 |
For details and further investigation, click here