| Full name: long intergenic non-protein coding RNA 682 | Alias Symbol: | ||
| Type: non-coding RNA | Cytoband: 4p13 | ||
| Entrez ID: 101927074 | HGNC ID: HGNC:44466 | Ensembl Gene: ENSG00000245870 | OMIM ID: |
Expression of LINC00682:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | LINC00682 | 101927074 | 1569418_at | -0.0531 | 0.8409 | |
| GSE26886 | LINC00682 | 101927074 | 1569418_at | 0.2124 | 0.1283 | |
| GSE45670 | LINC00682 | 101927074 | 1569418_at | 0.1902 | 0.6341 | |
| GSE53622 | LINC00682 | 101927074 | 6059 | -0.3119 | 0.0105 | |
| GSE53624 | LINC00682 | 101927074 | 41310 | 0.4555 | 0.0000 | |
| GSE63941 | LINC00682 | 101927074 | 1569418_at | 0.1309 | 0.3186 | |
| GSE77861 | LINC00682 | 101927074 | 1569418_at | -0.0438 | 0.6809 |
Upregulated datasets: 0; Downregulated datasets: 0.
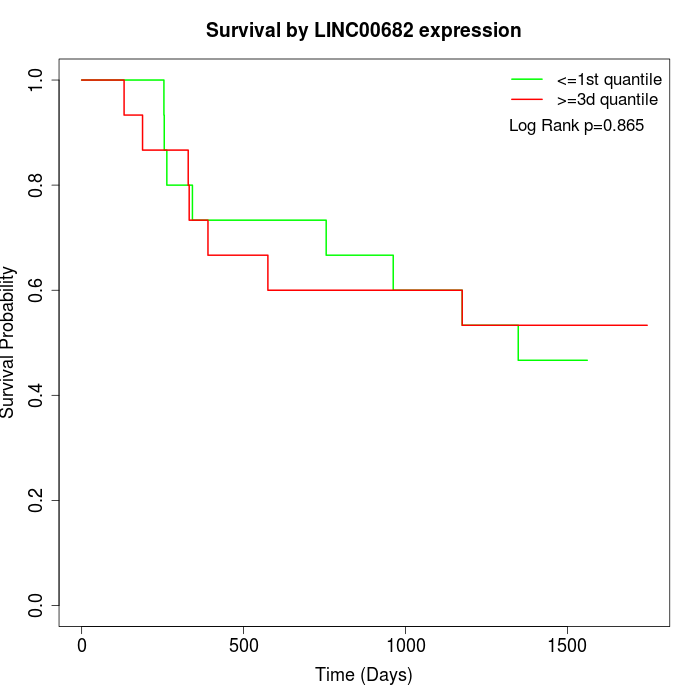
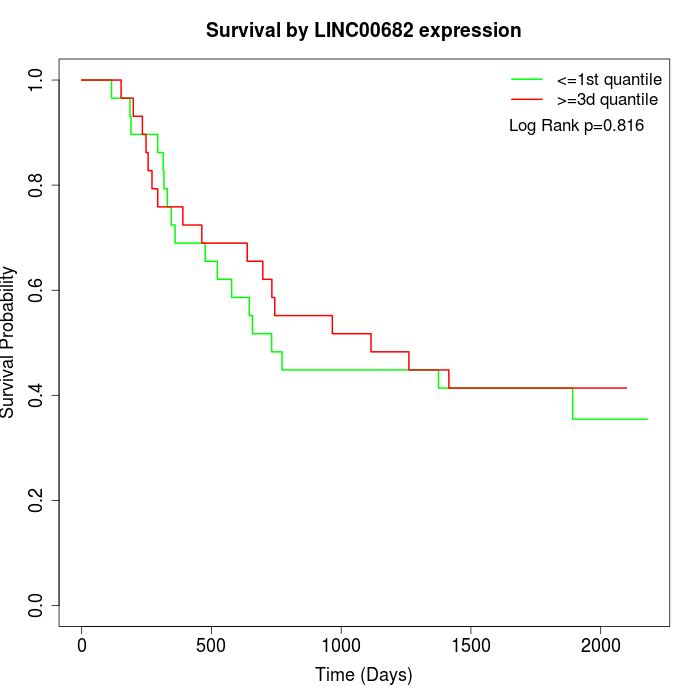
Survival by LINC00682 expression:
Note: Click image to view full size file.
Copy number change of LINC00682:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | LINC00682 | 101927074 | 0 | 16 | 14 | |
| GSE20123 | LINC00682 | 101927074 | 0 | 16 | 14 | |
| GSE43470 | LINC00682 | 101927074 | 0 | 14 | 29 | |
| GSE46452 | LINC00682 | 101927074 | 1 | 36 | 22 | |
| GSE47630 | LINC00682 | 101927074 | 1 | 19 | 20 | |
| GSE54993 | LINC00682 | 101927074 | 7 | 0 | 63 | |
| GSE54994 | LINC00682 | 101927074 | 4 | 9 | 40 | |
| GSE60625 | LINC00682 | 101927074 | 0 | 0 | 11 | |
| GSE74703 | LINC00682 | 101927074 | 0 | 12 | 24 | |
| GSE74704 | LINC00682 | 101927074 | 0 | 9 | 11 | |
| TCGA | LINC00682 | 101927074 | 11 | 42 | 43 |
Total number of gains: 24; Total number of losses: 173; Total Number of normals: 291.
Somatic mutations of LINC00682:
Generating mutation plots.
Highly correlated genes for LINC00682:
Showing top 20/27 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| LINC00682 | ATXN2L | 0.640019 | 3 | 0 | 3 |
| LINC00682 | EYS | 0.636693 | 3 | 0 | 3 |
| LINC00682 | PDZK1 | 0.622782 | 3 | 0 | 3 |
| LINC00682 | NFE4 | 0.61374 | 4 | 0 | 3 |
| LINC00682 | KLK15 | 0.609947 | 3 | 0 | 3 |
| LINC00682 | IL34 | 0.601025 | 4 | 0 | 3 |
| LINC00682 | NEUROD4 | 0.586264 | 3 | 0 | 3 |
| LINC00682 | ASMTL-AS1 | 0.585317 | 3 | 0 | 3 |
| LINC00682 | VPS9D1-AS1 | 0.581475 | 4 | 0 | 3 |
| LINC00682 | RILP | 0.579979 | 4 | 0 | 3 |
| LINC00682 | VAMP2 | 0.577006 | 4 | 0 | 3 |
| LINC00682 | PCAT4 | 0.566225 | 3 | 0 | 3 |
| LINC00682 | SLC25A2 | 0.561655 | 3 | 0 | 3 |
| LINC00682 | CBX4 | 0.560653 | 4 | 0 | 3 |
| LINC00682 | PRORY | 0.56016 | 4 | 0 | 3 |
| LINC00682 | TBKBP1 | 0.559077 | 4 | 0 | 3 |
| LINC00682 | RBP3 | 0.558248 | 4 | 0 | 3 |
| LINC00682 | GSK3A | 0.558189 | 5 | 0 | 3 |
| LINC00682 | EMD | 0.557078 | 3 | 0 | 3 |
| LINC00682 | SRL | 0.554872 | 3 | 0 | 3 |
For details and further investigation, click here