| Full name: tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein theta | Alias Symbol: HS1|14-3-3 | ||
| Type: protein-coding gene | Cytoband: 2p25.1 | ||
| Entrez ID: 10971 | HGNC ID: HGNC:12854 | Ensembl Gene: ENSG00000134308 | OMIM ID: 609009 |
YWHAQ involved pathways:
| KEGG pathway | Description | View |
|---|---|---|
| hsa04110 | Cell cycle | |
| hsa04114 | Oocyte meiosis | |
| hsa04151 | PI3K-Akt signaling pathway | |
| hsa04390 | Hippo signaling pathway |
Expression of YWHAQ:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | YWHAQ | 10971 | 213699_s_at | 0.1581 | 0.6135 | |
| GSE20347 | YWHAQ | 10971 | 213699_s_at | 0.2472 | 0.0341 | |
| GSE23400 | YWHAQ | 10971 | 213699_s_at | 0.4263 | 0.0000 | |
| GSE26886 | YWHAQ | 10971 | 212426_s_at | -0.1085 | 0.4249 | |
| GSE29001 | YWHAQ | 10971 | 213699_s_at | 0.2629 | 0.0835 | |
| GSE38129 | YWHAQ | 10971 | 213699_s_at | 0.3358 | 0.0001 | |
| GSE45670 | YWHAQ | 10971 | 213699_s_at | 0.3275 | 0.0038 | |
| GSE53622 | YWHAQ | 10971 | 34278 | 0.5820 | 0.0000 | |
| GSE53624 | YWHAQ | 10971 | 34278 | 0.4804 | 0.0000 | |
| GSE63941 | YWHAQ | 10971 | 213699_s_at | -0.4284 | 0.0655 | |
| GSE77861 | YWHAQ | 10971 | 213699_s_at | 0.2838 | 0.0859 | |
| GSE97050 | YWHAQ | 10971 | A_23_P131723 | 0.5069 | 0.1344 | |
| SRP007169 | YWHAQ | 10971 | RNAseq | -0.5774 | 0.1012 | |
| SRP008496 | YWHAQ | 10971 | RNAseq | -0.3756 | 0.1842 | |
| SRP064894 | YWHAQ | 10971 | RNAseq | 0.1176 | 0.4685 | |
| SRP133303 | YWHAQ | 10971 | RNAseq | 0.6560 | 0.0003 | |
| SRP159526 | YWHAQ | 10971 | RNAseq | 0.5254 | 0.0288 | |
| SRP193095 | YWHAQ | 10971 | RNAseq | 0.3251 | 0.0649 | |
| SRP219564 | YWHAQ | 10971 | RNAseq | 0.1540 | 0.6480 | |
| TCGA | YWHAQ | 10971 | RNAseq | 0.1356 | 0.0005 |
Upregulated datasets: 0; Downregulated datasets: 0.
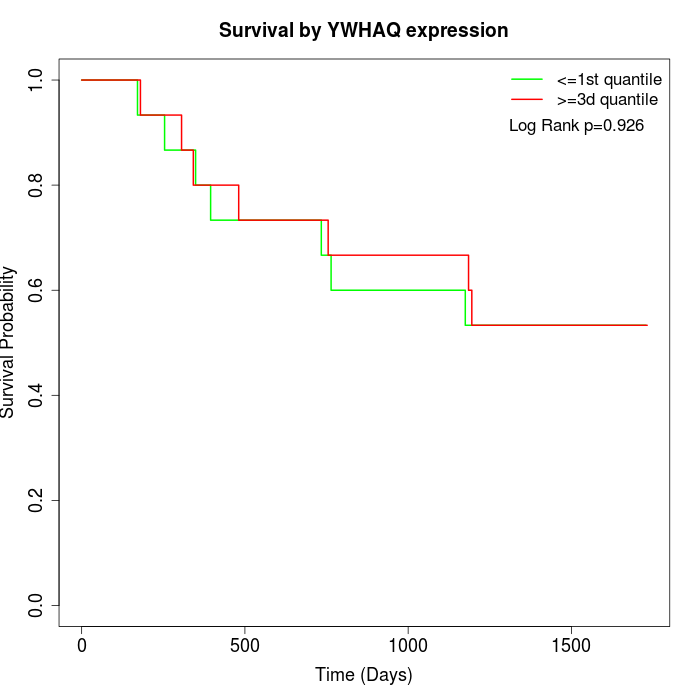
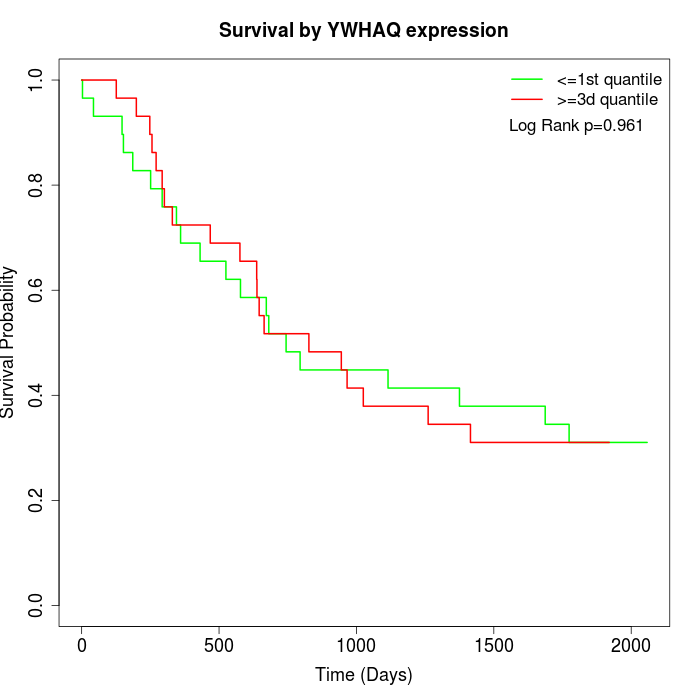
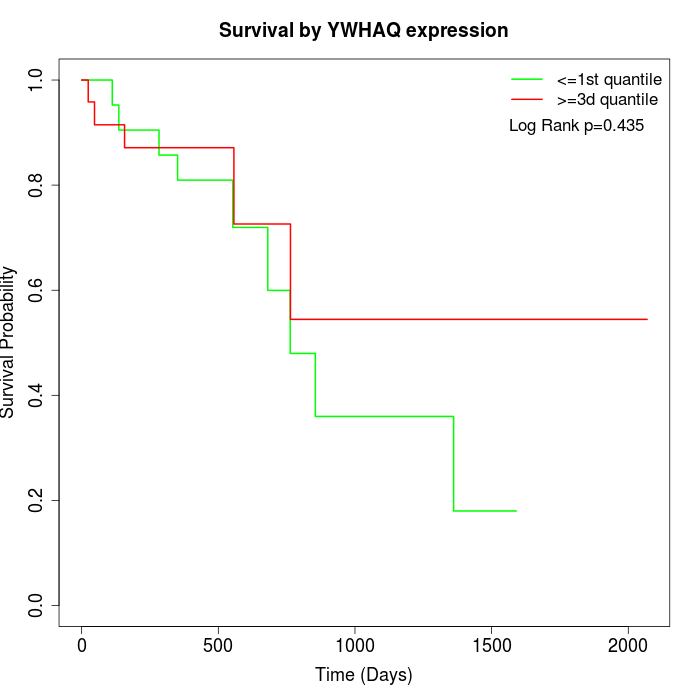
Survival by YWHAQ expression:
Note: Click image to view full size file.
Copy number change of YWHAQ:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | YWHAQ | 10971 | 11 | 2 | 17 | |
| GSE20123 | YWHAQ | 10971 | 11 | 2 | 17 | |
| GSE43470 | YWHAQ | 10971 | 2 | 1 | 40 | |
| GSE46452 | YWHAQ | 10971 | 3 | 4 | 52 | |
| GSE47630 | YWHAQ | 10971 | 7 | 0 | 33 | |
| GSE54993 | YWHAQ | 10971 | 0 | 7 | 63 | |
| GSE54994 | YWHAQ | 10971 | 11 | 0 | 42 | |
| GSE60625 | YWHAQ | 10971 | 0 | 3 | 8 | |
| GSE74703 | YWHAQ | 10971 | 2 | 0 | 34 | |
| GSE74704 | YWHAQ | 10971 | 9 | 1 | 10 | |
| TCGA | YWHAQ | 10971 | 34 | 5 | 57 |
Total number of gains: 90; Total number of losses: 25; Total Number of normals: 373.
Somatic mutations of YWHAQ:
Generating mutation plots.
Highly correlated genes for YWHAQ:
Showing top 20/573 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| YWHAQ | ALPK2 | 0.693328 | 3 | 0 | 3 |
| YWHAQ | TUBA4A | 0.673311 | 3 | 0 | 3 |
| YWHAQ | LINC00621 | 0.666576 | 3 | 0 | 3 |
| YWHAQ | GLIS2 | 0.666433 | 3 | 0 | 3 |
| YWHAQ | ADAM17 | 0.662669 | 10 | 0 | 9 |
| YWHAQ | SPRY4-IT1 | 0.661853 | 3 | 0 | 3 |
| YWHAQ | SH2D5 | 0.65302 | 5 | 0 | 5 |
| YWHAQ | SPRR2G | 0.64997 | 4 | 0 | 3 |
| YWHAQ | C16orf74 | 0.647041 | 4 | 0 | 4 |
| YWHAQ | KRTDAP | 0.646627 | 4 | 0 | 3 |
| YWHAQ | SH3RF3 | 0.641898 | 3 | 0 | 3 |
| YWHAQ | MSANTD3 | 0.629716 | 6 | 0 | 5 |
| YWHAQ | FBLIM1 | 0.628981 | 5 | 0 | 4 |
| YWHAQ | SH3KBP1 | 0.627241 | 3 | 0 | 3 |
| YWHAQ | PRIM2 | 0.625735 | 7 | 0 | 5 |
| YWHAQ | ADAMTS12 | 0.623771 | 6 | 0 | 5 |
| YWHAQ | SNRPG | 0.620803 | 10 | 0 | 9 |
| YWHAQ | TUBG1 | 0.620445 | 9 | 0 | 8 |
| YWHAQ | CKAP2L | 0.620148 | 6 | 0 | 4 |
| YWHAQ | PSMA4 | 0.619549 | 10 | 0 | 9 |
For details and further investigation, click here