| Full name: ATP6V0E2 antisense RNA 1 | Alias Symbol: | ||
| Type: non-coding RNA | Cytoband: 7q36.1 | ||
| Entrez ID: 401431 | HGNC ID: HGNC:44180 | Ensembl Gene: ENSG00000204934 | OMIM ID: |
Expression of ATP6V0E2-AS1:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | ATP6V0E2-AS1 | 401431 | 229094_at | -0.1995 | 0.5256 | |
| GSE26886 | ATP6V0E2-AS1 | 401431 | 229094_at | 0.3711 | 0.0013 | |
| GSE45670 | ATP6V0E2-AS1 | 401431 | 229094_at | -0.1874 | 0.1370 | |
| GSE53622 | ATP6V0E2-AS1 | 401431 | 2838 | -0.2184 | 0.1727 | |
| GSE53624 | ATP6V0E2-AS1 | 401431 | 2838 | -0.2871 | 0.0245 | |
| GSE63941 | ATP6V0E2-AS1 | 401431 | 229094_at | -0.2144 | 0.5458 | |
| GSE77861 | ATP6V0E2-AS1 | 401431 | 229094_at | -0.0400 | 0.7725 | |
| SRP133303 | ATP6V0E2-AS1 | 401431 | RNAseq | -0.6718 | 0.0420 | |
| SRP159526 | ATP6V0E2-AS1 | 401431 | RNAseq | 0.9678 | 0.2149 | |
| SRP219564 | ATP6V0E2-AS1 | 401431 | RNAseq | -0.2233 | 0.7418 |
Upregulated datasets: 0; Downregulated datasets: 0.
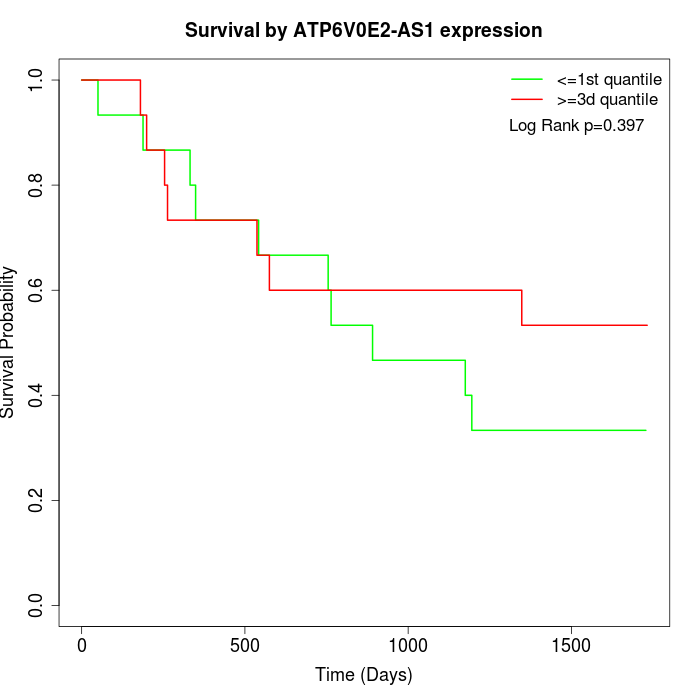
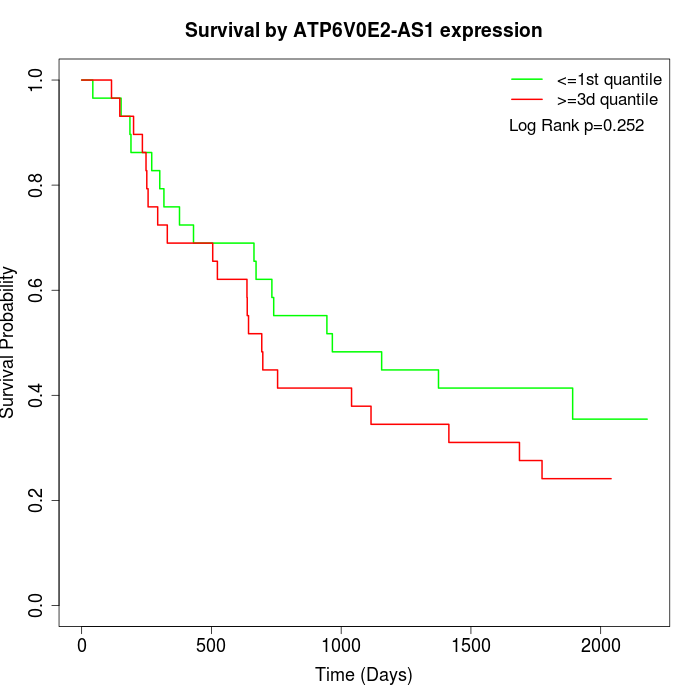
Survival by ATP6V0E2-AS1 expression:
Note: Click image to view full size file.
Copy number change of ATP6V0E2-AS1:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | ATP6V0E2-AS1 | 401431 | 2 | 3 | 25 | |
| GSE20123 | ATP6V0E2-AS1 | 401431 | 2 | 3 | 25 | |
| GSE43470 | ATP6V0E2-AS1 | 401431 | 2 | 5 | 36 | |
| GSE46452 | ATP6V0E2-AS1 | 401431 | 7 | 2 | 50 | |
| GSE47630 | ATP6V0E2-AS1 | 401431 | 6 | 8 | 26 | |
| GSE54993 | ATP6V0E2-AS1 | 401431 | 3 | 3 | 64 | |
| GSE54994 | ATP6V0E2-AS1 | 401431 | 6 | 8 | 39 | |
| GSE60625 | ATP6V0E2-AS1 | 401431 | 0 | 0 | 11 | |
| GSE74703 | ATP6V0E2-AS1 | 401431 | 2 | 4 | 30 | |
| GSE74704 | ATP6V0E2-AS1 | 401431 | 1 | 3 | 16 |
Total number of gains: 31; Total number of losses: 39; Total Number of normals: 322.
Somatic mutations of ATP6V0E2-AS1:
Generating mutation plots.
Highly correlated genes for ATP6V0E2-AS1:
Showing top 20/22 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| ATP6V0E2-AS1 | CMC4 | 0.662106 | 3 | 0 | 3 |
| ATP6V0E2-AS1 | FGF18 | 0.612948 | 3 | 0 | 3 |
| ATP6V0E2-AS1 | STARD13 | 0.610698 | 3 | 0 | 3 |
| ATP6V0E2-AS1 | FXYD6 | 0.605916 | 3 | 0 | 3 |
| ATP6V0E2-AS1 | PSIP1 | 0.605218 | 4 | 0 | 3 |
| ATP6V0E2-AS1 | LMO1 | 0.600656 | 3 | 0 | 3 |
| ATP6V0E2-AS1 | GSC | 0.600359 | 3 | 0 | 3 |
| ATP6V0E2-AS1 | YPEL1 | 0.596525 | 3 | 0 | 3 |
| ATP6V0E2-AS1 | LYRM9 | 0.590968 | 5 | 0 | 3 |
| ATP6V0E2-AS1 | MAGI2 | 0.575114 | 5 | 0 | 5 |
| ATP6V0E2-AS1 | TUB | 0.570249 | 4 | 0 | 3 |
| ATP6V0E2-AS1 | PCOLCE2 | 0.567852 | 3 | 0 | 3 |
| ATP6V0E2-AS1 | NT5M | 0.567452 | 4 | 0 | 3 |
| ATP6V0E2-AS1 | NUDT2 | 0.563182 | 4 | 0 | 3 |
| ATP6V0E2-AS1 | CACNA2D2 | 0.55709 | 4 | 0 | 4 |
| ATP6V0E2-AS1 | PLCB4 | 0.556565 | 4 | 0 | 3 |
| ATP6V0E2-AS1 | CTSF | 0.540944 | 5 | 0 | 3 |
| ATP6V0E2-AS1 | PRKAR2B | 0.533787 | 4 | 0 | 3 |
| ATP6V0E2-AS1 | RNF150 | 0.532104 | 6 | 0 | 3 |
| ATP6V0E2-AS1 | GNAZ | 0.530856 | 5 | 0 | 3 |
For details and further investigation, click here