| Full name: protein kinase cAMP-dependent type II regulatory subunit beta | Alias Symbol: | ||
| Type: protein-coding gene | Cytoband: 7q22.3 | ||
| Entrez ID: 5577 | HGNC ID: HGNC:9392 | Ensembl Gene: ENSG00000005249 | OMIM ID: 176912 |
| Related drugs: BX-795, CHELERYTHRINE, CYCLIC ADENOSINE MONOPHOSPHATE, METHYL NONYL KETONE, THEOPHYLLINE... [more] | |||
PRKAR2B involved pathways:
| KEGG pathway | Description | View |
|---|---|---|
| hsa04910 | Insulin signaling pathway |
Expression of PRKAR2B:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | PRKAR2B | 5577 | 203680_at | -2.0141 | 0.2162 | |
| GSE20347 | PRKAR2B | 5577 | 203680_at | -0.3759 | 0.5065 | |
| GSE23400 | PRKAR2B | 5577 | 203680_at | -0.9610 | 0.0001 | |
| GSE26886 | PRKAR2B | 5577 | 203680_at | 0.5716 | 0.4471 | |
| GSE29001 | PRKAR2B | 5577 | 203680_at | -0.3909 | 0.5797 | |
| GSE38129 | PRKAR2B | 5577 | 203680_at | -1.3769 | 0.0142 | |
| GSE45670 | PRKAR2B | 5577 | 203680_at | -2.9533 | 0.0000 | |
| GSE53622 | PRKAR2B | 5577 | 60361 | -1.2784 | 0.0000 | |
| GSE53624 | PRKAR2B | 5577 | 60361 | -0.8879 | 0.0000 | |
| GSE63941 | PRKAR2B | 5577 | 203680_at | -4.8853 | 0.0035 | |
| GSE77861 | PRKAR2B | 5577 | 203680_at | 0.5774 | 0.1588 | |
| GSE97050 | PRKAR2B | 5577 | A_33_P3304983 | -0.9054 | 0.1692 | |
| SRP007169 | PRKAR2B | 5577 | RNAseq | 0.2225 | 0.7168 | |
| SRP064894 | PRKAR2B | 5577 | RNAseq | -1.1641 | 0.0020 | |
| SRP133303 | PRKAR2B | 5577 | RNAseq | -1.1206 | 0.0022 | |
| SRP159526 | PRKAR2B | 5577 | RNAseq | -0.5804 | 0.0880 | |
| SRP193095 | PRKAR2B | 5577 | RNAseq | 0.2296 | 0.4811 | |
| SRP219564 | PRKAR2B | 5577 | RNAseq | -0.4636 | 0.6376 | |
| TCGA | PRKAR2B | 5577 | RNAseq | -0.7352 | 0.0002 |
Upregulated datasets: 0; Downregulated datasets: 6.
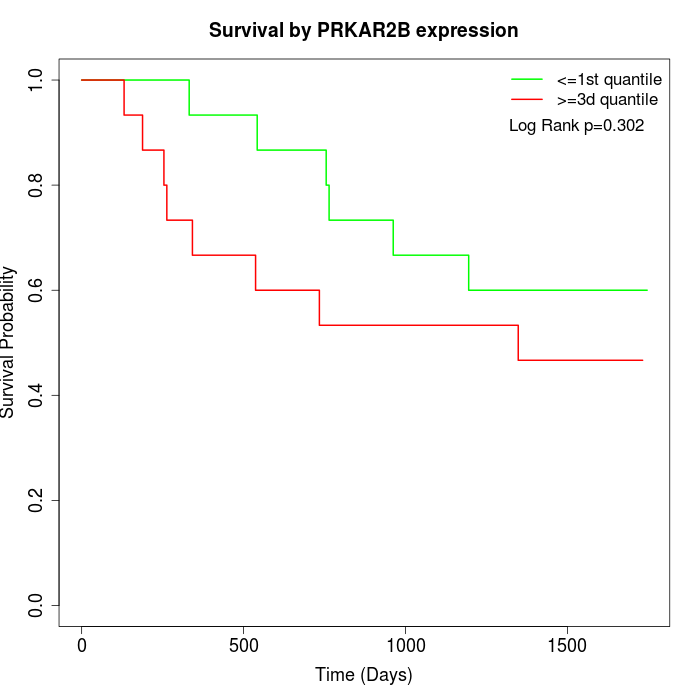
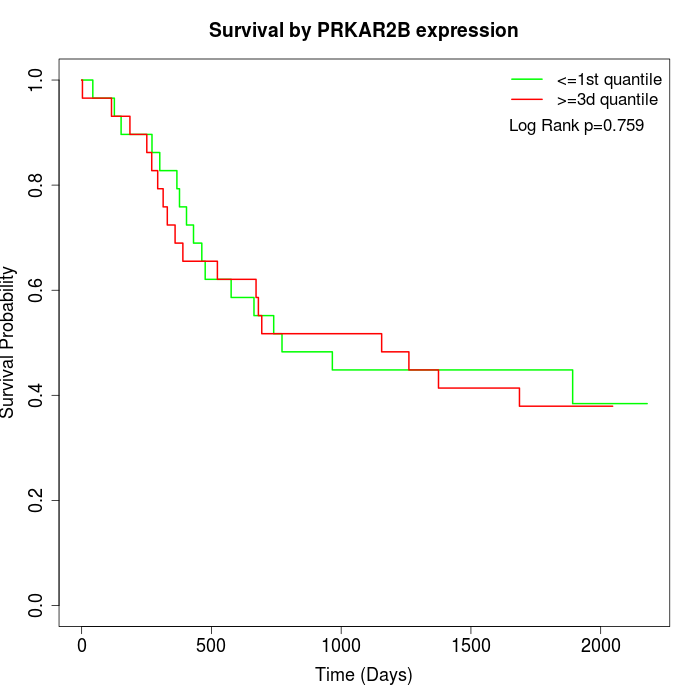
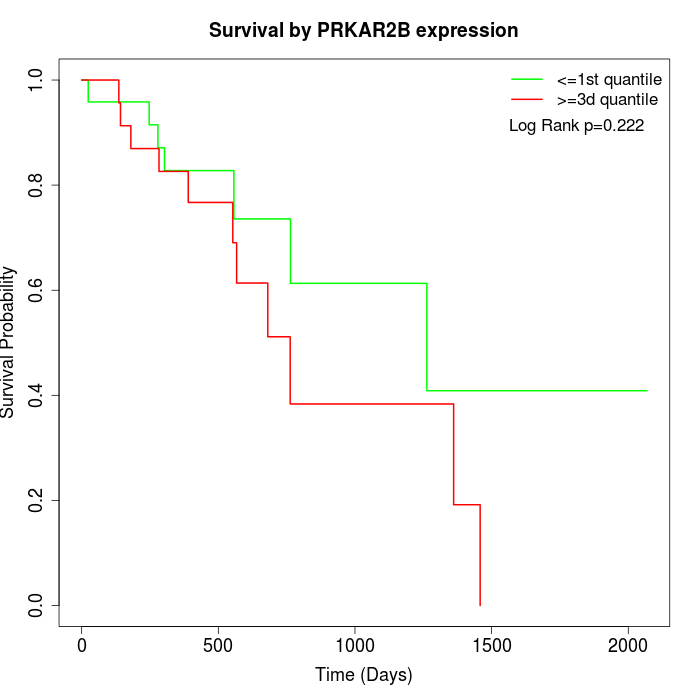
Survival by PRKAR2B expression:
Note: Click image to view full size file.
Copy number change of PRKAR2B:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | PRKAR2B | 5577 | 12 | 0 | 18 | |
| GSE20123 | PRKAR2B | 5577 | 12 | 0 | 18 | |
| GSE43470 | PRKAR2B | 5577 | 6 | 2 | 35 | |
| GSE46452 | PRKAR2B | 5577 | 10 | 1 | 48 | |
| GSE47630 | PRKAR2B | 5577 | 8 | 3 | 29 | |
| GSE54993 | PRKAR2B | 5577 | 3 | 8 | 59 | |
| GSE54994 | PRKAR2B | 5577 | 16 | 3 | 34 | |
| GSE60625 | PRKAR2B | 5577 | 0 | 0 | 11 | |
| GSE74703 | PRKAR2B | 5577 | 6 | 2 | 28 | |
| GSE74704 | PRKAR2B | 5577 | 8 | 0 | 12 | |
| TCGA | PRKAR2B | 5577 | 50 | 10 | 36 |
Total number of gains: 131; Total number of losses: 29; Total Number of normals: 328.
Somatic mutations of PRKAR2B:
Generating mutation plots.
Highly correlated genes for PRKAR2B:
Showing top 20/1228 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| PRKAR2B | AFF3 | 0.839461 | 3 | 0 | 3 |
| PRKAR2B | UBXN10 | 0.828904 | 3 | 0 | 3 |
| PRKAR2B | SYNPO2 | 0.827697 | 8 | 0 | 8 |
| PRKAR2B | ITGA7 | 0.813292 | 9 | 0 | 9 |
| PRKAR2B | PDE5A | 0.81326 | 9 | 0 | 9 |
| PRKAR2B | BHMT2 | 0.8126 | 9 | 0 | 9 |
| PRKAR2B | C3orf70 | 0.811813 | 7 | 0 | 7 |
| PRKAR2B | NEGR1 | 0.8091 | 6 | 0 | 6 |
| PRKAR2B | CBX7 | 0.805823 | 9 | 0 | 8 |
| PRKAR2B | FXYD1 | 0.802011 | 6 | 0 | 6 |
| PRKAR2B | RNF150 | 0.801166 | 7 | 0 | 7 |
| PRKAR2B | SORCS1 | 0.798877 | 4 | 0 | 4 |
| PRKAR2B | LONRF2 | 0.798013 | 5 | 0 | 5 |
| PRKAR2B | MAMDC2 | 0.797115 | 6 | 0 | 6 |
| PRKAR2B | FIBIN | 0.793107 | 4 | 0 | 4 |
| PRKAR2B | FENDRR | 0.792475 | 5 | 0 | 5 |
| PRKAR2B | TCEAL2 | 0.791385 | 10 | 0 | 9 |
| PRKAR2B | FILIP1 | 0.791022 | 5 | 0 | 5 |
| PRKAR2B | LMOD1 | 0.783193 | 11 | 0 | 11 |
| PRKAR2B | LINC01279 | 0.780894 | 4 | 0 | 4 |
For details and further investigation, click here