| Full name: dihydrolipoamide S-acetyltransferase | Alias Symbol: PDC-E2|E2 | ||
| Type: protein-coding gene | Cytoband: 11q23.1 | ||
| Entrez ID: 1737 | HGNC ID: HGNC:2896 | Ensembl Gene: ENSG00000150768 | OMIM ID: 608770 |
Screen Evidence:
| |||
Expression of DLAT:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | DLAT | 1737 | 212568_s_at | -0.3699 | 0.3463 | |
| GSE20347 | DLAT | 1737 | 212568_s_at | -0.6444 | 0.0004 | |
| GSE23400 | DLAT | 1737 | 212568_s_at | -0.0715 | 0.3786 | |
| GSE26886 | DLAT | 1737 | 213149_at | -0.7614 | 0.0124 | |
| GSE29001 | DLAT | 1737 | 212568_s_at | -0.3894 | 0.2134 | |
| GSE38129 | DLAT | 1737 | 212568_s_at | -0.4012 | 0.0089 | |
| GSE45670 | DLAT | 1737 | 212568_s_at | -0.1786 | 0.2631 | |
| GSE53622 | DLAT | 1737 | 122527 | -0.2866 | 0.0000 | |
| GSE53624 | DLAT | 1737 | 122527 | -0.0737 | 0.3007 | |
| GSE63941 | DLAT | 1737 | 212568_s_at | -0.0403 | 0.9310 | |
| GSE77861 | DLAT | 1737 | 212568_s_at | -0.3088 | 0.5071 | |
| GSE97050 | DLAT | 1737 | A_23_P203030 | -0.2504 | 0.3249 | |
| SRP007169 | DLAT | 1737 | RNAseq | -1.1430 | 0.0008 | |
| SRP008496 | DLAT | 1737 | RNAseq | -0.9473 | 0.0000 | |
| SRP064894 | DLAT | 1737 | RNAseq | -0.7539 | 0.0000 | |
| SRP133303 | DLAT | 1737 | RNAseq | -0.2786 | 0.0542 | |
| SRP159526 | DLAT | 1737 | RNAseq | -0.5821 | 0.1307 | |
| SRP193095 | DLAT | 1737 | RNAseq | -0.7194 | 0.0000 | |
| SRP219564 | DLAT | 1737 | RNAseq | -0.7562 | 0.0020 | |
| TCGA | DLAT | 1737 | RNAseq | -0.0975 | 0.0462 |
Upregulated datasets: 0; Downregulated datasets: 1.
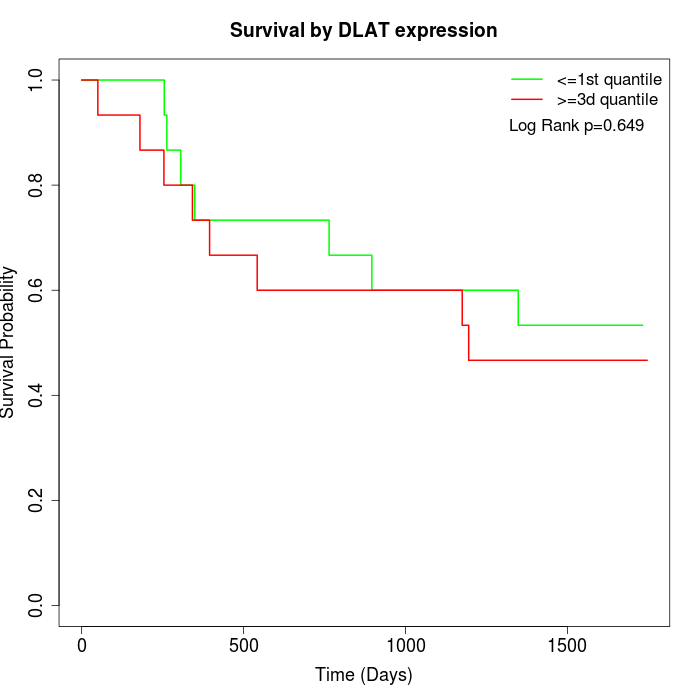
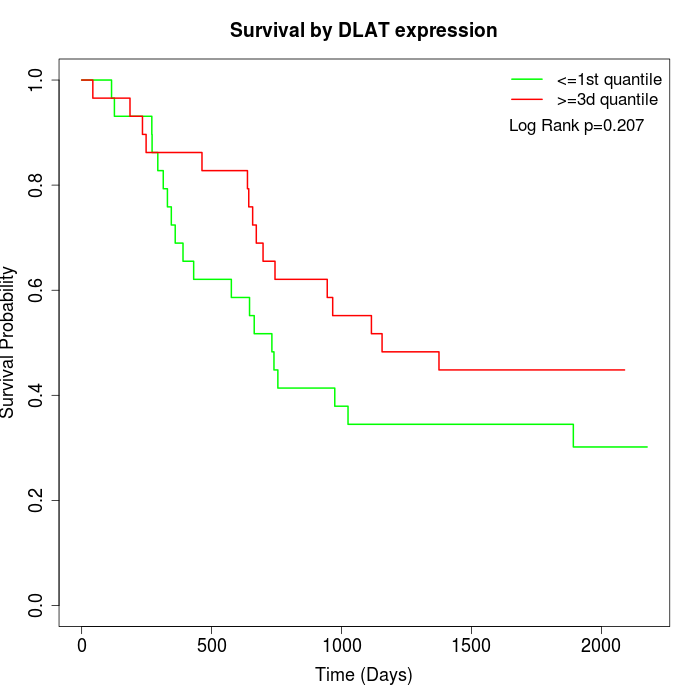
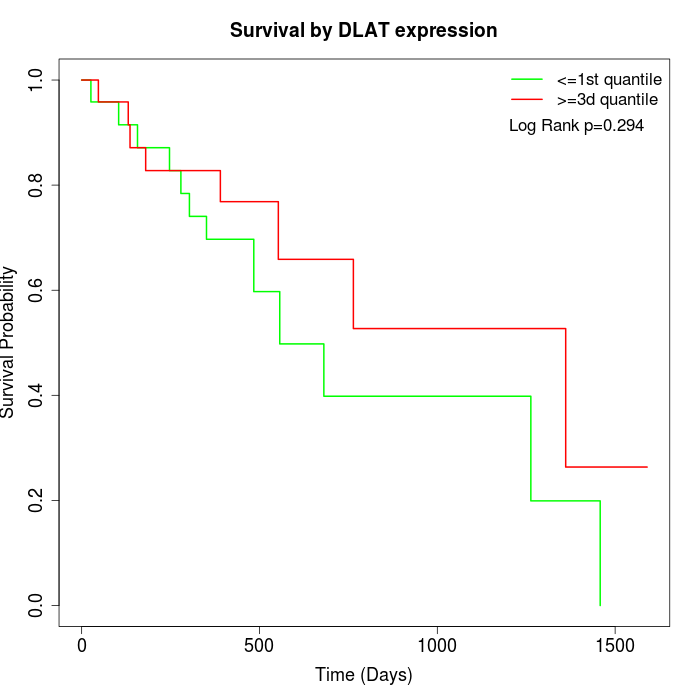
Survival by DLAT expression:
Note: Click image to view full size file.
Copy number change of DLAT:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | DLAT | 1737 | 0 | 12 | 18 | |
| GSE20123 | DLAT | 1737 | 0 | 12 | 18 | |
| GSE43470 | DLAT | 1737 | 2 | 6 | 35 | |
| GSE46452 | DLAT | 1737 | 3 | 26 | 30 | |
| GSE47630 | DLAT | 1737 | 2 | 19 | 19 | |
| GSE54993 | DLAT | 1737 | 10 | 0 | 60 | |
| GSE54994 | DLAT | 1737 | 5 | 19 | 29 | |
| GSE60625 | DLAT | 1737 | 0 | 3 | 8 | |
| GSE74703 | DLAT | 1737 | 1 | 4 | 31 | |
| GSE74704 | DLAT | 1737 | 0 | 8 | 12 | |
| TCGA | DLAT | 1737 | 7 | 48 | 41 |
Total number of gains: 30; Total number of losses: 157; Total Number of normals: 301.
Somatic mutations of DLAT:
Generating mutation plots.
Highly correlated genes for DLAT:
Showing top 20/470 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| DLAT | ACER3 | 0.709467 | 3 | 0 | 3 |
| DLAT | KATNBL1 | 0.706601 | 3 | 0 | 3 |
| DLAT | IMPACT | 0.705125 | 3 | 0 | 3 |
| DLAT | RPS17 | 0.695994 | 3 | 0 | 3 |
| DLAT | FAM174A | 0.688729 | 3 | 0 | 3 |
| DLAT | SENP8 | 0.683496 | 3 | 0 | 3 |
| DLAT | SDHD | 0.674695 | 13 | 0 | 12 |
| DLAT | FDX1 | 0.669047 | 13 | 0 | 11 |
| DLAT | ORMDL2 | 0.661886 | 3 | 0 | 3 |
| DLAT | C12orf74 | 0.659028 | 3 | 0 | 3 |
| DLAT | CMC1 | 0.650015 | 3 | 0 | 3 |
| DLAT | GSKIP | 0.649101 | 4 | 0 | 3 |
| DLAT | SPIN4 | 0.647531 | 4 | 0 | 3 |
| DLAT | TRMT1L | 0.635379 | 4 | 0 | 4 |
| DLAT | IREB2 | 0.635144 | 5 | 0 | 5 |
| DLAT | ZW10 | 0.632256 | 12 | 0 | 11 |
| DLAT | PTS | 0.631664 | 12 | 0 | 11 |
| DLAT | HOOK3 | 0.627524 | 4 | 0 | 3 |
| DLAT | SLC16A9 | 0.624526 | 4 | 0 | 3 |
| DLAT | FEM1B | 0.624431 | 4 | 0 | 3 |
For details and further investigation, click here