| Full name: long intergenic non-protein coding RNA 1208 | Alias Symbol: | ||
| Type: non-coding RNA | Cytoband: 3q26.32 | ||
| Entrez ID: 100505547 | HGNC ID: HGNC:49639 | Ensembl Gene: ENSG00000223715 | OMIM ID: |
Expression of LINC01208:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | LINC01208 | 100505547 | 237274_at | 0.0752 | 0.8150 | |
| GSE26886 | LINC01208 | 100505547 | 237274_at | 0.0808 | 0.6249 | |
| GSE45670 | LINC01208 | 100505547 | 237274_at | 0.1072 | 0.1975 | |
| GSE53622 | LINC01208 | 100505547 | 80112 | 0.2115 | 0.2405 | |
| GSE53624 | LINC01208 | 100505547 | 80112 | 0.3251 | 0.0269 | |
| GSE63941 | LINC01208 | 100505547 | 237274_at | 0.5917 | 0.0004 | |
| GSE77861 | LINC01208 | 100505547 | 237274_at | 0.1293 | 0.4708 |
Upregulated datasets: 0; Downregulated datasets: 0.
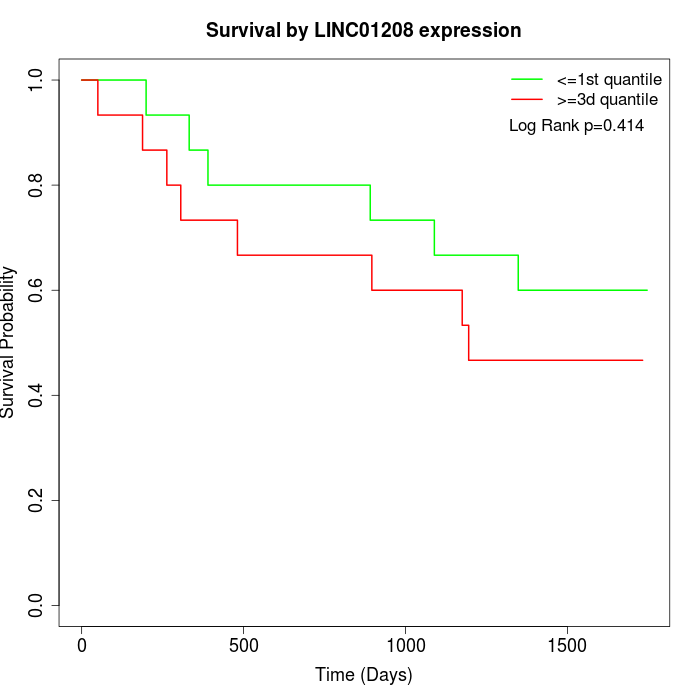
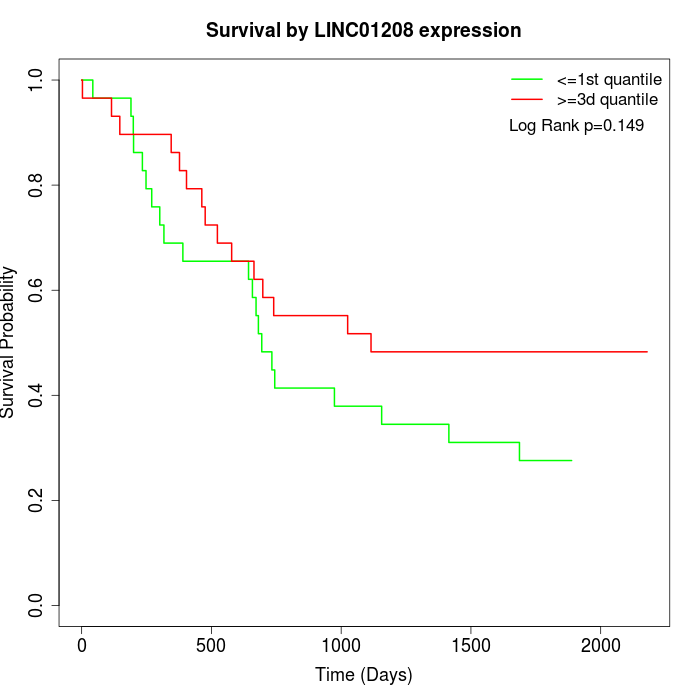
Survival by LINC01208 expression:
Note: Click image to view full size file.
Copy number change of LINC01208:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | LINC01208 | 100505547 | 24 | 0 | 6 | |
| GSE20123 | LINC01208 | 100505547 | 23 | 0 | 7 | |
| GSE43470 | LINC01208 | 100505547 | 24 | 0 | 19 | |
| GSE46452 | LINC01208 | 100505547 | 19 | 1 | 39 | |
| GSE47630 | LINC01208 | 100505547 | 27 | 2 | 11 | |
| GSE54993 | LINC01208 | 100505547 | 1 | 18 | 51 | |
| GSE54994 | LINC01208 | 100505547 | 40 | 1 | 12 | |
| GSE60625 | LINC01208 | 100505547 | 0 | 6 | 5 | |
| GSE74703 | LINC01208 | 100505547 | 20 | 0 | 16 | |
| GSE74704 | LINC01208 | 100505547 | 15 | 0 | 5 |
Total number of gains: 193; Total number of losses: 28; Total Number of normals: 171.
Somatic mutations of LINC01208:
Generating mutation plots.
Highly correlated genes for LINC01208:
Showing top 20/24 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| LINC01208 | HOXD8 | 0.697053 | 3 | 0 | 3 |
| LINC01208 | C17orf67 | 0.694431 | 3 | 0 | 3 |
| LINC01208 | GATA6-AS1 | 0.666563 | 3 | 0 | 3 |
| LINC01208 | DMWD | 0.651716 | 4 | 0 | 3 |
| LINC01208 | CNGB3 | 0.64877 | 3 | 0 | 3 |
| LINC01208 | EPN1 | 0.620072 | 3 | 0 | 3 |
| LINC01208 | GNAO1 | 0.614984 | 4 | 0 | 4 |
| LINC01208 | DMTN | 0.614476 | 3 | 0 | 3 |
| LINC01208 | IL17C | 0.614349 | 3 | 0 | 3 |
| LINC01208 | OR1A1 | 0.61329 | 3 | 0 | 3 |
| LINC01208 | SEMA4G | 0.600951 | 4 | 0 | 4 |
| LINC01208 | LINC00642 | 0.588235 | 4 | 0 | 3 |
| LINC01208 | MPV17L | 0.588123 | 3 | 0 | 3 |
| LINC01208 | BEST1 | 0.568523 | 4 | 0 | 3 |
| LINC01208 | RIMS2 | 0.556904 | 4 | 0 | 3 |
| LINC01208 | ITGAX | 0.55499 | 3 | 0 | 3 |
| LINC01208 | TPSG1 | 0.542367 | 3 | 0 | 3 |
| LINC01208 | BCL9L | 0.53737 | 4 | 0 | 3 |
| LINC01208 | MEGF10 | 0.520969 | 4 | 0 | 3 |
| LINC01208 | FAM178B | 0.519805 | 6 | 0 | 3 |
For details and further investigation, click here