| Full name: leucine rich repeat and Ig domain containing 3 | Alias Symbol: LERN2 | ||
| Type: protein-coding gene | Cytoband: 19p13.3 | ||
| Entrez ID: 645191 | HGNC ID: HGNC:21206 | Ensembl Gene: ENSG00000220008 | OMIM ID: 609792 |
Expression of LINGO3:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | LINGO3 | 645191 | 1560439_at | -0.1257 | 0.5672 | |
| GSE26886 | LINGO3 | 645191 | 1560439_at | -0.1899 | 0.0482 | |
| GSE45670 | LINGO3 | 645191 | 1560439_at | -0.0421 | 0.6432 | |
| GSE63941 | LINGO3 | 645191 | 1560439_at | 0.0439 | 0.8112 | |
| GSE77861 | LINGO3 | 645191 | 1560439_at | -0.1120 | 0.3122 | |
| SRP133303 | LINGO3 | 645191 | RNAseq | -0.7560 | 0.0068 | |
| SRP159526 | LINGO3 | 645191 | RNAseq | -0.8724 | 0.2871 | |
| SRP219564 | LINGO3 | 645191 | RNAseq | -0.0517 | 0.9213 | |
| TCGA | LINGO3 | 645191 | RNAseq | -0.4875 | 0.3757 |
Upregulated datasets: 0; Downregulated datasets: 0.
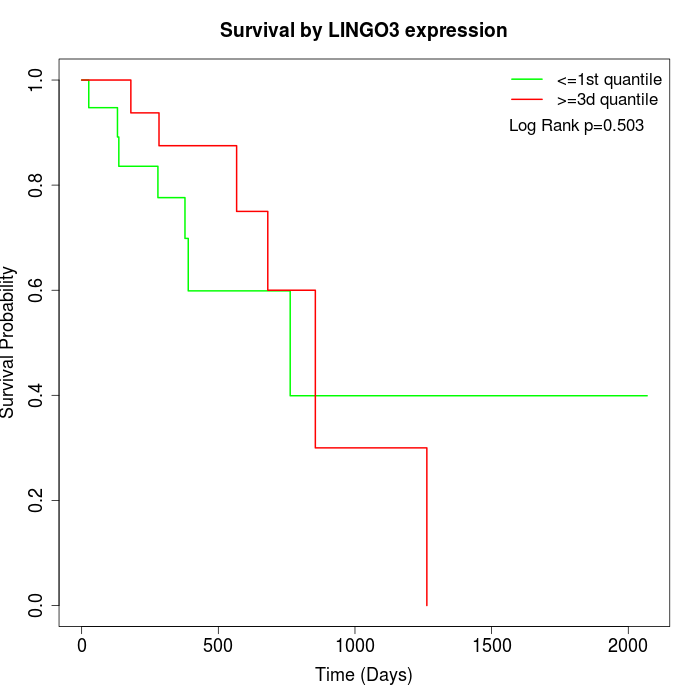
Survival by LINGO3 expression:
Note: Click image to view full size file.
Copy number change of LINGO3:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | LINGO3 | 645191 | 4 | 4 | 22 | |
| GSE20123 | LINGO3 | 645191 | 4 | 4 | 22 | |
| GSE43470 | LINGO3 | 645191 | 1 | 10 | 32 | |
| GSE46452 | LINGO3 | 645191 | 47 | 1 | 11 | |
| GSE47630 | LINGO3 | 645191 | 5 | 7 | 28 | |
| GSE54993 | LINGO3 | 645191 | 16 | 3 | 51 | |
| GSE54994 | LINGO3 | 645191 | 8 | 15 | 30 | |
| GSE60625 | LINGO3 | 645191 | 9 | 0 | 2 | |
| GSE74703 | LINGO3 | 645191 | 1 | 7 | 28 | |
| GSE74704 | LINGO3 | 645191 | 2 | 3 | 15 | |
| TCGA | LINGO3 | 645191 | 6 | 24 | 66 |
Total number of gains: 103; Total number of losses: 78; Total Number of normals: 307.
Somatic mutations of LINGO3:
Generating mutation plots.
Highly correlated genes for LINGO3:
Showing top 20/119 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| LINGO3 | RORC | 0.723821 | 3 | 0 | 3 |
| LINGO3 | ARMC4 | 0.692516 | 3 | 0 | 3 |
| LINGO3 | ZNF853 | 0.665997 | 3 | 0 | 3 |
| LINGO3 | TBATA | 0.665162 | 3 | 0 | 3 |
| LINGO3 | NANOS3 | 0.662315 | 3 | 0 | 3 |
| LINGO3 | PRRG3 | 0.655217 | 3 | 0 | 3 |
| LINGO3 | CYP11B1 | 0.652205 | 3 | 0 | 3 |
| LINGO3 | SIGLEC1 | 0.648348 | 4 | 0 | 3 |
| LINGO3 | PRSS58 | 0.646516 | 4 | 0 | 4 |
| LINGO3 | TGM7 | 0.641598 | 3 | 0 | 3 |
| LINGO3 | LMAN1L | 0.627409 | 5 | 0 | 4 |
| LINGO3 | CACNA2D4 | 0.626589 | 3 | 0 | 3 |
| LINGO3 | DNAI1 | 0.615538 | 3 | 0 | 3 |
| LINGO3 | HOXB1 | 0.614896 | 4 | 0 | 3 |
| LINGO3 | ITIH1 | 0.614676 | 4 | 0 | 3 |
| LINGO3 | INSL3 | 0.614261 | 5 | 0 | 4 |
| LINGO3 | DHRS7C | 0.61398 | 3 | 0 | 3 |
| LINGO3 | ACTR8 | 0.613871 | 4 | 0 | 3 |
| LINGO3 | CHIA | 0.613645 | 3 | 0 | 3 |
| LINGO3 | TEKT1 | 0.608517 | 4 | 0 | 3 |
For details and further investigation, click here