| Full name: LKAAEAR motif containing 1 | Alias Symbol: | ||
| Type: protein-coding gene | Cytoband: 20q13.33 | ||
| Entrez ID: 198437 | HGNC ID: HGNC:33718 | Ensembl Gene: ENSG00000171695 | OMIM ID: |
| Drug and gene relationship at DGIdb | |||
Expression of LKAAEAR1:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | LKAAEAR1 | 198437 | 1554977_at | -0.0082 | 0.9826 | |
| GSE26886 | LKAAEAR1 | 198437 | 1554977_at | 0.2441 | 0.1154 | |
| GSE45670 | LKAAEAR1 | 198437 | 1554977_at | 0.1136 | 0.2556 | |
| GSE53622 | LKAAEAR1 | 198437 | 64977 | 0.1597 | 0.0050 | |
| GSE53624 | LKAAEAR1 | 198437 | 64977 | 0.2906 | 0.0000 | |
| GSE63941 | LKAAEAR1 | 198437 | 1554977_at | 0.7744 | 0.0002 | |
| GSE77861 | LKAAEAR1 | 198437 | 1554977_at | -0.0850 | 0.5732 |
Upregulated datasets: 0; Downregulated datasets: 0.
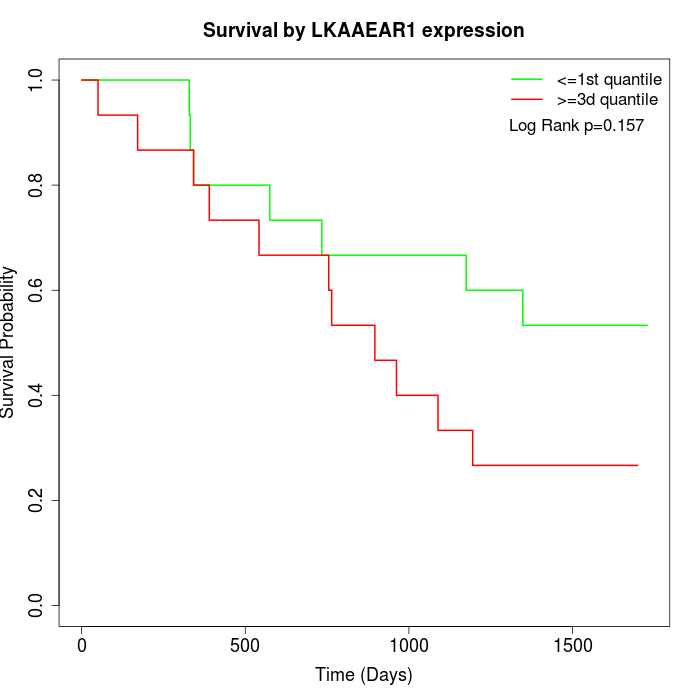
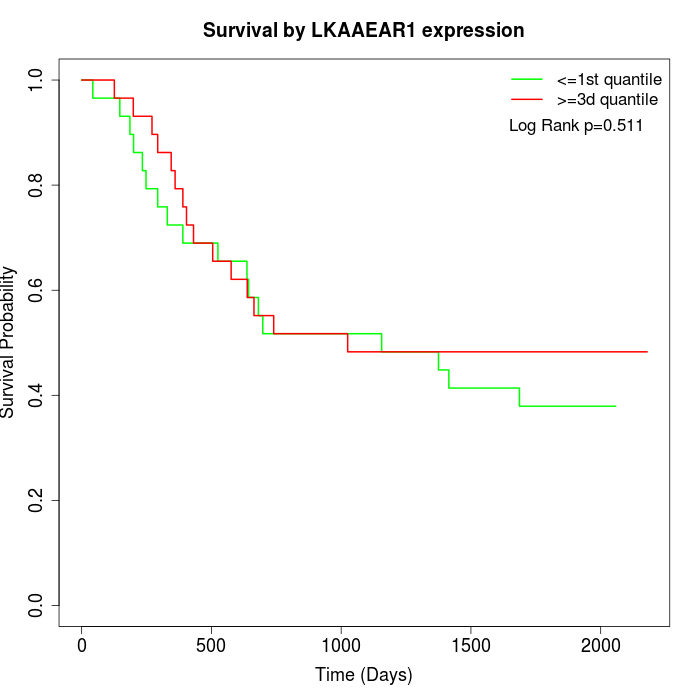
Survival by LKAAEAR1 expression:
Note: Click image to view full size file.
Copy number change of LKAAEAR1:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | LKAAEAR1 | 198437 | 13 | 2 | 15 | |
| GSE20123 | LKAAEAR1 | 198437 | 13 | 2 | 15 | |
| GSE43470 | LKAAEAR1 | 198437 | 11 | 1 | 31 | |
| GSE46452 | LKAAEAR1 | 198437 | 35 | 0 | 24 | |
| GSE47630 | LKAAEAR1 | 198437 | 25 | 1 | 14 | |
| GSE54993 | LKAAEAR1 | 198437 | 0 | 17 | 53 | |
| GSE54994 | LKAAEAR1 | 198437 | 29 | 0 | 24 | |
| GSE60625 | LKAAEAR1 | 198437 | 0 | 0 | 11 | |
| GSE74703 | LKAAEAR1 | 198437 | 10 | 0 | 26 | |
| GSE74704 | LKAAEAR1 | 198437 | 10 | 0 | 10 |
Total number of gains: 146; Total number of losses: 23; Total Number of normals: 223.
Somatic mutations of LKAAEAR1:
Generating mutation plots.
Highly correlated genes for LKAAEAR1:
Showing top 20/114 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| LKAAEAR1 | EFNA3 | 0.739104 | 3 | 0 | 3 |
| LKAAEAR1 | AATK | 0.713342 | 3 | 0 | 3 |
| LKAAEAR1 | DMPK | 0.701076 | 3 | 0 | 3 |
| LKAAEAR1 | GLP1R | 0.693352 | 3 | 0 | 3 |
| LKAAEAR1 | TSACC | 0.683412 | 3 | 0 | 3 |
| LKAAEAR1 | ELFN1 | 0.682897 | 3 | 0 | 3 |
| LKAAEAR1 | FHAD1 | 0.667713 | 3 | 0 | 3 |
| LKAAEAR1 | CACNA1A | 0.66338 | 3 | 0 | 3 |
| LKAAEAR1 | LINC00491 | 0.65544 | 3 | 0 | 3 |
| LKAAEAR1 | EPN1 | 0.63984 | 4 | 0 | 3 |
| LKAAEAR1 | DNHD1 | 0.639731 | 4 | 0 | 3 |
| LKAAEAR1 | UTS2R | 0.639247 | 7 | 0 | 5 |
| LKAAEAR1 | PHEX | 0.632996 | 3 | 0 | 3 |
| LKAAEAR1 | CNIH2 | 0.63268 | 4 | 0 | 3 |
| LKAAEAR1 | CCDC105 | 0.631739 | 3 | 0 | 3 |
| LKAAEAR1 | SLC5A2 | 0.62895 | 4 | 0 | 3 |
| LKAAEAR1 | CLPS | 0.628193 | 4 | 0 | 3 |
| LKAAEAR1 | PLCH2 | 0.62689 | 4 | 0 | 3 |
| LKAAEAR1 | SIRT6 | 0.625862 | 3 | 0 | 3 |
| LKAAEAR1 | LENEP | 0.621107 | 3 | 0 | 3 |
For details and further investigation, click here