| Full name: lymphocyte antigen 75 | Alias Symbol: DEC-205|CLEC13B|CD205 | ||
| Type: protein-coding gene | Cytoband: 2q24.2 | ||
| Entrez ID: 4065 | HGNC ID: HGNC:6729 | Ensembl Gene: ENSG00000054219 | OMIM ID: 604524 |
Expression of LY75:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | LY75 | 4065 | 205668_at | 0.7130 | 0.4361 | |
| GSE20347 | LY75 | 4065 | 205668_at | -0.6656 | 0.1936 | |
| GSE23400 | LY75 | 4065 | 205668_at | 0.2705 | 0.0965 | |
| GSE26886 | LY75 | 4065 | 205668_at | 1.2501 | 0.0703 | |
| GSE29001 | LY75 | 4065 | 205668_at | 0.2365 | 0.7627 | |
| GSE38129 | LY75 | 4065 | 205668_at | 0.0537 | 0.9069 | |
| GSE45670 | LY75 | 4065 | 205668_at | -0.2284 | 0.6126 | |
| GSE53622 | LY75 | 4065 | 113509 | -0.0019 | 0.9933 | |
| GSE53624 | LY75 | 4065 | 113509 | -0.2881 | 0.0726 | |
| GSE63941 | LY75 | 4065 | 205668_at | 1.0828 | 0.5677 | |
| GSE77861 | LY75 | 4065 | 205668_at | -0.5362 | 0.3645 | |
| GSE97050 | LY75 | 4065 | A_23_P334173 | 0.5042 | 0.3506 | |
| SRP007169 | LY75 | 4065 | RNAseq | 1.5744 | 0.0005 | |
| SRP008496 | LY75 | 4065 | RNAseq | 1.1520 | 0.0041 | |
| SRP064894 | LY75 | 4065 | RNAseq | 0.3287 | 0.3666 | |
| SRP133303 | LY75 | 4065 | RNAseq | 0.1397 | 0.5439 | |
| SRP159526 | LY75 | 4065 | RNAseq | -0.0205 | 0.9734 | |
| SRP193095 | LY75 | 4065 | RNAseq | -0.1641 | 0.5062 | |
| SRP219564 | LY75 | 4065 | RNAseq | -0.1458 | 0.8818 | |
| TCGA | LY75 | 4065 | RNAseq | -0.0503 | 0.7904 |
Upregulated datasets: 2; Downregulated datasets: 0.
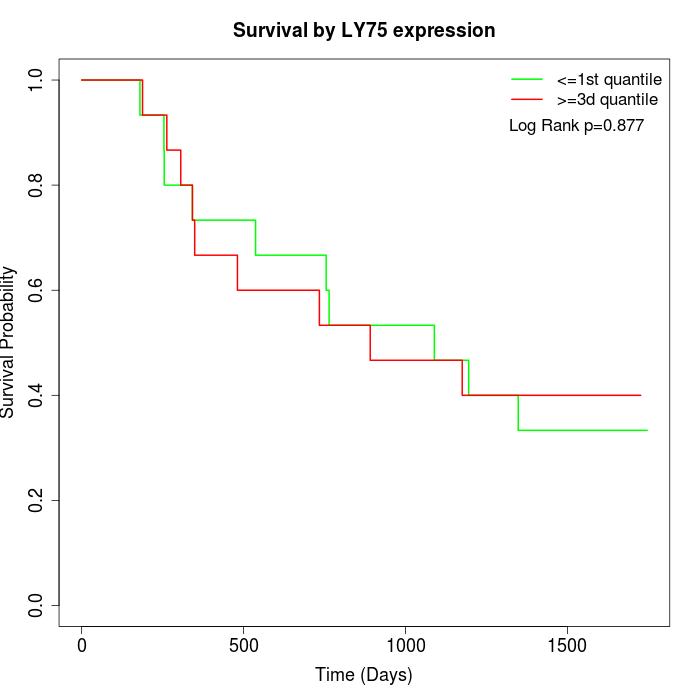
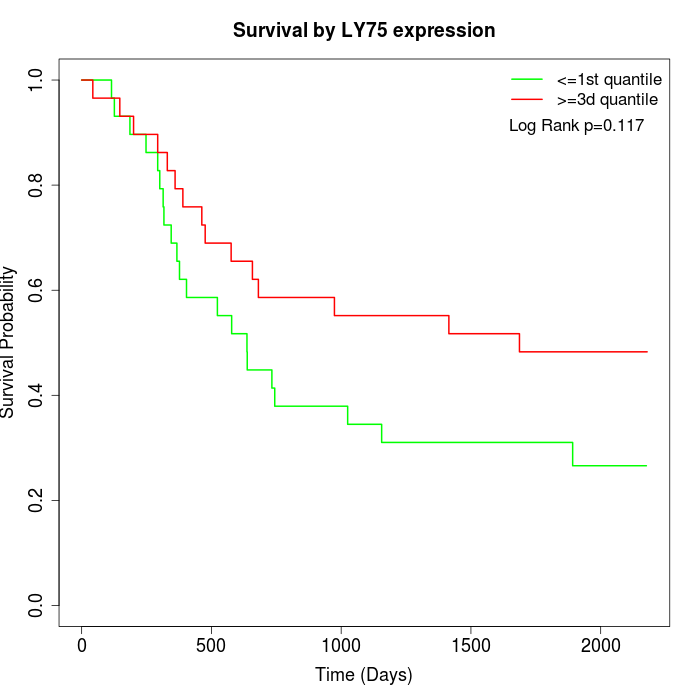
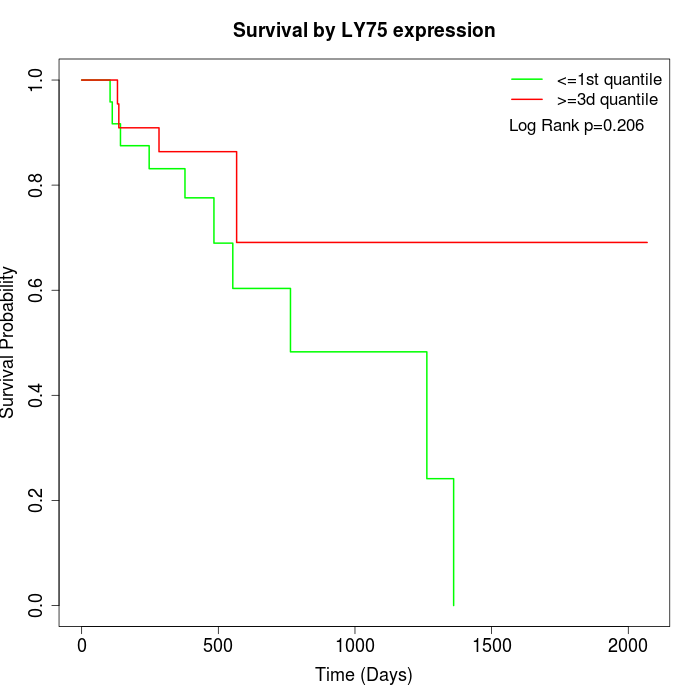
Survival by LY75 expression:
Note: Click image to view full size file.
Copy number change of LY75:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | LY75 | 4065 | 5 | 1 | 24 | |
| GSE20123 | LY75 | 4065 | 5 | 1 | 24 | |
| GSE43470 | LY75 | 4065 | 4 | 1 | 38 | |
| GSE46452 | LY75 | 4065 | 1 | 4 | 54 | |
| GSE47630 | LY75 | 4065 | 5 | 3 | 32 | |
| GSE54993 | LY75 | 4065 | 0 | 4 | 66 | |
| GSE54994 | LY75 | 4065 | 11 | 2 | 40 | |
| GSE60625 | LY75 | 4065 | 0 | 3 | 8 | |
| GSE74703 | LY75 | 4065 | 3 | 1 | 32 | |
| GSE74704 | LY75 | 4065 | 3 | 0 | 17 | |
| TCGA | LY75 | 4065 | 23 | 10 | 63 |
Total number of gains: 60; Total number of losses: 30; Total Number of normals: 398.
Somatic mutations of LY75:
Generating mutation plots.
Highly correlated genes for LY75:
Showing top 20/292 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| LY75 | SMARCA5 | 0.794353 | 3 | 0 | 3 |
| LY75 | PPIL1 | 0.784415 | 3 | 0 | 3 |
| LY75 | MGAT4B | 0.777318 | 3 | 0 | 3 |
| LY75 | OSTC | 0.769386 | 3 | 0 | 3 |
| LY75 | TET2 | 0.761709 | 3 | 0 | 3 |
| LY75 | EIF3L | 0.760512 | 3 | 0 | 3 |
| LY75 | ASXL2 | 0.755054 | 3 | 0 | 3 |
| LY75 | CNOT2 | 0.733052 | 3 | 0 | 3 |
| LY75 | SRP9 | 0.727534 | 3 | 0 | 3 |
| LY75 | RPP38 | 0.724039 | 4 | 0 | 4 |
| LY75 | B3GALT6 | 0.72398 | 3 | 0 | 3 |
| LY75 | NUDCD1 | 0.722352 | 3 | 0 | 3 |
| LY75 | AP1AR | 0.720865 | 3 | 0 | 3 |
| LY75 | UBE2A | 0.720457 | 3 | 0 | 3 |
| LY75 | UHRF1BP1 | 0.716535 | 3 | 0 | 3 |
| LY75 | FNDC3A | 0.713894 | 3 | 0 | 3 |
| LY75 | ANAPC10 | 0.712067 | 4 | 0 | 4 |
| LY75 | PRRC1 | 0.709669 | 3 | 0 | 3 |
| LY75 | ZNF614 | 0.707801 | 3 | 0 | 3 |
| LY75 | ANKRD13C | 0.706185 | 3 | 0 | 3 |
For details and further investigation, click here