| Full name: mitogen-activated protein kinase kinase 6 | Alias Symbol: MEK6|MKK6|SAPKK3|MAPKK6 | ||
| Type: protein-coding gene | Cytoband: 17q24.3 | ||
| Entrez ID: 5608 | HGNC ID: HGNC:6846 | Ensembl Gene: ENSG00000108984 | OMIM ID: 601254 |
| Related drugs: BINIMETINIB, COBIMETINIB, DABRAFENIB MESYLATE, PD-0325901, Refametinib, SELUMETINIB, STAUROSPORINE, TRAMETINIB, WX-554... [more] | |||
MAP2K6 involved pathways:
Expression of MAP2K6:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | MAP2K6 | 5608 | 205698_s_at | -0.1483 | 0.7429 | |
| GSE20347 | MAP2K6 | 5608 | 205698_s_at | -0.2379 | 0.0589 | |
| GSE23400 | MAP2K6 | 5608 | 205698_s_at | -0.1586 | 0.0027 | |
| GSE26886 | MAP2K6 | 5608 | 205698_s_at | -0.6834 | 0.0397 | |
| GSE29001 | MAP2K6 | 5608 | 205698_s_at | -0.1855 | 0.5403 | |
| GSE38129 | MAP2K6 | 5608 | 205698_s_at | -0.1650 | 0.3234 | |
| GSE45670 | MAP2K6 | 5608 | 205698_s_at | -0.1099 | 0.7574 | |
| GSE53622 | MAP2K6 | 5608 | 28139 | -0.8402 | 0.0000 | |
| GSE53624 | MAP2K6 | 5608 | 28139 | -0.5517 | 0.0001 | |
| GSE63941 | MAP2K6 | 5608 | 205698_s_at | 0.9152 | 0.3796 | |
| GSE77861 | MAP2K6 | 5608 | 205698_s_at | -0.2283 | 0.4306 | |
| GSE97050 | MAP2K6 | 5608 | A_23_P207445 | -0.3353 | 0.2495 | |
| SRP007169 | MAP2K6 | 5608 | RNAseq | -1.4145 | 0.0006 | |
| SRP008496 | MAP2K6 | 5608 | RNAseq | -0.5408 | 0.1378 | |
| SRP064894 | MAP2K6 | 5608 | RNAseq | -0.0602 | 0.7649 | |
| SRP133303 | MAP2K6 | 5608 | RNAseq | -0.7072 | 0.0811 | |
| SRP159526 | MAP2K6 | 5608 | RNAseq | -0.4394 | 0.3191 | |
| SRP193095 | MAP2K6 | 5608 | RNAseq | -0.8598 | 0.0001 | |
| SRP219564 | MAP2K6 | 5608 | RNAseq | -0.1052 | 0.8428 | |
| TCGA | MAP2K6 | 5608 | RNAseq | 0.0700 | 0.7482 |
Upregulated datasets: 0; Downregulated datasets: 1.
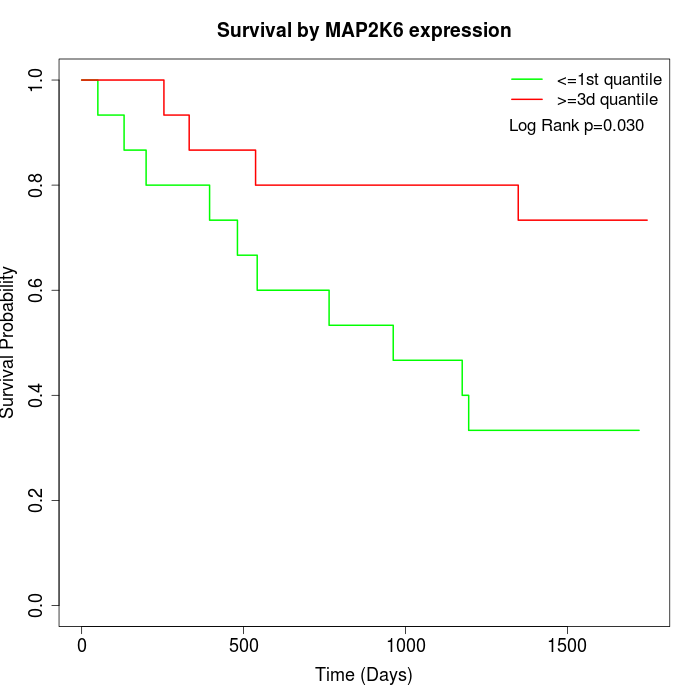
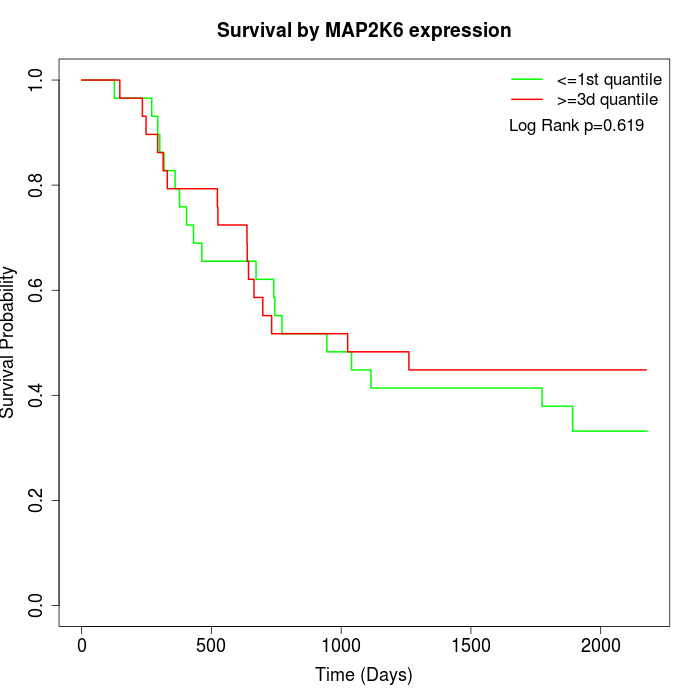
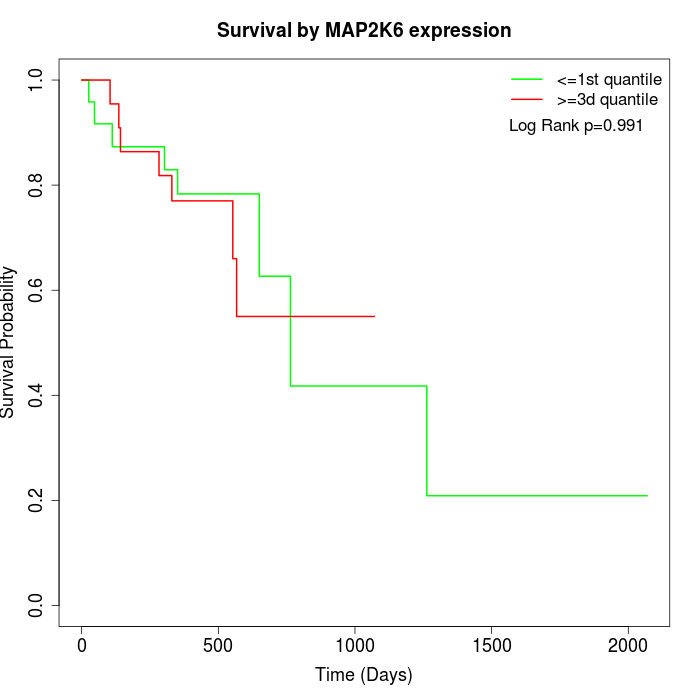
Survival by MAP2K6 expression:
Note: Click image to view full size file.
Copy number change of MAP2K6:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | MAP2K6 | 5608 | 3 | 1 | 26 | |
| GSE20123 | MAP2K6 | 5608 | 3 | 1 | 26 | |
| GSE43470 | MAP2K6 | 5608 | 6 | 0 | 37 | |
| GSE46452 | MAP2K6 | 5608 | 31 | 1 | 27 | |
| GSE47630 | MAP2K6 | 5608 | 7 | 1 | 32 | |
| GSE54993 | MAP2K6 | 5608 | 2 | 5 | 63 | |
| GSE54994 | MAP2K6 | 5608 | 9 | 4 | 40 | |
| GSE60625 | MAP2K6 | 5608 | 4 | 3 | 4 | |
| GSE74703 | MAP2K6 | 5608 | 6 | 0 | 30 | |
| GSE74704 | MAP2K6 | 5608 | 3 | 1 | 16 | |
| TCGA | MAP2K6 | 5608 | 31 | 10 | 55 |
Total number of gains: 105; Total number of losses: 27; Total Number of normals: 356.
Somatic mutations of MAP2K6:
Generating mutation plots.
Highly correlated genes for MAP2K6:
Showing top 20/231 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| MAP2K6 | KCNE3 | 0.771725 | 3 | 0 | 3 |
| MAP2K6 | KHK | 0.723178 | 3 | 0 | 3 |
| MAP2K6 | SFT2D3 | 0.711747 | 3 | 0 | 3 |
| MAP2K6 | GSTT1 | 0.709292 | 3 | 0 | 3 |
| MAP2K6 | SETX | 0.700265 | 3 | 0 | 3 |
| MAP2K6 | ZNF175 | 0.690639 | 4 | 0 | 3 |
| MAP2K6 | RBM4B | 0.688573 | 3 | 0 | 3 |
| MAP2K6 | ZNF564 | 0.680028 | 3 | 0 | 3 |
| MAP2K6 | RNF14 | 0.675279 | 3 | 0 | 3 |
| MAP2K6 | TSPAN3 | 0.672922 | 3 | 0 | 3 |
| MAP2K6 | GPRASP2 | 0.666834 | 3 | 0 | 3 |
| MAP2K6 | ZDHHC2 | 0.655568 | 5 | 0 | 4 |
| MAP2K6 | FBXL17 | 0.655204 | 3 | 0 | 3 |
| MAP2K6 | ANAPC7 | 0.654735 | 3 | 0 | 3 |
| MAP2K6 | NFE2L2 | 0.654733 | 4 | 0 | 4 |
| MAP2K6 | PARP16 | 0.650538 | 6 | 0 | 5 |
| MAP2K6 | RAB39B | 0.649977 | 4 | 0 | 3 |
| MAP2K6 | MKRN1 | 0.649327 | 4 | 0 | 4 |
| MAP2K6 | WDFY1 | 0.643169 | 3 | 0 | 3 |
| MAP2K6 | ACTR1A | 0.642657 | 3 | 0 | 3 |
For details and further investigation, click here