| Full name: potassium voltage-gated channel subfamily E regulatory subunit 3 | Alias Symbol: MiRP2|HOKPP | ||
| Type: protein-coding gene | Cytoband: 11q13.4 | ||
| Entrez ID: 10008 | HGNC ID: HGNC:6243 | Ensembl Gene: ENSG00000175538 | OMIM ID: 604433 |
Expression of KCNE3:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | KCNE3 | 10008 | 227647_at | 0.0703 | 0.9578 | |
| GSE26886 | KCNE3 | 10008 | 227647_at | 1.3073 | 0.0220 | |
| GSE45670 | KCNE3 | 10008 | 227647_at | -0.0339 | 0.9668 | |
| GSE63941 | KCNE3 | 10008 | 222923_s_at | 0.0357 | 0.8851 | |
| GSE77861 | KCNE3 | 10008 | 227647_at | 1.0555 | 0.1505 | |
| GSE97050 | KCNE3 | 10008 | A_23_P24948 | -0.1306 | 0.6253 | |
| SRP007169 | KCNE3 | 10008 | RNAseq | 2.2818 | 0.0021 | |
| SRP064894 | KCNE3 | 10008 | RNAseq | 0.5381 | 0.0726 | |
| SRP133303 | KCNE3 | 10008 | RNAseq | -0.0497 | 0.8434 | |
| SRP159526 | KCNE3 | 10008 | RNAseq | 1.6919 | 0.0110 | |
| SRP193095 | KCNE3 | 10008 | RNAseq | -0.5181 | 0.2398 | |
| SRP219564 | KCNE3 | 10008 | RNAseq | 1.3322 | 0.0491 | |
| TCGA | KCNE3 | 10008 | RNAseq | -0.1321 | 0.5710 |
Upregulated datasets: 4; Downregulated datasets: 0.
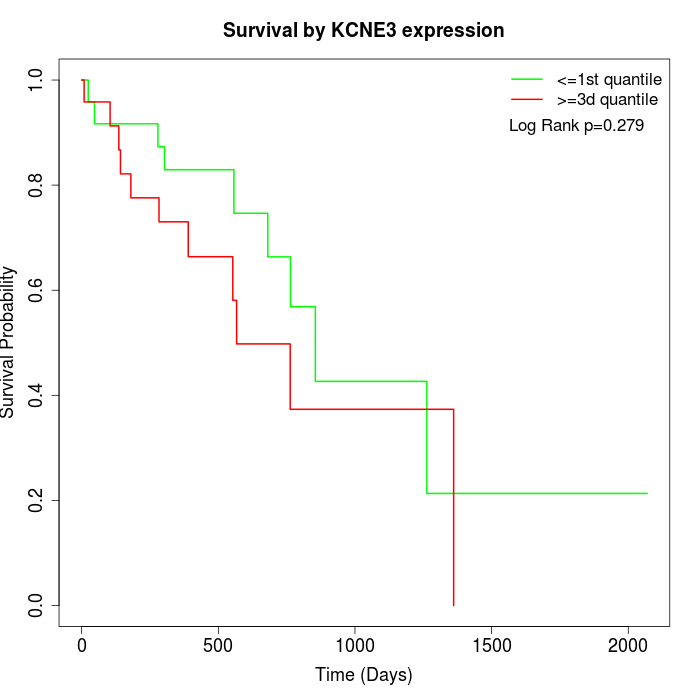
Survival by KCNE3 expression:
Note: Click image to view full size file.
Copy number change of KCNE3:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | KCNE3 | 10008 | 6 | 9 | 15 | |
| GSE20123 | KCNE3 | 10008 | 6 | 8 | 16 | |
| GSE43470 | KCNE3 | 10008 | 5 | 4 | 34 | |
| GSE46452 | KCNE3 | 10008 | 7 | 22 | 30 | |
| GSE47630 | KCNE3 | 10008 | 6 | 14 | 20 | |
| GSE54993 | KCNE3 | 10008 | 8 | 3 | 59 | |
| GSE54994 | KCNE3 | 10008 | 8 | 15 | 30 | |
| GSE60625 | KCNE3 | 10008 | 4 | 3 | 4 | |
| GSE74703 | KCNE3 | 10008 | 4 | 3 | 29 | |
| GSE74704 | KCNE3 | 10008 | 4 | 4 | 12 | |
| TCGA | KCNE3 | 10008 | 23 | 32 | 41 |
Total number of gains: 81; Total number of losses: 117; Total Number of normals: 290.
Somatic mutations of KCNE3:
Generating mutation plots.
Highly correlated genes for KCNE3:
Showing top 20/368 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| KCNE3 | DHRSX | 0.827585 | 3 | 0 | 3 |
| KCNE3 | LIFR | 0.810465 | 3 | 0 | 3 |
| KCNE3 | TMEM38B | 0.804311 | 3 | 0 | 3 |
| KCNE3 | NUDCD1 | 0.793717 | 3 | 0 | 3 |
| KCNE3 | SPRY2 | 0.773618 | 3 | 0 | 3 |
| KCNE3 | MAP2K6 | 0.771725 | 3 | 0 | 3 |
| KCNE3 | SACS | 0.771563 | 3 | 0 | 3 |
| KCNE3 | PTDSS1 | 0.76624 | 3 | 0 | 3 |
| KCNE3 | FZD6 | 0.762786 | 3 | 0 | 3 |
| KCNE3 | NAPEPLD | 0.76149 | 3 | 0 | 3 |
| KCNE3 | FBXL4 | 0.751818 | 3 | 0 | 3 |
| KCNE3 | HSF2 | 0.747716 | 3 | 0 | 3 |
| KCNE3 | MED30 | 0.747465 | 3 | 0 | 3 |
| KCNE3 | LIG3 | 0.747217 | 3 | 0 | 3 |
| KCNE3 | DACT2 | 0.740469 | 3 | 0 | 3 |
| KCNE3 | ZXDA | 0.735745 | 3 | 0 | 3 |
| KCNE3 | BRD9 | 0.733127 | 3 | 0 | 3 |
| KCNE3 | TMTC4 | 0.732293 | 4 | 0 | 4 |
| KCNE3 | DAP | 0.730924 | 4 | 0 | 4 |
| KCNE3 | SGCB | 0.729812 | 3 | 0 | 3 |
For details and further investigation, click here