| Full name: mutL homolog 1 | Alias Symbol: HNPCC|FCC2|HNPCC2 | ||
| Type: protein-coding gene | Cytoband: 3p22.2 | ||
| Entrez ID: 4292 | HGNC ID: HGNC:7127 | Ensembl Gene: ENSG00000076242 | OMIM ID: 120436 |
| Related drugs: ATEZOLIZUMAB, IPILIMUMAB, NIVOLUMAB, PEMBROLIZUMAB... [more] | |||
Screen Evidence:
| |||
MLH1 involved pathways:
| KEGG pathway | Description | View |
|---|---|---|
| hsa03460 | Fanconi anemia pathway |
Expression of MLH1:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | MLH1 | 4292 | 202520_s_at | -0.0311 | 0.9367 | |
| GSE20347 | MLH1 | 4292 | 202520_s_at | -0.2041 | 0.0800 | |
| GSE23400 | MLH1 | 4292 | 202520_s_at | 0.0971 | 0.0791 | |
| GSE26886 | MLH1 | 4292 | 202520_s_at | -0.3138 | 0.1453 | |
| GSE29001 | MLH1 | 4292 | 202520_s_at | 0.0507 | 0.8986 | |
| GSE38129 | MLH1 | 4292 | 202520_s_at | -0.2220 | 0.0064 | |
| GSE45670 | MLH1 | 4292 | 202520_s_at | -0.1683 | 0.2187 | |
| GSE53622 | MLH1 | 4292 | 17044 | -0.1517 | 0.0085 | |
| GSE53624 | MLH1 | 4292 | 17044 | -0.1139 | 0.1799 | |
| GSE63941 | MLH1 | 4292 | 202520_s_at | -0.2913 | 0.3531 | |
| GSE77861 | MLH1 | 4292 | 202520_s_at | -0.0657 | 0.8130 | |
| GSE97050 | MLH1 | 4292 | A_23_P69058 | 0.0234 | 0.9454 | |
| SRP007169 | MLH1 | 4292 | RNAseq | -0.2294 | 0.5255 | |
| SRP008496 | MLH1 | 4292 | RNAseq | -0.1320 | 0.5321 | |
| SRP064894 | MLH1 | 4292 | RNAseq | 0.0491 | 0.6462 | |
| SRP133303 | MLH1 | 4292 | RNAseq | -0.1213 | 0.2174 | |
| SRP159526 | MLH1 | 4292 | RNAseq | -0.2423 | 0.3340 | |
| SRP193095 | MLH1 | 4292 | RNAseq | -0.2056 | 0.0188 | |
| SRP219564 | MLH1 | 4292 | RNAseq | -0.1906 | 0.6643 | |
| TCGA | MLH1 | 4292 | RNAseq | -0.1301 | 0.0145 |
Upregulated datasets: 0; Downregulated datasets: 0.
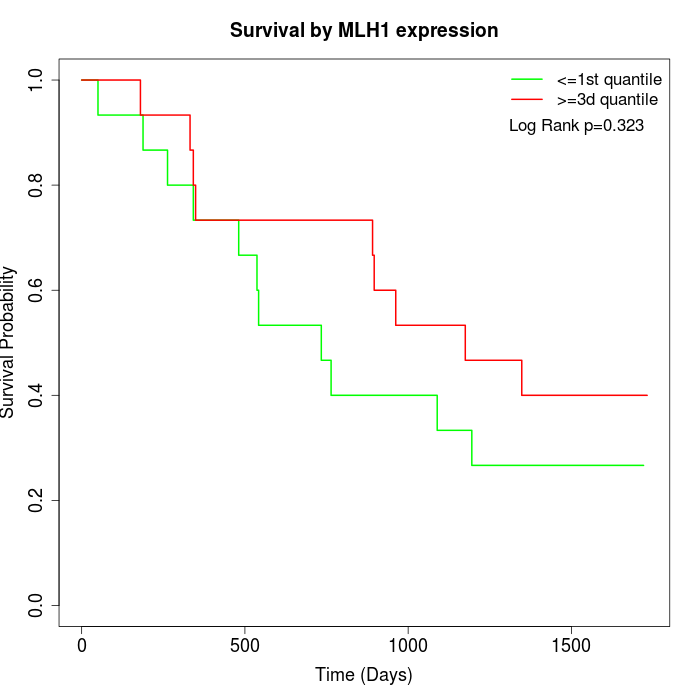
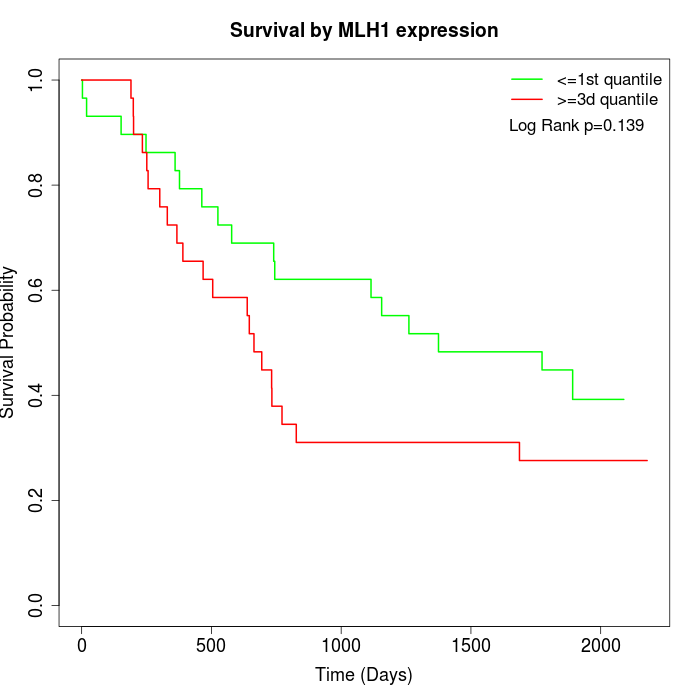
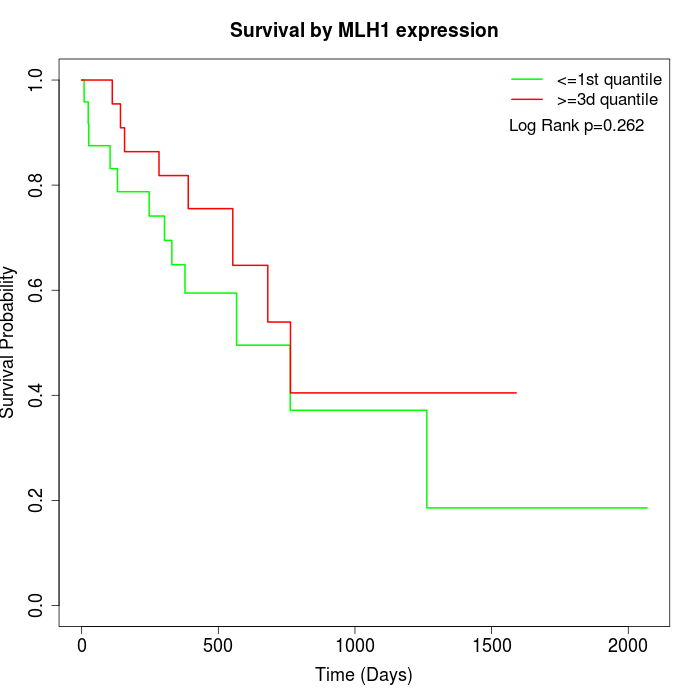
Survival by MLH1 expression:
Note: Click image to view full size file.
Copy number change of MLH1:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | MLH1 | 4292 | 0 | 18 | 12 | |
| GSE20123 | MLH1 | 4292 | 0 | 18 | 12 | |
| GSE43470 | MLH1 | 4292 | 0 | 20 | 23 | |
| GSE46452 | MLH1 | 4292 | 2 | 17 | 40 | |
| GSE47630 | MLH1 | 4292 | 1 | 25 | 14 | |
| GSE54993 | MLH1 | 4292 | 6 | 3 | 61 | |
| GSE54994 | MLH1 | 4292 | 1 | 32 | 20 | |
| GSE60625 | MLH1 | 4292 | 5 | 0 | 6 | |
| GSE74703 | MLH1 | 4292 | 0 | 16 | 20 | |
| GSE74704 | MLH1 | 4292 | 0 | 12 | 8 | |
| TCGA | MLH1 | 4292 | 0 | 71 | 25 |
Total number of gains: 15; Total number of losses: 232; Total Number of normals: 241.
Somatic mutations of MLH1:
Generating mutation plots.
Highly correlated genes for MLH1:
Showing top 20/429 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| MLH1 | TUBGCP3 | 0.76867 | 3 | 0 | 3 |
| MLH1 | CHD1 | 0.759542 | 4 | 0 | 4 |
| MLH1 | FARS2 | 0.73675 | 3 | 0 | 3 |
| MLH1 | ZNF383 | 0.730737 | 3 | 0 | 3 |
| MLH1 | B3GALNT2 | 0.727917 | 3 | 0 | 3 |
| MLH1 | NXT2 | 0.71977 | 3 | 0 | 3 |
| MLH1 | BIVM | 0.707322 | 3 | 0 | 3 |
| MLH1 | ATG2B | 0.702475 | 3 | 0 | 3 |
| MLH1 | KIF16B | 0.702123 | 3 | 0 | 3 |
| MLH1 | PARP8 | 0.697512 | 3 | 0 | 3 |
| MLH1 | ASXL2 | 0.696339 | 3 | 0 | 3 |
| MLH1 | MRPL48 | 0.695601 | 3 | 0 | 3 |
| MLH1 | TUBGCP4 | 0.694578 | 3 | 0 | 3 |
| MLH1 | FPGT | 0.691621 | 3 | 0 | 3 |
| MLH1 | ZBTB11 | 0.687798 | 3 | 0 | 3 |
| MLH1 | CYCS | 0.685096 | 3 | 0 | 3 |
| MLH1 | UBE2Q2 | 0.683582 | 4 | 0 | 4 |
| MLH1 | SNTB2 | 0.679349 | 4 | 0 | 4 |
| MLH1 | ATF6 | 0.677586 | 4 | 0 | 3 |
| MLH1 | C1QBP | 0.677392 | 3 | 0 | 3 |
For details and further investigation, click here