| Full name: SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily d, member 2 | Alias Symbol: BAF60B|Rsc6p|CRACD2|PRO2451 | ||
| Type: protein-coding gene | Cytoband: 17q23.3 | ||
| Entrez ID: 6603 | HGNC ID: HGNC:11107 | Ensembl Gene: ENSG00000108604 | OMIM ID: 601736 |
Expression of SMARCD2:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | SMARCD2 | 6603 | 201827_at | 0.0844 | 0.8501 | |
| GSE20347 | SMARCD2 | 6603 | 201827_at | -0.1590 | 0.1634 | |
| GSE23400 | SMARCD2 | 6603 | 201827_at | -0.0293 | 0.6572 | |
| GSE26886 | SMARCD2 | 6603 | 201827_at | 0.2179 | 0.3161 | |
| GSE29001 | SMARCD2 | 6603 | 201827_at | -0.0958 | 0.8550 | |
| GSE38129 | SMARCD2 | 6603 | 201827_at | 0.0800 | 0.5684 | |
| GSE45670 | SMARCD2 | 6603 | 201827_at | 0.1253 | 0.5266 | |
| GSE63941 | SMARCD2 | 6603 | 201827_at | -0.0598 | 0.8864 | |
| GSE77861 | SMARCD2 | 6603 | 201827_at | 0.2677 | 0.3101 | |
| GSE97050 | SMARCD2 | 6603 | A_23_P66487 | 0.2185 | 0.5248 | |
| SRP007169 | SMARCD2 | 6603 | RNAseq | -0.5525 | 0.0585 | |
| SRP008496 | SMARCD2 | 6603 | RNAseq | -0.3244 | 0.1502 | |
| SRP064894 | SMARCD2 | 6603 | RNAseq | -0.0404 | 0.8245 | |
| SRP133303 | SMARCD2 | 6603 | RNAseq | 0.0639 | 0.8017 | |
| SRP159526 | SMARCD2 | 6603 | RNAseq | 0.2643 | 0.0799 | |
| SRP193095 | SMARCD2 | 6603 | RNAseq | -0.1977 | 0.0598 | |
| SRP219564 | SMARCD2 | 6603 | RNAseq | 0.3422 | 0.4255 | |
| TCGA | SMARCD2 | 6603 | RNAseq | 0.0094 | 0.8392 |
Upregulated datasets: 0; Downregulated datasets: 0.
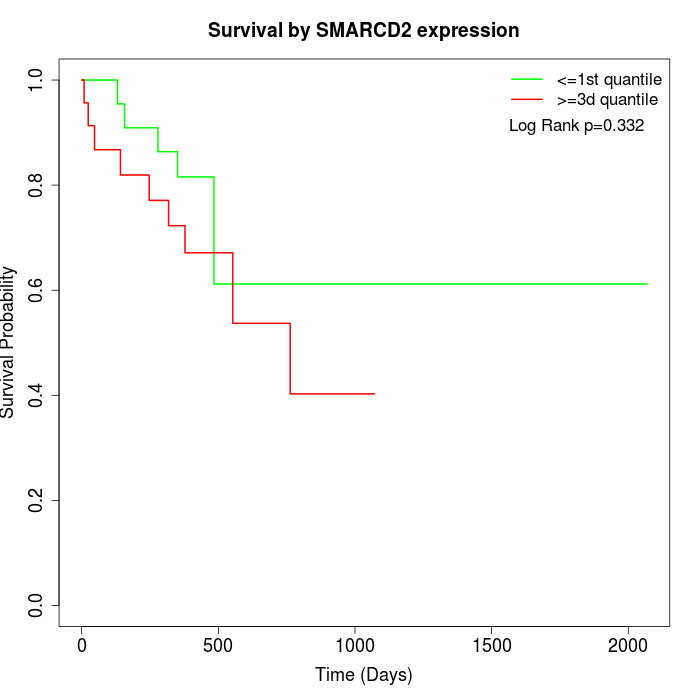
Survival by SMARCD2 expression:
Note: Click image to view full size file.
Copy number change of SMARCD2:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | SMARCD2 | 6603 | 3 | 1 | 26 | |
| GSE20123 | SMARCD2 | 6603 | 3 | 1 | 26 | |
| GSE43470 | SMARCD2 | 6603 | 4 | 0 | 39 | |
| GSE46452 | SMARCD2 | 6603 | 31 | 0 | 28 | |
| GSE47630 | SMARCD2 | 6603 | 7 | 1 | 32 | |
| GSE54993 | SMARCD2 | 6603 | 2 | 5 | 63 | |
| GSE54994 | SMARCD2 | 6603 | 9 | 4 | 40 | |
| GSE60625 | SMARCD2 | 6603 | 4 | 0 | 7 | |
| GSE74703 | SMARCD2 | 6603 | 4 | 0 | 32 | |
| GSE74704 | SMARCD2 | 6603 | 3 | 1 | 16 | |
| TCGA | SMARCD2 | 6603 | 31 | 9 | 56 |
Total number of gains: 101; Total number of losses: 22; Total Number of normals: 365.
Somatic mutations of SMARCD2:
Generating mutation plots.
Highly correlated genes for SMARCD2:
Showing top 20/345 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| SMARCD2 | SBK1 | 0.764014 | 3 | 0 | 3 |
| SMARCD2 | EID2 | 0.763761 | 3 | 0 | 3 |
| SMARCD2 | POLR2A | 0.753334 | 4 | 0 | 4 |
| SMARCD2 | PRR13 | 0.748699 | 4 | 0 | 3 |
| SMARCD2 | RHBDD2 | 0.736563 | 3 | 0 | 3 |
| SMARCD2 | PRKCSH | 0.728886 | 3 | 0 | 3 |
| SMARCD2 | GTF2IRD1 | 0.728295 | 4 | 0 | 4 |
| SMARCD2 | CUL7 | 0.724963 | 3 | 0 | 3 |
| SMARCD2 | ZNF766 | 0.723921 | 3 | 0 | 3 |
| SMARCD2 | DMAP1 | 0.719064 | 3 | 0 | 3 |
| SMARCD2 | PPP1R37 | 0.718365 | 3 | 0 | 3 |
| SMARCD2 | LEMD2 | 0.713672 | 3 | 0 | 3 |
| SMARCD2 | SLAIN1 | 0.71037 | 3 | 0 | 3 |
| SMARCD2 | AKT2 | 0.709675 | 3 | 0 | 3 |
| SMARCD2 | POP5 | 0.705128 | 3 | 0 | 3 |
| SMARCD2 | TNFSF13 | 0.703026 | 4 | 0 | 3 |
| SMARCD2 | SFXN4 | 0.700668 | 3 | 0 | 3 |
| SMARCD2 | SLC25A19 | 0.700268 | 3 | 0 | 3 |
| SMARCD2 | FAM98C | 0.695068 | 3 | 0 | 3 |
| SMARCD2 | PTPRF | 0.693876 | 4 | 0 | 3 |
For details and further investigation, click here