| Full name: Ras related GTP binding C | Alias Symbol: GTR2|FLJ13311 | ||
| Type: protein-coding gene | Cytoband: 1p34.3 | ||
| Entrez ID: 64121 | HGNC ID: HGNC:19902 | Ensembl Gene: ENSG00000116954 | OMIM ID: 608267 |
Screen Evidence:
| |||
RRAGC involved pathways:
| KEGG pathway | Description | View |
|---|---|---|
| hsa04150 | mTOR signaling pathway |
Expression of RRAGC:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | RRAGC | 64121 | 218088_s_at | 0.0354 | 0.9563 | |
| GSE20347 | RRAGC | 64121 | 218088_s_at | -0.3109 | 0.0384 | |
| GSE23400 | RRAGC | 64121 | 218088_s_at | 0.0179 | 0.8368 | |
| GSE26886 | RRAGC | 64121 | 218088_s_at | -0.5611 | 0.0527 | |
| GSE29001 | RRAGC | 64121 | 218088_s_at | -0.3654 | 0.1421 | |
| GSE38129 | RRAGC | 64121 | 218088_s_at | -0.2005 | 0.1179 | |
| GSE45670 | RRAGC | 64121 | 218088_s_at | 0.0264 | 0.9029 | |
| GSE63941 | RRAGC | 64121 | 218088_s_at | -0.4284 | 0.2618 | |
| GSE77861 | RRAGC | 64121 | 218088_s_at | 0.0274 | 0.9413 | |
| GSE97050 | RRAGC | 64121 | A_23_P97623 | -0.0176 | 0.9369 | |
| SRP007169 | RRAGC | 64121 | RNAseq | -0.8532 | 0.0152 | |
| SRP008496 | RRAGC | 64121 | RNAseq | -0.9365 | 0.0002 | |
| SRP064894 | RRAGC | 64121 | RNAseq | -0.2585 | 0.1244 | |
| SRP133303 | RRAGC | 64121 | RNAseq | 0.2139 | 0.3249 | |
| SRP159526 | RRAGC | 64121 | RNAseq | -0.6567 | 0.0015 | |
| SRP193095 | RRAGC | 64121 | RNAseq | -0.2739 | 0.0361 | |
| SRP219564 | RRAGC | 64121 | RNAseq | -0.1267 | 0.7873 | |
| TCGA | RRAGC | 64121 | RNAseq | 0.1368 | 0.0664 |
Upregulated datasets: 0; Downregulated datasets: 0.
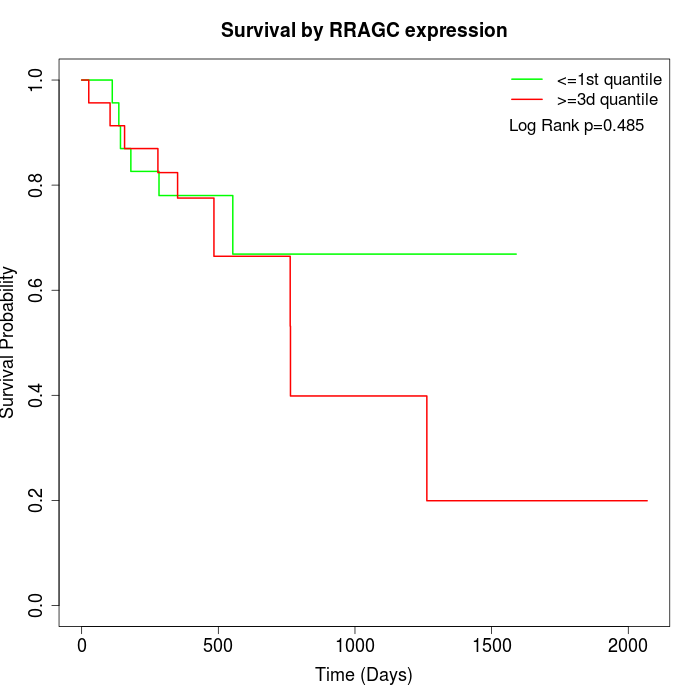
Survival by RRAGC expression:
Note: Click image to view full size file.
Copy number change of RRAGC:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | RRAGC | 64121 | 2 | 4 | 24 | |
| GSE20123 | RRAGC | 64121 | 2 | 3 | 25 | |
| GSE43470 | RRAGC | 64121 | 7 | 2 | 34 | |
| GSE46452 | RRAGC | 64121 | 4 | 1 | 54 | |
| GSE47630 | RRAGC | 64121 | 8 | 4 | 28 | |
| GSE54993 | RRAGC | 64121 | 0 | 1 | 69 | |
| GSE54994 | RRAGC | 64121 | 13 | 2 | 38 | |
| GSE60625 | RRAGC | 64121 | 0 | 0 | 11 | |
| GSE74703 | RRAGC | 64121 | 6 | 1 | 29 | |
| GSE74704 | RRAGC | 64121 | 1 | 0 | 19 | |
| TCGA | RRAGC | 64121 | 14 | 16 | 66 |
Total number of gains: 57; Total number of losses: 34; Total Number of normals: 397.
Somatic mutations of RRAGC:
Generating mutation plots.
Highly correlated genes for RRAGC:
Showing top 20/310 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| RRAGC | TRAPPC6B | 0.819318 | 3 | 0 | 3 |
| RRAGC | PPM1A | 0.814885 | 3 | 0 | 3 |
| RRAGC | SOS2 | 0.813431 | 3 | 0 | 3 |
| RRAGC | MAP4K5 | 0.807097 | 3 | 0 | 3 |
| RRAGC | ZCCHC8 | 0.79957 | 3 | 0 | 3 |
| RRAGC | ZNF766 | 0.787991 | 3 | 0 | 3 |
| RRAGC | TMEM50A | 0.771203 | 3 | 0 | 3 |
| RRAGC | TRIT1 | 0.771018 | 3 | 0 | 3 |
| RRAGC | GET4 | 0.761096 | 3 | 0 | 3 |
| RRAGC | PSMC6 | 0.758105 | 3 | 0 | 3 |
| RRAGC | ZNF436 | 0.755675 | 3 | 0 | 3 |
| RRAGC | STYX | 0.75433 | 3 | 0 | 3 |
| RRAGC | SNX6 | 0.75318 | 3 | 0 | 3 |
| RRAGC | PSEN1 | 0.751338 | 3 | 0 | 3 |
| RRAGC | CNOT6 | 0.743954 | 3 | 0 | 3 |
| RRAGC | PEX13 | 0.74314 | 4 | 0 | 4 |
| RRAGC | SEC22A | 0.741843 | 3 | 0 | 3 |
| RRAGC | VPS52 | 0.740208 | 3 | 0 | 3 |
| RRAGC | HARS2 | 0.738207 | 3 | 0 | 3 |
| RRAGC | IMPACT | 0.737593 | 3 | 0 | 3 |
For details and further investigation, click here