| Full name: C-C motif chemokine ligand 4 | Alias Symbol: MIP-1-beta|Act-2|AT744.1 | ||
| Type: protein-coding gene | Cytoband: 17q12 | ||
| Entrez ID: 6351 | HGNC ID: HGNC:10630 | Ensembl Gene: ENSG00000275302 | OMIM ID: 182284 |
| Drug and gene relationship at DGIdb | |||
CCL4 involved pathways:
| KEGG pathway | Description | View |
|---|---|---|
| hsa04062 | Chemokine signaling pathway | |
| hsa04064 | NF-kappa B signaling pathway | |
| hsa04620 | Toll-like receptor signaling pathway |
Expression of CCL4:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | CCL4 | 6351 | 204103_at | 1.1304 | 0.3984 | |
| GSE20347 | CCL4 | 6351 | 204103_at | 0.1780 | 0.5451 | |
| GSE23400 | CCL4 | 6351 | 204103_at | 0.2965 | 0.0050 | |
| GSE26886 | CCL4 | 6351 | 204103_at | 0.4040 | 0.0369 | |
| GSE29001 | CCL4 | 6351 | 204103_at | 0.3796 | 0.2033 | |
| GSE38129 | CCL4 | 6351 | 204103_at | 0.2147 | 0.5119 | |
| GSE45670 | CCL4 | 6351 | 204103_at | 0.1853 | 0.6785 | |
| GSE53622 | CCL4 | 6351 | 42359 | -1.3782 | 0.0000 | |
| GSE53624 | CCL4 | 6351 | 99195 | -1.7191 | 0.0000 | |
| GSE63941 | CCL4 | 6351 | 204103_at | 0.0681 | 0.6750 | |
| GSE77861 | CCL4 | 6351 | 204103_at | 0.7045 | 0.0146 | |
| GSE97050 | CCL4 | 6351 | A_33_P3354607 | -1.9911 | 0.1806 | |
| SRP007169 | CCL4 | 6351 | RNAseq | 2.0061 | 0.0242 | |
| SRP064894 | CCL4 | 6351 | RNAseq | 1.2756 | 0.0000 | |
| SRP133303 | CCL4 | 6351 | RNAseq | 2.0014 | 0.0000 | |
| SRP159526 | CCL4 | 6351 | RNAseq | 0.7793 | 0.2952 | |
| SRP193095 | CCL4 | 6351 | RNAseq | 2.2532 | 0.0000 | |
| SRP219564 | CCL4 | 6351 | RNAseq | 1.4573 | 0.0742 | |
| TCGA | CCL4 | 6351 | RNAseq | 0.8081 | 0.0001 |
Upregulated datasets: 4; Downregulated datasets: 2.
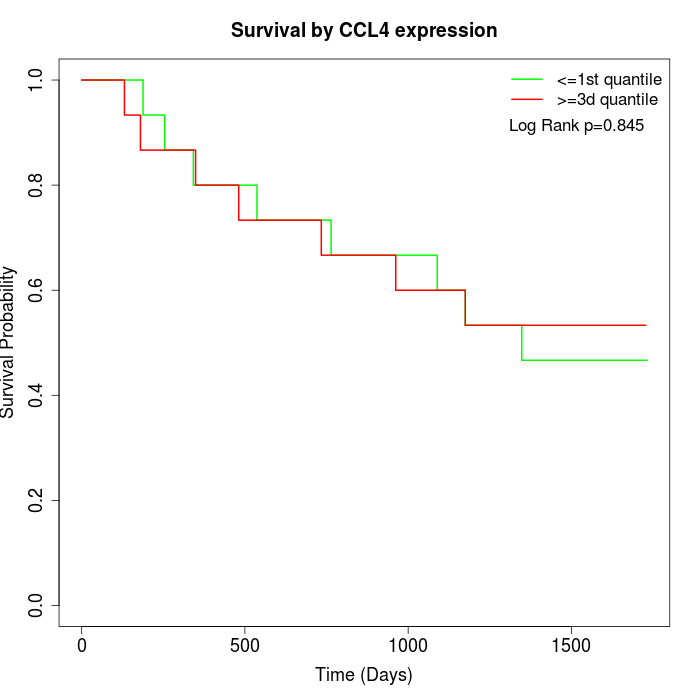
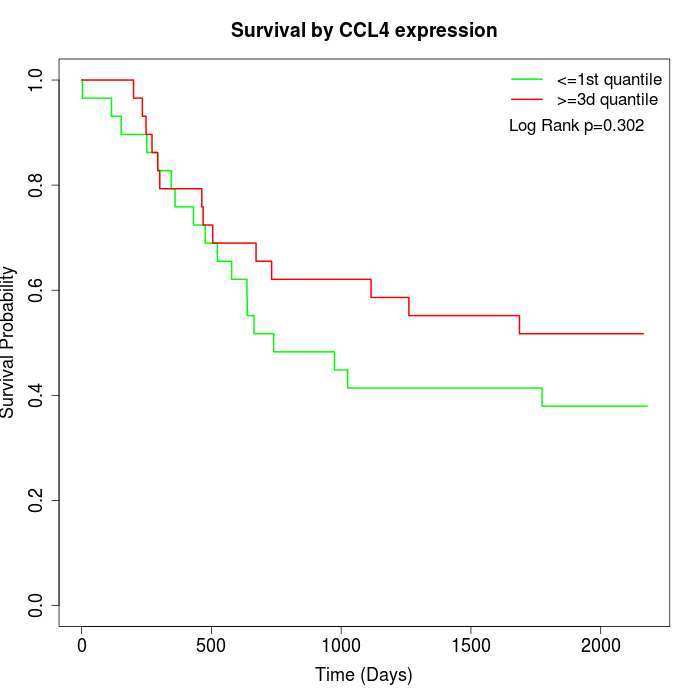
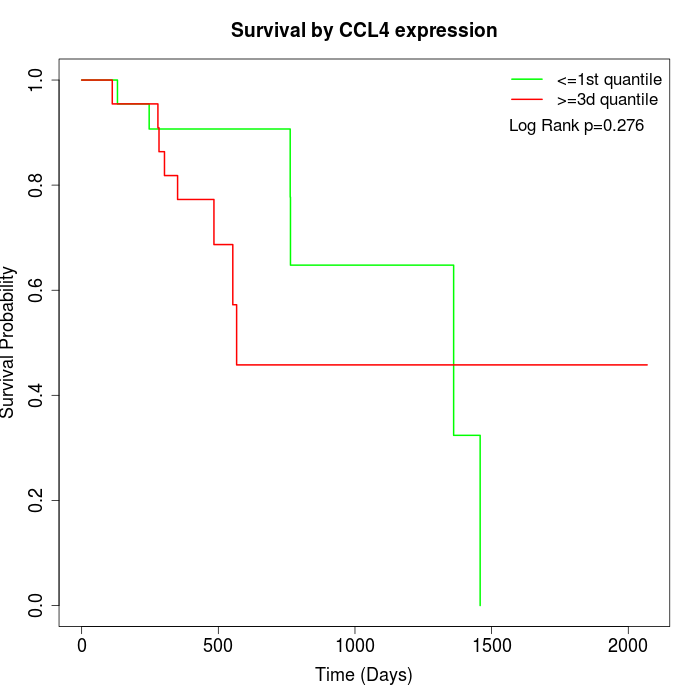
Survival by CCL4 expression:
Note: Click image to view full size file.
Copy number change of CCL4:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | CCL4 | 6351 | 5 | 1 | 24 | |
| GSE20123 | CCL4 | 6351 | 5 | 1 | 24 | |
| GSE43470 | CCL4 | 6351 | 1 | 2 | 40 | |
| GSE46452 | CCL4 | 6351 | 34 | 0 | 25 | |
| GSE47630 | CCL4 | 6351 | 7 | 1 | 32 | |
| GSE54993 | CCL4 | 6351 | 3 | 3 | 64 | |
| GSE54994 | CCL4 | 6351 | 8 | 6 | 39 | |
| GSE60625 | CCL4 | 6351 | 4 | 0 | 7 | |
| GSE74703 | CCL4 | 6351 | 1 | 1 | 34 | |
| GSE74704 | CCL4 | 6351 | 3 | 1 | 16 | |
| TCGA | CCL4 | 6351 | 20 | 9 | 67 |
Total number of gains: 91; Total number of losses: 25; Total Number of normals: 372.
Somatic mutations of CCL4:
Generating mutation plots.
Highly correlated genes for CCL4:
Showing top 20/95 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| CCL4 | TMPRSS11A | 0.690024 | 3 | 0 | 3 |
| CCL4 | C3orf20 | 0.676777 | 3 | 0 | 3 |
| CCL4 | NCCRP1 | 0.667339 | 3 | 0 | 3 |
| CCL4 | OTOF | 0.66123 | 3 | 0 | 3 |
| CCL4 | BTBD19 | 0.660782 | 4 | 0 | 4 |
| CCL4 | UPB1 | 0.657868 | 4 | 0 | 4 |
| CCL4 | TFEC | 0.643896 | 7 | 0 | 7 |
| CCL4 | FAM163B | 0.640149 | 3 | 0 | 3 |
| CCL4 | GZMB | 0.634623 | 8 | 0 | 7 |
| CCL4 | OR10G8 | 0.634435 | 3 | 0 | 3 |
| CCL4 | HYAL1 | 0.626986 | 5 | 0 | 4 |
| CCL4 | ARID5A | 0.624189 | 7 | 0 | 6 |
| CCL4 | FCGR3B | 0.622762 | 7 | 0 | 5 |
| CCL4 | MIXL1 | 0.620667 | 3 | 0 | 3 |
| CCL4 | STAT4 | 0.617737 | 7 | 0 | 7 |
| CCL4 | LAS1L | 0.616077 | 4 | 0 | 4 |
| CCL4 | RAD9A | 0.615394 | 5 | 0 | 4 |
| CCL4 | LEP | 0.615012 | 7 | 0 | 5 |
| CCL4 | CXCL11 | 0.613176 | 7 | 0 | 6 |
| CCL4 | RNF222 | 0.612949 | 3 | 0 | 3 |
For details and further investigation, click here