| Full name: caseinolytic mitochondrial matrix peptidase chaperone subunit | Alias Symbol: | ||
| Type: protein-coding gene | Cytoband: 15q22.31 | ||
| Entrez ID: 10845 | HGNC ID: HGNC:2088 | Ensembl Gene: ENSG00000166855 | OMIM ID: 615611 |
| Drug and gene relationship at DGIdb | |||
Expression of CLPX:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | CLPX | 10845 | 204809_at | -0.3681 | 0.5325 | |
| GSE20347 | CLPX | 10845 | 204809_at | -0.3754 | 0.0829 | |
| GSE23400 | CLPX | 10845 | 204809_at | -0.4227 | 0.0000 | |
| GSE26886 | CLPX | 10845 | 204809_at | -1.2462 | 0.0000 | |
| GSE29001 | CLPX | 10845 | 204809_at | -0.5864 | 0.0601 | |
| GSE38129 | CLPX | 10845 | 204809_at | -0.1185 | 0.5904 | |
| GSE45670 | CLPX | 10845 | 204809_at | -0.2138 | 0.3190 | |
| GSE53622 | CLPX | 10845 | 19176 | -0.6027 | 0.0000 | |
| GSE53624 | CLPX | 10845 | 19176 | -0.6079 | 0.0000 | |
| GSE63941 | CLPX | 10845 | 204809_at | 0.1267 | 0.7335 | |
| GSE77861 | CLPX | 10845 | 204809_at | -0.5037 | 0.2507 | |
| GSE97050 | CLPX | 10845 | A_23_P140511 | -0.0965 | 0.8012 | |
| SRP007169 | CLPX | 10845 | RNAseq | -0.7594 | 0.0220 | |
| SRP008496 | CLPX | 10845 | RNAseq | -0.7466 | 0.0010 | |
| SRP064894 | CLPX | 10845 | RNAseq | -0.8989 | 0.0000 | |
| SRP133303 | CLPX | 10845 | RNAseq | -0.3880 | 0.0017 | |
| SRP159526 | CLPX | 10845 | RNAseq | -1.0489 | 0.0017 | |
| SRP193095 | CLPX | 10845 | RNAseq | -0.7414 | 0.0000 | |
| SRP219564 | CLPX | 10845 | RNAseq | -0.8729 | 0.0469 | |
| TCGA | CLPX | 10845 | RNAseq | -0.0339 | 0.4975 |
Upregulated datasets: 0; Downregulated datasets: 2.
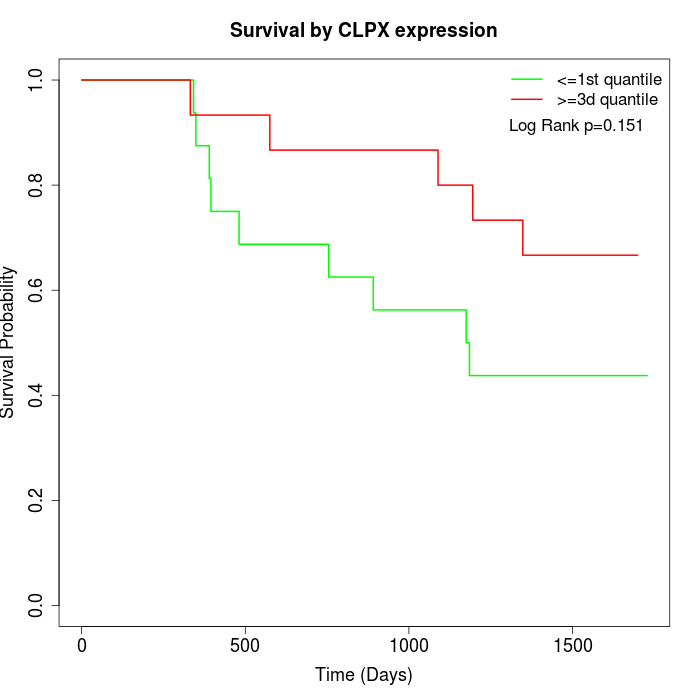
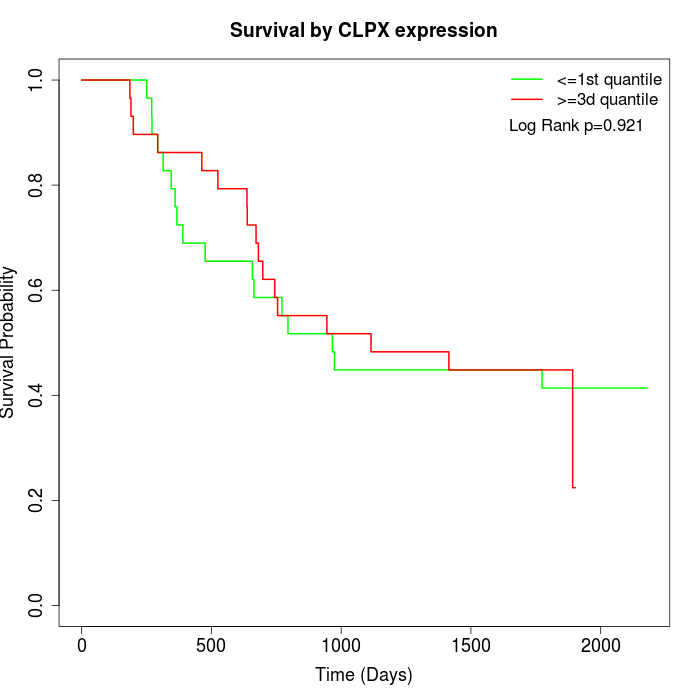
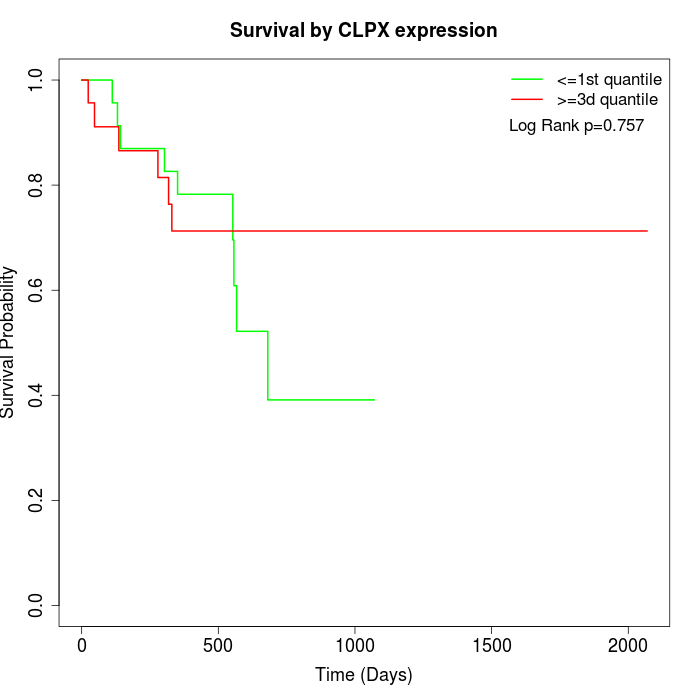
Survival by CLPX expression:
Note: Click image to view full size file.
Copy number change of CLPX:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | CLPX | 10845 | 7 | 3 | 20 | |
| GSE20123 | CLPX | 10845 | 7 | 3 | 20 | |
| GSE43470 | CLPX | 10845 | 4 | 6 | 33 | |
| GSE46452 | CLPX | 10845 | 3 | 7 | 49 | |
| GSE47630 | CLPX | 10845 | 8 | 10 | 22 | |
| GSE54993 | CLPX | 10845 | 5 | 6 | 59 | |
| GSE54994 | CLPX | 10845 | 6 | 7 | 40 | |
| GSE60625 | CLPX | 10845 | 4 | 0 | 7 | |
| GSE74703 | CLPX | 10845 | 4 | 3 | 29 | |
| GSE74704 | CLPX | 10845 | 3 | 3 | 14 | |
| TCGA | CLPX | 10845 | 10 | 16 | 70 |
Total number of gains: 61; Total number of losses: 64; Total Number of normals: 363.
Somatic mutations of CLPX:
Generating mutation plots.
Highly correlated genes for CLPX:
Showing top 20/1449 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| CLPX | CCDC25 | 0.835493 | 3 | 0 | 3 |
| CLPX | TTC39B | 0.78546 | 4 | 0 | 4 |
| CLPX | WDFY1 | 0.783571 | 3 | 0 | 3 |
| CLPX | IL34 | 0.747851 | 3 | 0 | 3 |
| CLPX | EPG5 | 0.746653 | 4 | 0 | 4 |
| CLPX | PRSS27 | 0.742271 | 6 | 0 | 6 |
| CLPX | BLOC1S2 | 0.735789 | 4 | 0 | 4 |
| CLPX | C15orf48 | 0.733009 | 6 | 0 | 6 |
| CLPX | CNFN | 0.732395 | 6 | 0 | 6 |
| CLPX | FAR1 | 0.721817 | 6 | 0 | 5 |
| CLPX | C12orf66 | 0.721542 | 3 | 0 | 3 |
| CLPX | PGM2 | 0.720539 | 8 | 0 | 8 |
| CLPX | CAPN14 | 0.720194 | 6 | 0 | 6 |
| CLPX | SLC10A7 | 0.718204 | 5 | 0 | 5 |
| CLPX | EPGN | 0.716321 | 3 | 0 | 3 |
| CLPX | LYNX1 | 0.715906 | 6 | 0 | 6 |
| CLPX | ZNF92 | 0.714872 | 3 | 0 | 3 |
| CLPX | CYSRT1 | 0.710415 | 6 | 0 | 6 |
| CLPX | SCP2 | 0.710365 | 9 | 0 | 8 |
| CLPX | HDHD2 | 0.708917 | 4 | 0 | 3 |
For details and further investigation, click here