| Full name: clavesin 1 | Alias Symbol: MGC34646|CRALBPL|C6orf212L | ||
| Type: protein-coding gene | Cytoband: 8q12.2-q12.3 | ||
| Entrez ID: 157807 | HGNC ID: HGNC:23139 | Ensembl Gene: ENSG00000177182 | OMIM ID: 611292 |
Expression of CLVS1:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | CLVS1 | 157807 | 230217_at | -0.1893 | 0.3242 | |
| GSE26886 | CLVS1 | 157807 | 230217_at | -0.2516 | 0.0714 | |
| GSE45670 | CLVS1 | 157807 | 230217_at | -0.0696 | 0.4661 | |
| GSE63941 | CLVS1 | 157807 | 230217_at | 0.2177 | 0.2405 | |
| GSE77861 | CLVS1 | 157807 | 230217_at | -0.1323 | 0.3146 | |
| SRP007169 | CLVS1 | 157807 | RNAseq | -7.1905 | 0.0000 | |
| SRP064894 | CLVS1 | 157807 | RNAseq | -2.8206 | 0.0000 | |
| SRP133303 | CLVS1 | 157807 | RNAseq | -1.2969 | 0.0428 | |
| TCGA | CLVS1 | 157807 | RNAseq | -0.0603 | 0.8703 |
Upregulated datasets: 0; Downregulated datasets: 3.
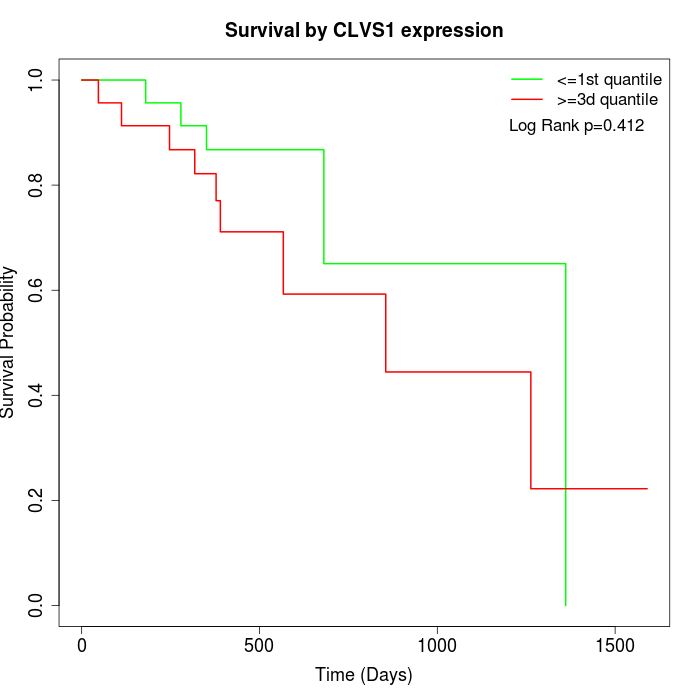
Survival by CLVS1 expression:
Note: Click image to view full size file.
Copy number change of CLVS1:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | CLVS1 | 157807 | 13 | 1 | 16 | |
| GSE20123 | CLVS1 | 157807 | 13 | 1 | 16 | |
| GSE43470 | CLVS1 | 157807 | 14 | 3 | 26 | |
| GSE46452 | CLVS1 | 157807 | 20 | 2 | 37 | |
| GSE47630 | CLVS1 | 157807 | 23 | 1 | 16 | |
| GSE54993 | CLVS1 | 157807 | 1 | 18 | 51 | |
| GSE54994 | CLVS1 | 157807 | 30 | 0 | 23 | |
| GSE60625 | CLVS1 | 157807 | 0 | 4 | 7 | |
| GSE74703 | CLVS1 | 157807 | 12 | 2 | 22 | |
| GSE74704 | CLVS1 | 157807 | 9 | 0 | 11 | |
| TCGA | CLVS1 | 157807 | 45 | 6 | 45 |
Total number of gains: 180; Total number of losses: 38; Total Number of normals: 270.
Somatic mutations of CLVS1:
Generating mutation plots.
Highly correlated genes for CLVS1:
Showing top 20/101 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| CLVS1 | CRB3 | 0.714084 | 3 | 0 | 3 |
| CLVS1 | ATP6V0C | 0.698473 | 3 | 0 | 3 |
| CLVS1 | ZNF135 | 0.693755 | 3 | 0 | 3 |
| CLVS1 | NAPA | 0.691668 | 3 | 0 | 3 |
| CLVS1 | TJP2 | 0.688364 | 3 | 0 | 3 |
| CLVS1 | NDUFS7 | 0.686583 | 4 | 0 | 4 |
| CLVS1 | LINC01134 | 0.660282 | 3 | 0 | 3 |
| CLVS1 | CLDN17 | 0.658158 | 3 | 0 | 3 |
| CLVS1 | GPR82 | 0.65624 | 3 | 0 | 3 |
| CLVS1 | SEC14L5 | 0.652916 | 3 | 0 | 3 |
| CLVS1 | LCN8 | 0.651006 | 3 | 0 | 3 |
| CLVS1 | PHF7 | 0.648117 | 4 | 0 | 3 |
| CLVS1 | C10orf67 | 0.647484 | 3 | 0 | 3 |
| CLVS1 | MYBPC3 | 0.640582 | 3 | 0 | 3 |
| CLVS1 | KIF25 | 0.63868 | 4 | 0 | 3 |
| CLVS1 | KRT36 | 0.638638 | 3 | 0 | 3 |
| CLVS1 | ALKBH7 | 0.635272 | 3 | 0 | 3 |
| CLVS1 | GOT1 | 0.633452 | 3 | 0 | 3 |
| CLVS1 | FAM71C | 0.633029 | 3 | 0 | 3 |
| CLVS1 | ANKRD30B | 0.627846 | 4 | 0 | 3 |
For details and further investigation, click here