| Full name: cornichon family AMPA receptor auxiliary protein 1 | Alias Symbol: TGAM77|CNIL | ||
| Type: protein-coding gene | Cytoband: 14q22.2 | ||
| Entrez ID: 10175 | HGNC ID: HGNC:19431 | Ensembl Gene: ENSG00000100528 | OMIM ID: 611287 |
| Drug and gene relationship at DGIdb | |||
Expression of CNIH1:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | CNIH1 | 10175 | 201653_at | 0.0500 | 0.9348 | |
| GSE20347 | CNIH1 | 10175 | 201653_at | 0.1276 | 0.4136 | |
| GSE23400 | CNIH1 | 10175 | 201653_at | 0.4825 | 0.0000 | |
| GSE26886 | CNIH1 | 10175 | 201653_at | -0.0492 | 0.8301 | |
| GSE29001 | CNIH1 | 10175 | 201653_at | 0.1971 | 0.6004 | |
| GSE38129 | CNIH1 | 10175 | 201653_at | 0.2396 | 0.0444 | |
| GSE45670 | CNIH1 | 10175 | 201653_at | 0.1971 | 0.2668 | |
| GSE53622 | CNIH1 | 10175 | 30376 | 0.1601 | 0.0519 | |
| GSE53624 | CNIH1 | 10175 | 30376 | 0.2022 | 0.0077 | |
| GSE63941 | CNIH1 | 10175 | 201653_at | -0.0281 | 0.9284 | |
| GSE77861 | CNIH1 | 10175 | 201653_at | 0.0774 | 0.8738 | |
| SRP007169 | CNIH1 | 10175 | RNAseq | -0.1010 | 0.7423 | |
| SRP008496 | CNIH1 | 10175 | RNAseq | -0.2065 | 0.2683 | |
| SRP064894 | CNIH1 | 10175 | RNAseq | 0.1062 | 0.4657 | |
| SRP133303 | CNIH1 | 10175 | RNAseq | 0.3381 | 0.0702 | |
| SRP159526 | CNIH1 | 10175 | RNAseq | 0.0199 | 0.9341 | |
| SRP193095 | CNIH1 | 10175 | RNAseq | -0.1978 | 0.1800 | |
| SRP219564 | CNIH1 | 10175 | RNAseq | 0.0411 | 0.8890 |
Upregulated datasets: 0; Downregulated datasets: 0.
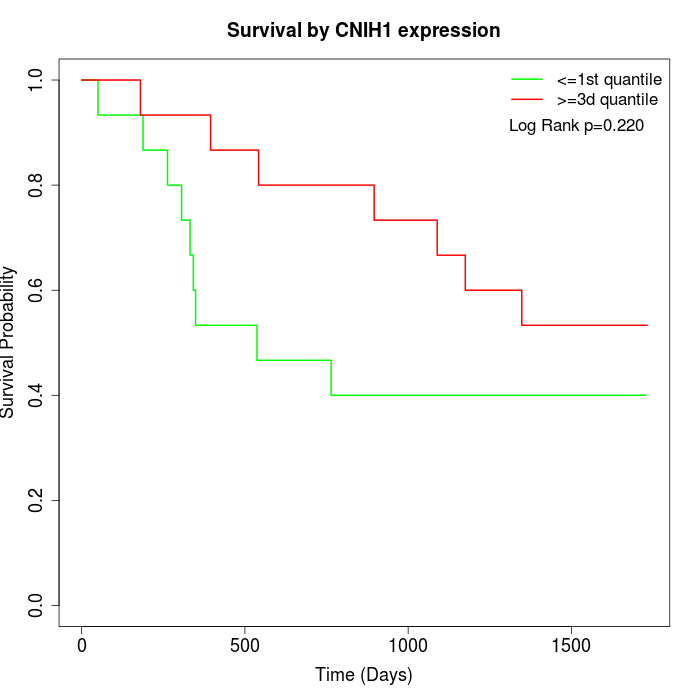
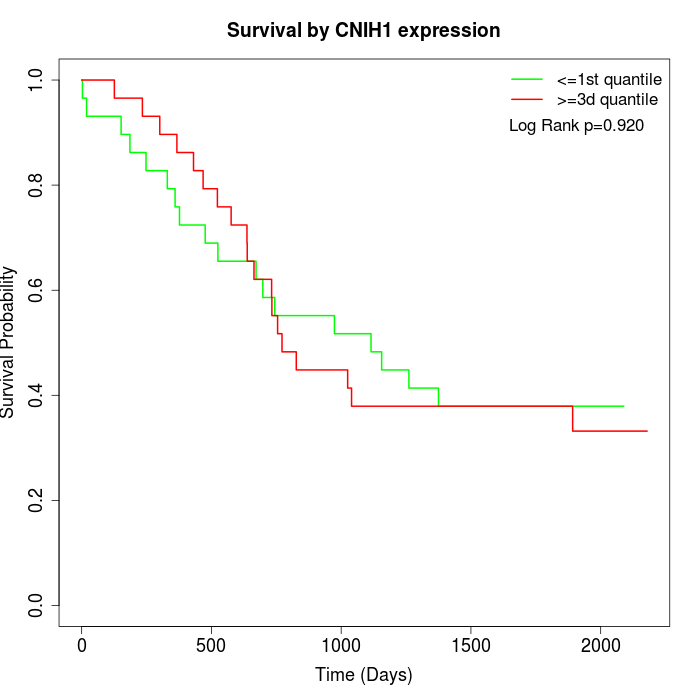
Survival by CNIH1 expression:
Note: Click image to view full size file.
Copy number change of CNIH1:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | CNIH1 | 10175 | 8 | 3 | 19 | |
| GSE20123 | CNIH1 | 10175 | 8 | 3 | 19 | |
| GSE43470 | CNIH1 | 10175 | 9 | 1 | 33 | |
| GSE46452 | CNIH1 | 10175 | 16 | 3 | 40 | |
| GSE47630 | CNIH1 | 10175 | 11 | 10 | 19 | |
| GSE54993 | CNIH1 | 10175 | 3 | 8 | 59 | |
| GSE54994 | CNIH1 | 10175 | 19 | 4 | 30 | |
| GSE60625 | CNIH1 | 10175 | 0 | 2 | 9 | |
| GSE74703 | CNIH1 | 10175 | 8 | 1 | 27 | |
| GSE74704 | CNIH1 | 10175 | 3 | 2 | 15 |
Total number of gains: 85; Total number of losses: 37; Total Number of normals: 270.
Somatic mutations of CNIH1:
Generating mutation plots.
Highly correlated genes for CNIH1:
Showing top 20/255 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| CNIH1 | WDR36 | 0.728632 | 3 | 0 | 3 |
| CNIH1 | USP1 | 0.722808 | 4 | 0 | 4 |
| CNIH1 | TMX1 | 0.700291 | 11 | 0 | 11 |
| CNIH1 | ASCC3 | 0.699185 | 4 | 0 | 4 |
| CNIH1 | MRPL13 | 0.667172 | 5 | 0 | 4 |
| CNIH1 | RDH11 | 0.663326 | 10 | 0 | 9 |
| CNIH1 | INTS12 | 0.658208 | 3 | 0 | 3 |
| CNIH1 | KTN1 | 0.655798 | 8 | 0 | 7 |
| CNIH1 | DUSP11 | 0.653453 | 4 | 0 | 4 |
| CNIH1 | MMGT1 | 0.653238 | 3 | 0 | 3 |
| CNIH1 | POLR2K | 0.647732 | 4 | 0 | 3 |
| CNIH1 | GMFB | 0.643447 | 9 | 0 | 7 |
| CNIH1 | NAA30 | 0.638657 | 6 | 0 | 4 |
| CNIH1 | COX16 | 0.637432 | 4 | 0 | 3 |
| CNIH1 | PRDX4 | 0.635936 | 3 | 0 | 3 |
| CNIH1 | JKAMP | 0.635121 | 7 | 0 | 6 |
| CNIH1 | MED6 | 0.634822 | 9 | 0 | 8 |
| CNIH1 | ASCC1 | 0.632103 | 4 | 0 | 3 |
| CNIH1 | PNO1 | 0.63021 | 3 | 0 | 3 |
| CNIH1 | ILF2 | 0.629899 | 4 | 0 | 4 |
For details and further investigation, click here