| Full name: free fatty acid receptor 2 | Alias Symbol: FFA2R | ||
| Type: protein-coding gene | Cytoband: 19q13.12 | ||
| Entrez ID: 2867 | HGNC ID: HGNC:4501 | Ensembl Gene: ENSG00000126262 | OMIM ID: 603823 |
| Related drugs: ACETIC ACID, BUTANOIC ACID, CHEMBL52416, ISOBUTYRIC ACID, PROPIONIC ACID, TIRATRICOL, VALERIC ACID... [more] | |||
FFAR2 involved pathways:
| KEGG pathway | Description | View |
|---|---|---|
| hsa04024 | cAMP signaling pathway |
Expression of FFAR2:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | FFAR2 | 2867 | 221345_at | 0.5078 | 0.6188 | |
| GSE20347 | FFAR2 | 2867 | 221345_at | 0.0207 | 0.8583 | |
| GSE23400 | FFAR2 | 2867 | 221345_at | -0.1162 | 0.0376 | |
| GSE26886 | FFAR2 | 2867 | 221345_at | -0.2917 | 0.0200 | |
| GSE29001 | FFAR2 | 2867 | 221345_at | 0.0174 | 0.9132 | |
| GSE38129 | FFAR2 | 2867 | 221345_at | 0.0453 | 0.6663 | |
| GSE45670 | FFAR2 | 2867 | 221345_at | 0.6542 | 0.0128 | |
| GSE53622 | FFAR2 | 2867 | 15220 | 0.4219 | 0.0000 | |
| GSE53624 | FFAR2 | 2867 | 15220 | 0.0888 | 0.3001 | |
| GSE63941 | FFAR2 | 2867 | 221345_at | 0.2005 | 0.5553 | |
| GSE77861 | FFAR2 | 2867 | 221345_at | -0.1283 | 0.2274 | |
| GSE97050 | FFAR2 | 2867 | A_23_P397391 | 0.3252 | 0.1657 | |
| SRP007169 | FFAR2 | 2867 | RNAseq | -0.0841 | 0.8975 | |
| SRP064894 | FFAR2 | 2867 | RNAseq | 0.7259 | 0.0490 | |
| SRP133303 | FFAR2 | 2867 | RNAseq | 0.8471 | 0.0000 | |
| SRP159526 | FFAR2 | 2867 | RNAseq | -0.1622 | 0.8434 | |
| SRP193095 | FFAR2 | 2867 | RNAseq | 1.1162 | 0.0000 | |
| SRP219564 | FFAR2 | 2867 | RNAseq | -0.6942 | 0.1399 | |
| TCGA | FFAR2 | 2867 | RNAseq | 0.8679 | 0.0161 |
Upregulated datasets: 1; Downregulated datasets: 0.
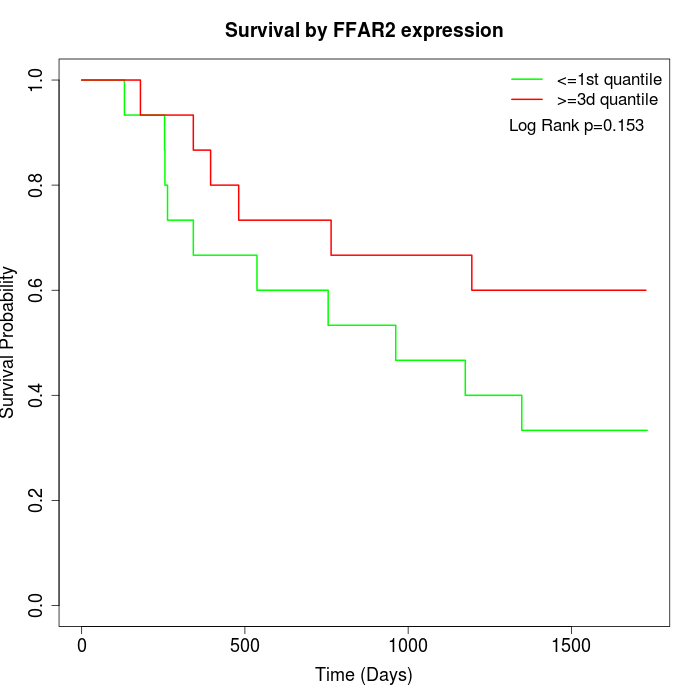
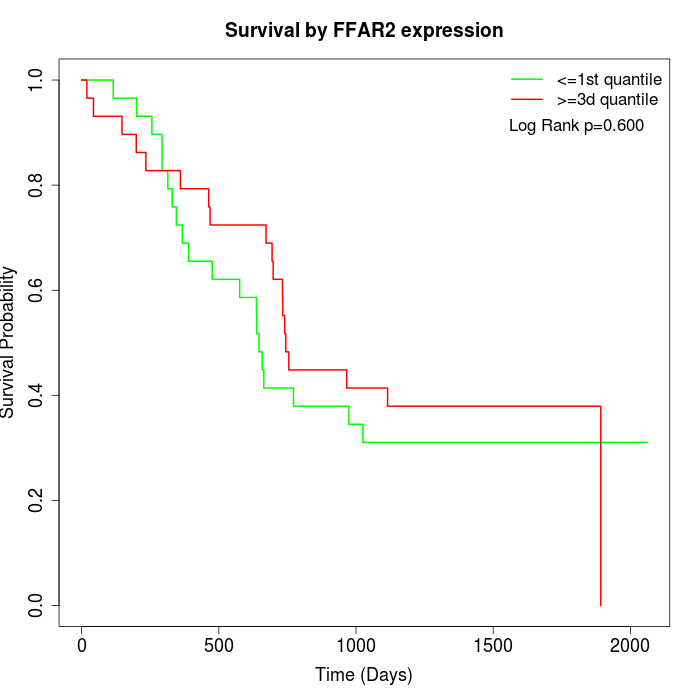
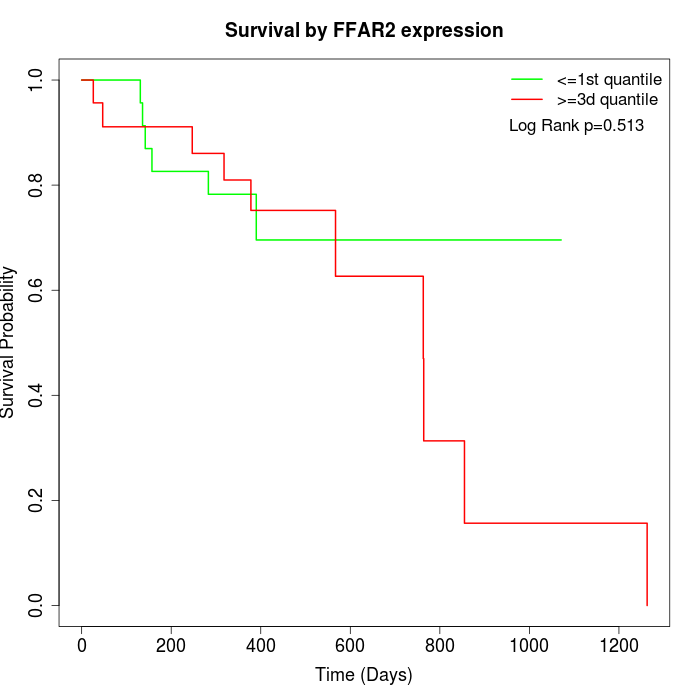
Survival by FFAR2 expression:
Note: Click image to view full size file.
Copy number change of FFAR2:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | FFAR2 | 2867 | 6 | 4 | 20 | |
| GSE20123 | FFAR2 | 2867 | 6 | 3 | 21 | |
| GSE43470 | FFAR2 | 2867 | 3 | 7 | 33 | |
| GSE46452 | FFAR2 | 2867 | 48 | 1 | 10 | |
| GSE47630 | FFAR2 | 2867 | 9 | 5 | 26 | |
| GSE54993 | FFAR2 | 2867 | 17 | 3 | 50 | |
| GSE54994 | FFAR2 | 2867 | 8 | 9 | 36 | |
| GSE60625 | FFAR2 | 2867 | 9 | 0 | 2 | |
| GSE74703 | FFAR2 | 2867 | 3 | 4 | 29 | |
| GSE74704 | FFAR2 | 2867 | 6 | 1 | 13 | |
| TCGA | FFAR2 | 2867 | 21 | 11 | 64 |
Total number of gains: 136; Total number of losses: 48; Total Number of normals: 304.
Somatic mutations of FFAR2:
Generating mutation plots.
Highly correlated genes for FFAR2:
Showing top 20/297 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| FFAR2 | ARL4D | 0.777065 | 3 | 0 | 3 |
| FFAR2 | LDHC | 0.764362 | 3 | 0 | 3 |
| FFAR2 | MPP2 | 0.751105 | 3 | 0 | 3 |
| FFAR2 | NHLRC4 | 0.738087 | 3 | 0 | 3 |
| FFAR2 | STK32C | 0.731407 | 3 | 0 | 3 |
| FFAR2 | LHFPL4 | 0.723869 | 3 | 0 | 3 |
| FFAR2 | GJD4 | 0.710602 | 4 | 0 | 4 |
| FFAR2 | CRYBB3 | 0.710395 | 3 | 0 | 3 |
| FFAR2 | LIMD1 | 0.709339 | 3 | 0 | 3 |
| FFAR2 | LDHAL6A | 0.706304 | 3 | 0 | 3 |
| FFAR2 | ANKRD33 | 0.705208 | 3 | 0 | 3 |
| FFAR2 | ACSBG2 | 0.702034 | 3 | 0 | 3 |
| FFAR2 | FCN2 | 0.700892 | 3 | 0 | 3 |
| FFAR2 | SFMBT2 | 0.700465 | 3 | 0 | 3 |
| FFAR2 | KCNF1 | 0.698707 | 3 | 0 | 3 |
| FFAR2 | LCN8 | 0.697959 | 4 | 0 | 3 |
| FFAR2 | TRAPPC9 | 0.689597 | 4 | 0 | 4 |
| FFAR2 | IL1F10 | 0.688402 | 4 | 0 | 3 |
| FFAR2 | TP73 | 0.684682 | 4 | 0 | 3 |
| FFAR2 | MCEMP1 | 0.680691 | 5 | 0 | 4 |
For details and further investigation, click here