| Full name: hypocretin receptor 2 | Alias Symbol: OX2R | ||
| Type: protein-coding gene | Cytoband: 6p12.1 | ||
| Entrez ID: 3062 | HGNC ID: HGNC:4849 | Ensembl Gene: ENSG00000137252 | OMIM ID: 602393 |
| Related drugs: ALMOREXANT, LEMBOREXANT, SB-649868, SUVOREXANT... [more] | |||
Expression of HCRTR2:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | HCRTR2 | 3062 | 207393_at | 0.0108 | 0.9766 | |
| GSE20347 | HCRTR2 | 3062 | 207393_at | 0.0133 | 0.8473 | |
| GSE23400 | HCRTR2 | 3062 | 207393_at | -0.1299 | 0.0001 | |
| GSE26886 | HCRTR2 | 3062 | 207393_at | -0.2283 | 0.0614 | |
| GSE29001 | HCRTR2 | 3062 | 207393_at | -0.1102 | 0.3802 | |
| GSE38129 | HCRTR2 | 3062 | 207393_at | -0.0597 | 0.3127 | |
| GSE45670 | HCRTR2 | 3062 | 207393_at | 0.0781 | 0.3772 | |
| GSE53622 | HCRTR2 | 3062 | 7575 | -0.1747 | 0.1916 | |
| GSE53624 | HCRTR2 | 3062 | 7575 | 0.1386 | 0.0773 | |
| GSE63941 | HCRTR2 | 3062 | 207393_at | 0.0199 | 0.9124 | |
| GSE77861 | HCRTR2 | 3062 | 207393_at | 0.0021 | 0.9822 | |
| TCGA | HCRTR2 | 3062 | RNAseq | 1.6422 | 0.2374 |
Upregulated datasets: 0; Downregulated datasets: 0.
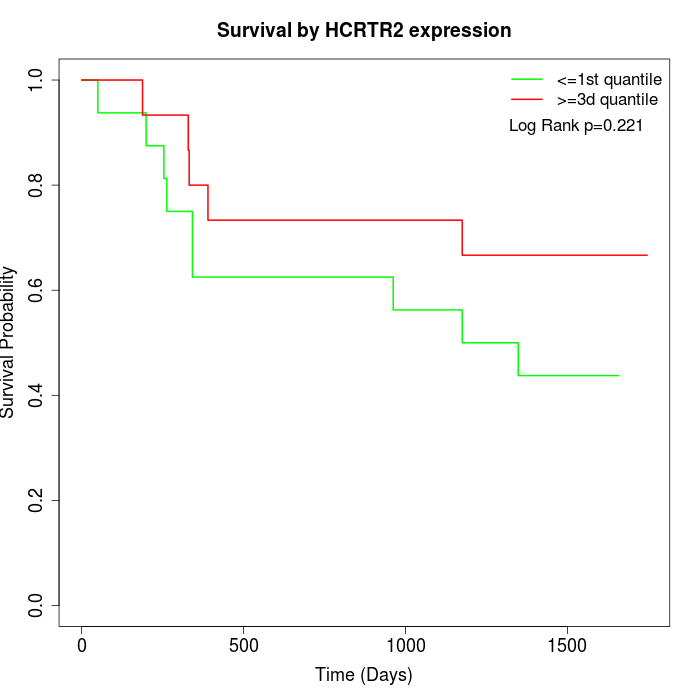
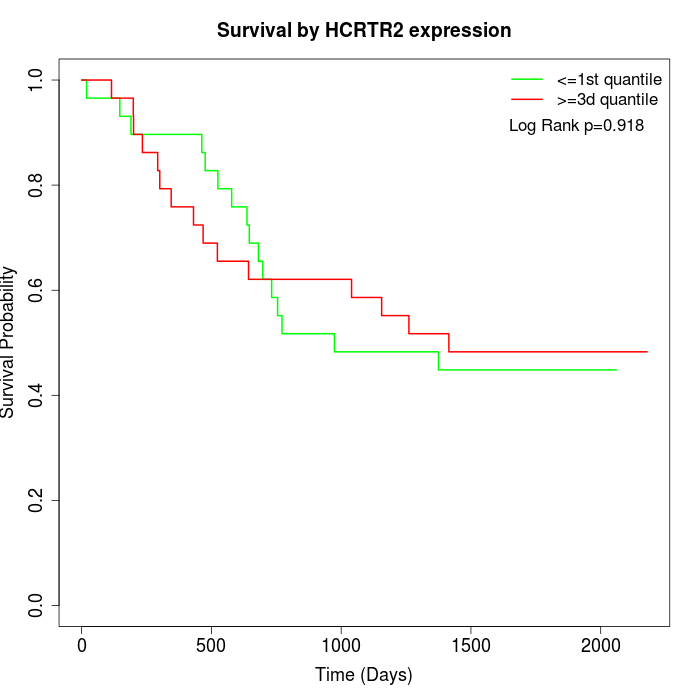
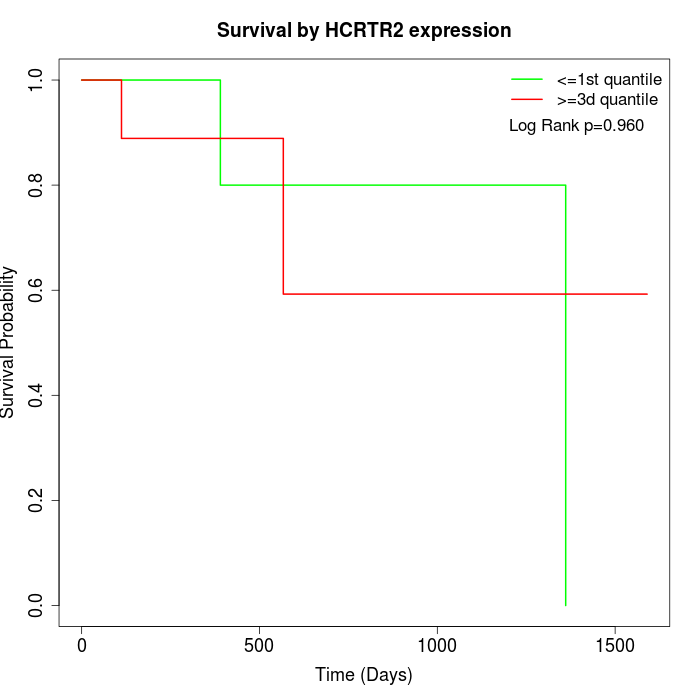
Survival by HCRTR2 expression:
Note: Click image to view full size file.
Copy number change of HCRTR2:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | HCRTR2 | 3062 | 8 | 1 | 21 | |
| GSE20123 | HCRTR2 | 3062 | 8 | 1 | 21 | |
| GSE43470 | HCRTR2 | 3062 | 5 | 0 | 38 | |
| GSE46452 | HCRTR2 | 3062 | 2 | 10 | 47 | |
| GSE47630 | HCRTR2 | 3062 | 7 | 5 | 28 | |
| GSE54993 | HCRTR2 | 3062 | 3 | 2 | 65 | |
| GSE54994 | HCRTR2 | 3062 | 8 | 4 | 41 | |
| GSE60625 | HCRTR2 | 3062 | 0 | 3 | 8 | |
| GSE74703 | HCRTR2 | 3062 | 5 | 0 | 31 | |
| GSE74704 | HCRTR2 | 3062 | 4 | 0 | 16 | |
| TCGA | HCRTR2 | 3062 | 23 | 12 | 61 |
Total number of gains: 73; Total number of losses: 38; Total Number of normals: 377.
Somatic mutations of HCRTR2:
Generating mutation plots.
Highly correlated genes for HCRTR2:
Showing top 20/304 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| HCRTR2 | CA8 | 0.711187 | 3 | 0 | 3 |
| HCRTR2 | SEC14L3 | 0.674325 | 3 | 0 | 3 |
| HCRTR2 | FOXN3-AS2 | 0.667768 | 4 | 0 | 4 |
| HCRTR2 | NPAP1 | 0.664486 | 3 | 0 | 3 |
| HCRTR2 | GLP1R | 0.653715 | 4 | 0 | 3 |
| HCRTR2 | NPHS2 | 0.648994 | 4 | 0 | 3 |
| HCRTR2 | SLC28A2 | 0.646209 | 3 | 0 | 3 |
| HCRTR2 | FGA | 0.643865 | 3 | 0 | 3 |
| HCRTR2 | TAC3 | 0.632454 | 4 | 0 | 3 |
| HCRTR2 | ADRA1A | 0.630964 | 3 | 0 | 3 |
| HCRTR2 | TNFSF11 | 0.628953 | 4 | 0 | 3 |
| HCRTR2 | ART1 | 0.628145 | 6 | 0 | 4 |
| HCRTR2 | MATN1 | 0.62789 | 4 | 0 | 3 |
| HCRTR2 | MMP24 | 0.623875 | 4 | 0 | 4 |
| HCRTR2 | GREB1L | 0.62272 | 4 | 0 | 4 |
| HCRTR2 | CCR4 | 0.621852 | 3 | 0 | 3 |
| HCRTR2 | CCL1 | 0.621533 | 4 | 0 | 3 |
| HCRTR2 | ZNF549 | 0.620793 | 3 | 0 | 3 |
| HCRTR2 | CPS1-IT1 | 0.620751 | 3 | 0 | 3 |
| HCRTR2 | NOTUM | 0.618293 | 3 | 0 | 3 |
For details and further investigation, click here