| Full name: intraflagellar transport 57 | Alias Symbol: FLJ10147|HIPPI|MHS4R2 | ||
| Type: protein-coding gene | Cytoband: 3q13.12-q13.13 | ||
| Entrez ID: 55081 | HGNC ID: HGNC:17367 | Ensembl Gene: ENSG00000114446 | OMIM ID: 606621 |
Expression of IFT57:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | IFT57 | 55081 | 218100_s_at | -0.0780 | 0.8421 | |
| GSE20347 | IFT57 | 55081 | 218100_s_at | 0.2541 | 0.3806 | |
| GSE23400 | IFT57 | 55081 | 218100_s_at | 0.3952 | 0.0000 | |
| GSE26886 | IFT57 | 55081 | 218100_s_at | 1.2127 | 0.0000 | |
| GSE29001 | IFT57 | 55081 | 218100_s_at | 0.0491 | 0.9328 | |
| GSE38129 | IFT57 | 55081 | 218100_s_at | 0.1964 | 0.3227 | |
| GSE45670 | IFT57 | 55081 | 218100_s_at | -0.0223 | 0.9311 | |
| GSE53622 | IFT57 | 55081 | 41086 | -1.1465 | 0.0000 | |
| GSE53624 | IFT57 | 55081 | 41086 | -1.0492 | 0.0000 | |
| GSE63941 | IFT57 | 55081 | 218100_s_at | -0.6896 | 0.3540 | |
| GSE77861 | IFT57 | 55081 | 218100_s_at | -0.0525 | 0.8797 | |
| GSE97050 | IFT57 | 55081 | A_24_P362881 | -0.3871 | 0.2041 | |
| SRP007169 | IFT57 | 55081 | RNAseq | 0.7394 | 0.2587 | |
| SRP008496 | IFT57 | 55081 | RNAseq | 1.6859 | 0.0005 | |
| SRP064894 | IFT57 | 55081 | RNAseq | 0.1240 | 0.4126 | |
| SRP133303 | IFT57 | 55081 | RNAseq | 0.1413 | 0.4727 | |
| SRP159526 | IFT57 | 55081 | RNAseq | 0.4429 | 0.0682 | |
| SRP193095 | IFT57 | 55081 | RNAseq | 0.2654 | 0.0433 | |
| SRP219564 | IFT57 | 55081 | RNAseq | -0.0025 | 0.9953 | |
| TCGA | IFT57 | 55081 | RNAseq | -0.0447 | 0.4546 |
Upregulated datasets: 2; Downregulated datasets: 2.
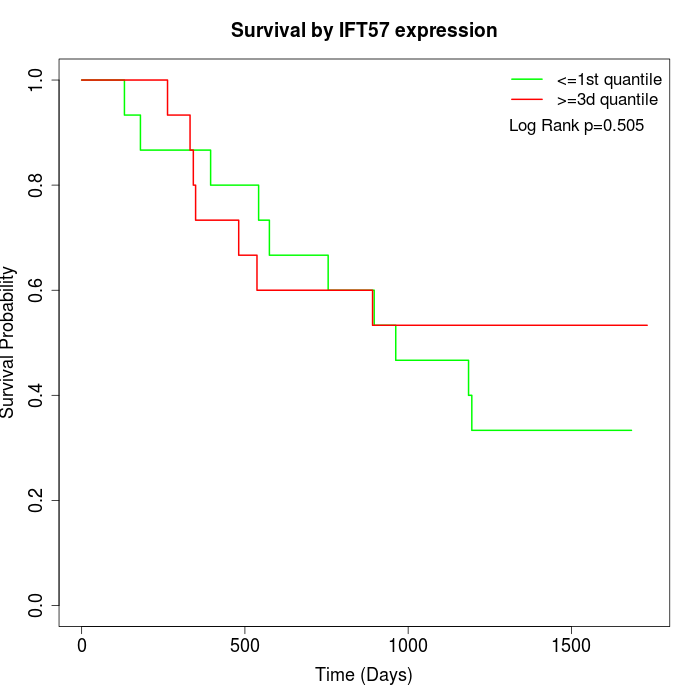
Survival by IFT57 expression:
Note: Click image to view full size file.
Copy number change of IFT57:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | IFT57 | 55081 | 18 | 1 | 11 | |
| GSE20123 | IFT57 | 55081 | 17 | 1 | 12 | |
| GSE43470 | IFT57 | 55081 | 18 | 1 | 24 | |
| GSE46452 | IFT57 | 55081 | 12 | 6 | 41 | |
| GSE47630 | IFT57 | 55081 | 16 | 6 | 18 | |
| GSE54993 | IFT57 | 55081 | 2 | 5 | 63 | |
| GSE54994 | IFT57 | 55081 | 28 | 5 | 20 | |
| GSE60625 | IFT57 | 55081 | 0 | 6 | 5 | |
| GSE74703 | IFT57 | 55081 | 15 | 1 | 20 | |
| GSE74704 | IFT57 | 55081 | 14 | 1 | 5 | |
| TCGA | IFT57 | 55081 | 50 | 8 | 38 |
Total number of gains: 190; Total number of losses: 41; Total Number of normals: 257.
Somatic mutations of IFT57:
Generating mutation plots.
Highly correlated genes for IFT57:
Showing top 20/325 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| IFT57 | CCL14 | 0.718905 | 3 | 0 | 3 |
| IFT57 | RCAN2 | 0.716195 | 3 | 0 | 3 |
| IFT57 | ARL6 | 0.701053 | 3 | 0 | 3 |
| IFT57 | CLEC3B | 0.695185 | 3 | 0 | 3 |
| IFT57 | SHPRH | 0.685108 | 3 | 0 | 3 |
| IFT57 | PYGM | 0.667358 | 3 | 0 | 3 |
| IFT57 | RERGL | 0.667076 | 3 | 0 | 3 |
| IFT57 | SPARCL1 | 0.664266 | 3 | 0 | 3 |
| IFT57 | RGN | 0.660529 | 3 | 0 | 3 |
| IFT57 | RFX7 | 0.660021 | 3 | 0 | 3 |
| IFT57 | DYRK1A | 0.657978 | 3 | 0 | 3 |
| IFT57 | HEXA | 0.656219 | 3 | 0 | 3 |
| IFT57 | TMF1 | 0.655543 | 4 | 0 | 4 |
| IFT57 | ZNF616 | 0.655058 | 4 | 0 | 3 |
| IFT57 | KIF9 | 0.654889 | 5 | 0 | 3 |
| IFT57 | WHAMM | 0.652687 | 4 | 0 | 3 |
| IFT57 | PPIL6 | 0.651616 | 4 | 0 | 4 |
| IFT57 | FDX1 | 0.650384 | 3 | 0 | 3 |
| IFT57 | ITIH5 | 0.646515 | 3 | 0 | 3 |
| IFT57 | USP9X | 0.644928 | 6 | 0 | 5 |
For details and further investigation, click here