| Full name: inhibin subunit beta E | Alias Symbol: activin|MGC4638 | ||
| Type: protein-coding gene | Cytoband: 12q13.3 | ||
| Entrez ID: 83729 | HGNC ID: HGNC:24029 | Ensembl Gene: ENSG00000139269 | OMIM ID: 612031 |
| Related drugs: GONADORELIN ACETATE... [more] | |||
INHBE involved pathways:
| KEGG pathway | Description | View |
|---|---|---|
| hsa04350 | TGF-beta signaling pathway | |
| hsa04550 | Signaling pathways regulating pluripotency of stem cells |
Expression of INHBE:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | INHBE | 83729 | 210587_at | 0.1415 | 0.6265 | |
| GSE20347 | INHBE | 83729 | 210587_at | 0.1601 | 0.0236 | |
| GSE23400 | INHBE | 83729 | 210587_at | 0.0198 | 0.5139 | |
| GSE26886 | INHBE | 83729 | 210587_at | 0.1030 | 0.3718 | |
| GSE29001 | INHBE | 83729 | 210587_at | 0.4105 | 0.2789 | |
| GSE38129 | INHBE | 83729 | 210587_at | 0.2664 | 0.1828 | |
| GSE45670 | INHBE | 83729 | 210587_at | 0.4077 | 0.3809 | |
| GSE53622 | INHBE | 83729 | 62709 | 0.5959 | 0.0001 | |
| GSE53624 | INHBE | 83729 | 62709 | 0.8702 | 0.0000 | |
| GSE63941 | INHBE | 83729 | 210587_at | 2.1738 | 0.3190 | |
| GSE77861 | INHBE | 83729 | 210587_at | -0.0661 | 0.6027 | |
| GSE97050 | INHBE | 83729 | A_24_P141707 | 0.0937 | 0.6481 | |
| SRP133303 | INHBE | 83729 | RNAseq | 0.2296 | 0.4170 | |
| SRP159526 | INHBE | 83729 | RNAseq | 0.9040 | 0.2497 | |
| TCGA | INHBE | 83729 | RNAseq | 2.7868 | 0.0000 |
Upregulated datasets: 1; Downregulated datasets: 0.
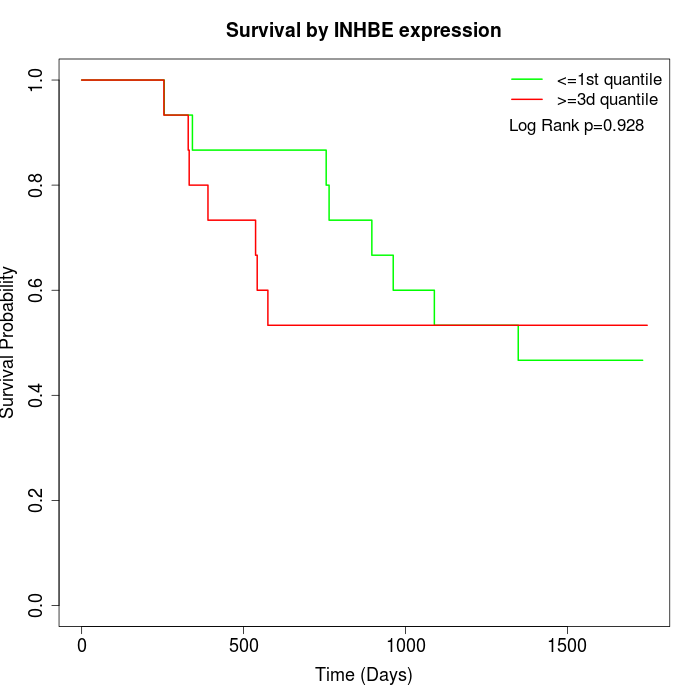
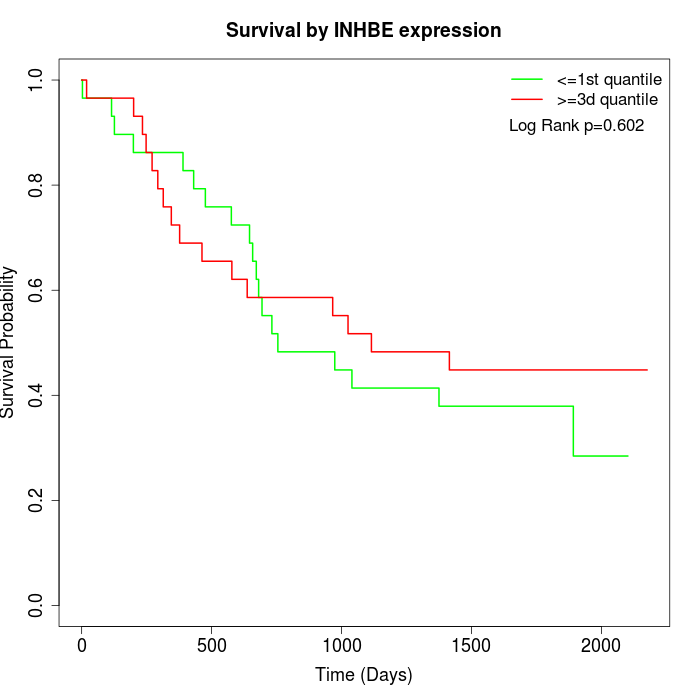
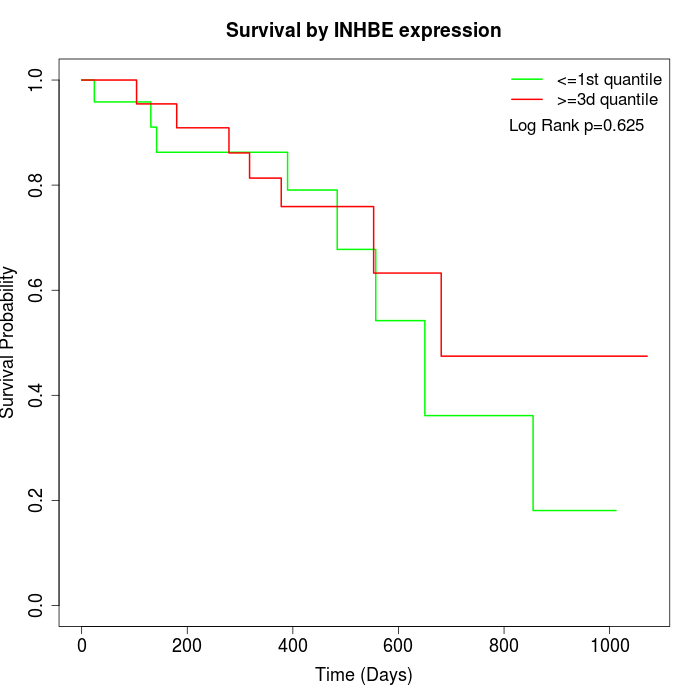
Survival by INHBE expression:
Note: Click image to view full size file.
Copy number change of INHBE:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | INHBE | 83729 | 6 | 1 | 23 | |
| GSE20123 | INHBE | 83729 | 6 | 0 | 24 | |
| GSE43470 | INHBE | 83729 | 2 | 0 | 41 | |
| GSE46452 | INHBE | 83729 | 8 | 1 | 50 | |
| GSE47630 | INHBE | 83729 | 10 | 2 | 28 | |
| GSE54993 | INHBE | 83729 | 0 | 5 | 65 | |
| GSE54994 | INHBE | 83729 | 4 | 1 | 48 | |
| GSE60625 | INHBE | 83729 | 0 | 0 | 11 | |
| GSE74703 | INHBE | 83729 | 2 | 0 | 34 | |
| GSE74704 | INHBE | 83729 | 4 | 0 | 16 | |
| TCGA | INHBE | 83729 | 14 | 10 | 72 |
Total number of gains: 56; Total number of losses: 20; Total Number of normals: 412.
Somatic mutations of INHBE:
Generating mutation plots.
Highly correlated genes for INHBE:
Showing top 20/90 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| INHBE | PAPPA | 0.715615 | 3 | 0 | 3 |
| INHBE | EIF3J | 0.712974 | 3 | 0 | 3 |
| INHBE | FOXS1 | 0.70287 | 3 | 0 | 3 |
| INHBE | PNO1 | 0.702798 | 4 | 0 | 4 |
| INHBE | ACP1 | 0.688249 | 3 | 0 | 3 |
| INHBE | MYO3B | 0.687928 | 3 | 0 | 3 |
| INHBE | PIGH | 0.668474 | 3 | 0 | 3 |
| INHBE | FANCM | 0.656227 | 3 | 0 | 3 |
| INHBE | ELK4 | 0.648605 | 3 | 0 | 3 |
| INHBE | E2F5 | 0.646324 | 4 | 0 | 4 |
| INHBE | CCR10 | 0.643372 | 4 | 0 | 3 |
| INHBE | MYADML2 | 0.636411 | 3 | 0 | 3 |
| INHBE | DHX38 | 0.634634 | 3 | 0 | 3 |
| INHBE | INO80B | 0.63129 | 4 | 0 | 3 |
| INHBE | ESCO1 | 0.630983 | 3 | 0 | 3 |
| INHBE | S1PR5 | 0.628918 | 3 | 0 | 3 |
| INHBE | CYCS | 0.627593 | 3 | 0 | 3 |
| INHBE | CDH7 | 0.625903 | 3 | 0 | 3 |
| INHBE | ZNF628 | 0.622925 | 3 | 0 | 3 |
| INHBE | SLC2A9 | 0.619617 | 4 | 0 | 4 |
For details and further investigation, click here