| Full name: phosphatidylserine synthase 1 | Alias Symbol: KIAA0024|PSSA|PSS1 | ||
| Type: protein-coding gene | Cytoband: 8q22.1 | ||
| Entrez ID: 9791 | HGNC ID: HGNC:9587 | Ensembl Gene: ENSG00000156471 | OMIM ID: 612792 |
Screen Evidence:
| |||
Expression of PTDSS1:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | PTDSS1 | 9791 | 201433_s_at | 1.1075 | 0.0089 | |
| GSE20347 | PTDSS1 | 9791 | 201433_s_at | 1.1647 | 0.0000 | |
| GSE23400 | PTDSS1 | 9791 | 201433_s_at | 1.2029 | 0.0000 | |
| GSE26886 | PTDSS1 | 9791 | 201433_s_at | 0.7598 | 0.0186 | |
| GSE29001 | PTDSS1 | 9791 | 201433_s_at | 1.0538 | 0.0002 | |
| GSE38129 | PTDSS1 | 9791 | 201433_s_at | 1.2122 | 0.0000 | |
| GSE45670 | PTDSS1 | 9791 | 201433_s_at | 0.7082 | 0.0092 | |
| GSE53622 | PTDSS1 | 9791 | 41979 | 1.0866 | 0.0000 | |
| GSE53624 | PTDSS1 | 9791 | 41979 | 1.3463 | 0.0000 | |
| GSE63941 | PTDSS1 | 9791 | 201433_s_at | -0.6062 | 0.2630 | |
| GSE77861 | PTDSS1 | 9791 | 201433_s_at | 0.9396 | 0.0319 | |
| GSE97050 | PTDSS1 | 9791 | A_23_P168868 | 0.5447 | 0.1163 | |
| SRP007169 | PTDSS1 | 9791 | RNAseq | 2.2969 | 0.0000 | |
| SRP008496 | PTDSS1 | 9791 | RNAseq | 2.3073 | 0.0000 | |
| SRP064894 | PTDSS1 | 9791 | RNAseq | 1.2026 | 0.0000 | |
| SRP133303 | PTDSS1 | 9791 | RNAseq | 0.7386 | 0.0000 | |
| SRP159526 | PTDSS1 | 9791 | RNAseq | 1.5436 | 0.0013 | |
| SRP193095 | PTDSS1 | 9791 | RNAseq | 0.8790 | 0.0000 | |
| SRP219564 | PTDSS1 | 9791 | RNAseq | 0.8211 | 0.0009 | |
| TCGA | PTDSS1 | 9791 | RNAseq | 0.3115 | 0.0000 |
Upregulated datasets: 11; Downregulated datasets: 0.
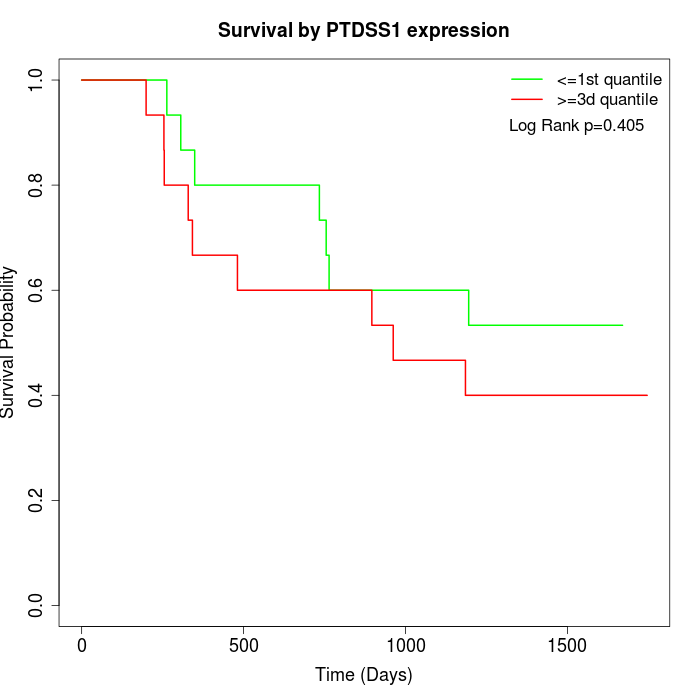
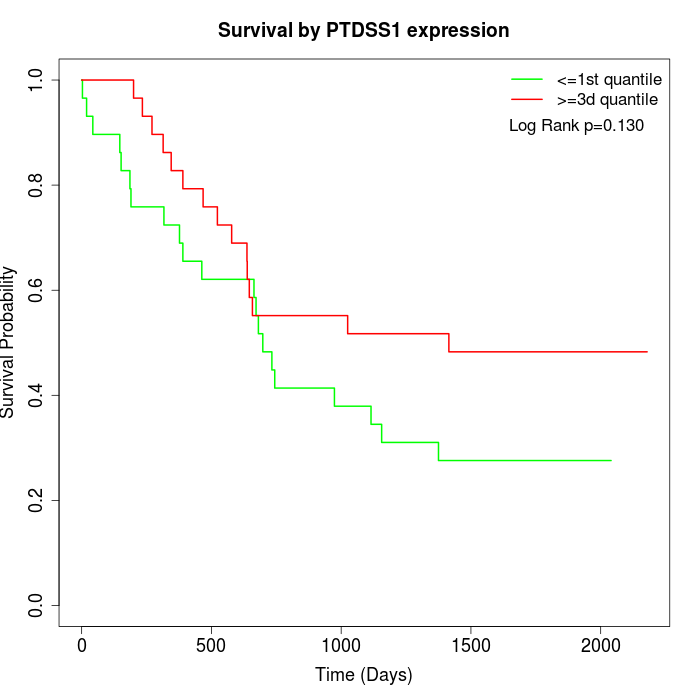
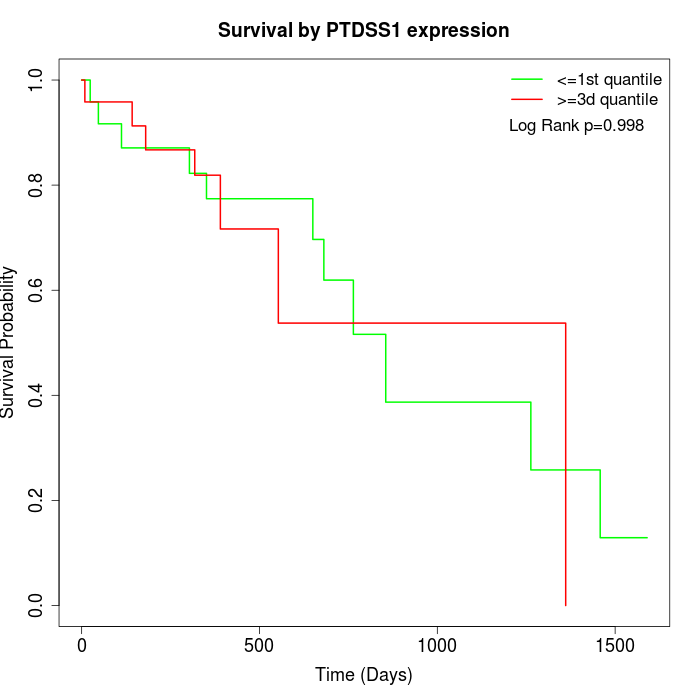
Survival by PTDSS1 expression:
Note: Click image to view full size file.
Copy number change of PTDSS1:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | PTDSS1 | 9791 | 16 | 0 | 14 | |
| GSE20123 | PTDSS1 | 9791 | 16 | 0 | 14 | |
| GSE43470 | PTDSS1 | 9791 | 21 | 1 | 21 | |
| GSE46452 | PTDSS1 | 9791 | 23 | 0 | 36 | |
| GSE47630 | PTDSS1 | 9791 | 24 | 0 | 16 | |
| GSE54993 | PTDSS1 | 9791 | 0 | 20 | 50 | |
| GSE54994 | PTDSS1 | 9791 | 35 | 1 | 17 | |
| GSE60625 | PTDSS1 | 9791 | 0 | 4 | 7 | |
| GSE74703 | PTDSS1 | 9791 | 18 | 1 | 17 | |
| GSE74704 | PTDSS1 | 9791 | 12 | 0 | 8 | |
| TCGA | PTDSS1 | 9791 | 56 | 1 | 39 |
Total number of gains: 221; Total number of losses: 28; Total Number of normals: 239.
Somatic mutations of PTDSS1:
Generating mutation plots.
Highly correlated genes for PTDSS1:
Showing top 20/2140 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| PTDSS1 | CDCA2 | 0.807195 | 4 | 0 | 4 |
| PTDSS1 | FAM89A | 0.804696 | 3 | 0 | 3 |
| PTDSS1 | SNRPE | 0.784649 | 11 | 0 | 10 |
| PTDSS1 | LAPTM4B | 0.779987 | 12 | 0 | 12 |
| PTDSS1 | ECT2 | 0.778416 | 9 | 0 | 9 |
| PTDSS1 | PUF60 | 0.773467 | 11 | 0 | 11 |
| PTDSS1 | CAPN12 | 0.772857 | 3 | 0 | 3 |
| PTDSS1 | APMAP | 0.770622 | 10 | 0 | 10 |
| PTDSS1 | NEK2 | 0.768937 | 10 | 0 | 10 |
| PTDSS1 | KCNE3 | 0.76624 | 3 | 0 | 3 |
| PTDSS1 | HSPD1 | 0.763273 | 10 | 0 | 10 |
| PTDSS1 | TFRC | 0.76283 | 11 | 0 | 11 |
| PTDSS1 | PDCD2L | 0.762726 | 7 | 0 | 7 |
| PTDSS1 | BUB1 | 0.761723 | 9 | 0 | 8 |
| PTDSS1 | FZD6 | 0.761573 | 10 | 0 | 10 |
| PTDSS1 | DHX33 | 0.76042 | 6 | 0 | 6 |
| PTDSS1 | CCT5 | 0.759992 | 8 | 0 | 8 |
| PTDSS1 | FOXM1 | 0.759294 | 9 | 0 | 9 |
| PTDSS1 | MMS22L | 0.755834 | 3 | 0 | 3 |
| PTDSS1 | ZNF707 | 0.754253 | 3 | 0 | 3 |
For details and further investigation, click here