| Full name: keratin associated protein 5-9 | Alias Symbol: KRTAP5.9|KRTAP5-1 | ||
| Type: protein-coding gene | Cytoband: 11q13.4 | ||
| Entrez ID: 3846 | HGNC ID: HGNC:23604 | Ensembl Gene: ENSG00000254997 | OMIM ID: 148021 |
Expression of KRTAP5-9:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | KRTAP5-9 | 3846 | 214517_at | -0.0746 | 0.7888 | |
| GSE20347 | KRTAP5-9 | 3846 | 214517_at | 0.0280 | 0.7762 | |
| GSE23400 | KRTAP5-9 | 3846 | 214517_at | -0.1897 | 0.0000 | |
| GSE26886 | KRTAP5-9 | 3846 | 214517_at | 0.1873 | 0.0846 | |
| GSE29001 | KRTAP5-9 | 3846 | 214517_at | -0.3399 | 0.0351 | |
| GSE38129 | KRTAP5-9 | 3846 | 214517_at | -0.0654 | 0.3753 | |
| GSE45670 | KRTAP5-9 | 3846 | 214517_at | 0.0669 | 0.5209 | |
| GSE63941 | KRTAP5-9 | 3846 | 214517_at | 0.1060 | 0.5735 | |
| GSE77861 | KRTAP5-9 | 3846 | 214517_at | 0.0049 | 0.9711 | |
| GSE97050 | KRTAP5-9 | 3846 | A_33_P3271087 | 0.1295 | 0.6699 | |
| TCGA | KRTAP5-9 | 3846 | RNAseq | 1.0980 | 0.0624 |
Upregulated datasets: 0; Downregulated datasets: 0.
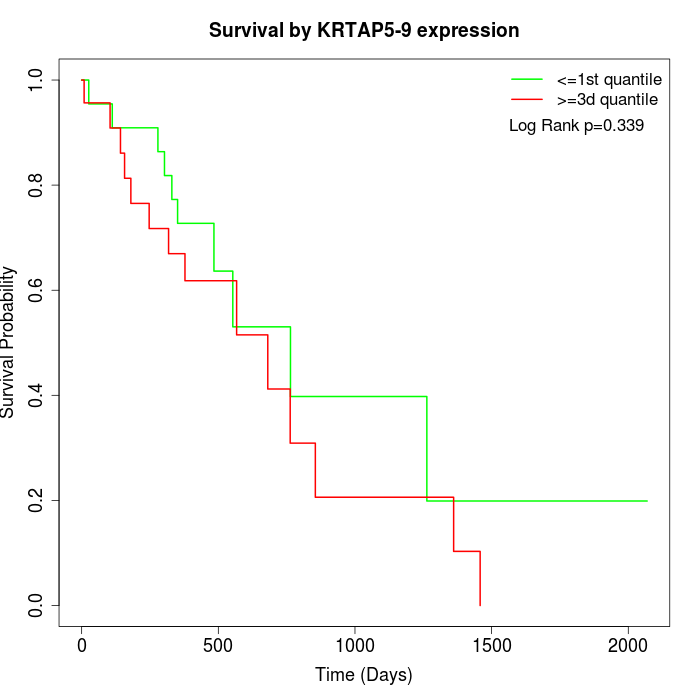
Survival by KRTAP5-9 expression:
Note: Click image to view full size file.
Copy number change of KRTAP5-9:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | KRTAP5-9 | 3846 | 11 | 6 | 13 | |
| GSE20123 | KRTAP5-9 | 3846 | 11 | 6 | 13 | |
| GSE43470 | KRTAP5-9 | 3846 | 7 | 4 | 32 | |
| GSE46452 | KRTAP5-9 | 3846 | 18 | 12 | 29 | |
| GSE47630 | KRTAP5-9 | 3846 | 11 | 9 | 20 | |
| GSE54993 | KRTAP5-9 | 3846 | 6 | 6 | 58 | |
| GSE54994 | KRTAP5-9 | 3846 | 11 | 12 | 30 | |
| GSE60625 | KRTAP5-9 | 3846 | 0 | 7 | 4 | |
| GSE74703 | KRTAP5-9 | 3846 | 6 | 3 | 27 | |
| GSE74704 | KRTAP5-9 | 3846 | 7 | 4 | 9 |
Total number of gains: 88; Total number of losses: 69; Total Number of normals: 235.
Somatic mutations of KRTAP5-9:
Generating mutation plots.
Highly correlated genes for KRTAP5-9:
Showing top 20/812 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| KRTAP5-9 | HRC | 0.751454 | 3 | 0 | 3 |
| KRTAP5-9 | ACR | 0.742735 | 4 | 0 | 4 |
| KRTAP5-9 | ADM2 | 0.741027 | 4 | 0 | 4 |
| KRTAP5-9 | ETV2 | 0.723039 | 3 | 0 | 3 |
| KRTAP5-9 | LALBA | 0.721093 | 3 | 0 | 3 |
| KRTAP5-9 | PTPRN2 | 0.717068 | 3 | 0 | 3 |
| KRTAP5-9 | EDA2R | 0.716697 | 4 | 0 | 4 |
| KRTAP5-9 | CGA | 0.71352 | 4 | 0 | 4 |
| KRTAP5-9 | ZNF835 | 0.710935 | 3 | 0 | 3 |
| KRTAP5-9 | CRHR2 | 0.707237 | 4 | 0 | 4 |
| KRTAP5-9 | EPS8L3 | 0.704486 | 4 | 0 | 4 |
| KRTAP5-9 | PSG1 | 0.703098 | 5 | 0 | 5 |
| KRTAP5-9 | RGR | 0.702941 | 4 | 0 | 4 |
| KRTAP5-9 | PRDM12 | 0.699783 | 3 | 0 | 3 |
| KRTAP5-9 | CYP2A7 | 0.6977 | 4 | 0 | 4 |
| KRTAP5-9 | LINC01260 | 0.695761 | 4 | 0 | 4 |
| KRTAP5-9 | CLCA1 | 0.695439 | 4 | 0 | 4 |
| KRTAP5-9 | CER1 | 0.694868 | 5 | 0 | 5 |
| KRTAP5-9 | LMAN1L | 0.690406 | 4 | 0 | 4 |
| KRTAP5-9 | PAX7 | 0.689083 | 5 | 0 | 5 |
For details and further investigation, click here