| Full name: galectin like | Alias Symbol: HSPC159|GRP | ||
| Type: protein-coding gene | Cytoband: 2p14 | ||
| Entrez ID: 29094 | HGNC ID: HGNC:25012 | Ensembl Gene: ENSG00000119862 | OMIM ID: 617902 |
| Drug and gene relationship at DGIdb | |||
Expression of LGALSL:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | LGALSL | 29094 | 226188_at | -0.1775 | 0.9089 | |
| GSE20347 | LGALSL | 29094 | 219998_at | -0.5808 | 0.0074 | |
| GSE23400 | LGALSL | 29094 | 219998_at | -0.3196 | 0.0064 | |
| GSE26886 | LGALSL | 29094 | 226188_at | -1.2538 | 0.0003 | |
| GSE29001 | LGALSL | 29094 | 219998_at | -0.8183 | 0.1744 | |
| GSE38129 | LGALSL | 29094 | 219998_at | -0.1106 | 0.6057 | |
| GSE45670 | LGALSL | 29094 | 226188_at | 0.3378 | 0.2550 | |
| GSE53622 | LGALSL | 29094 | 28006 | -0.6842 | 0.0000 | |
| GSE53624 | LGALSL | 29094 | 28006 | -0.8699 | 0.0000 | |
| GSE63941 | LGALSL | 29094 | 226188_at | 0.6928 | 0.3490 | |
| GSE77861 | LGALSL | 29094 | 226188_at | -0.0562 | 0.9022 | |
| GSE97050 | LGALSL | 29094 | A_23_P210330 | -0.2668 | 0.3770 | |
| SRP007169 | LGALSL | 29094 | RNAseq | -0.4301 | 0.3709 | |
| SRP008496 | LGALSL | 29094 | RNAseq | -0.5030 | 0.1309 | |
| SRP064894 | LGALSL | 29094 | RNAseq | -0.7220 | 0.0028 | |
| SRP133303 | LGALSL | 29094 | RNAseq | -0.5620 | 0.0759 | |
| SRP159526 | LGALSL | 29094 | RNAseq | 0.1372 | 0.7155 | |
| SRP193095 | LGALSL | 29094 | RNAseq | -0.7655 | 0.0012 | |
| SRP219564 | LGALSL | 29094 | RNAseq | -0.9542 | 0.0326 |
Upregulated datasets: 0; Downregulated datasets: 1.
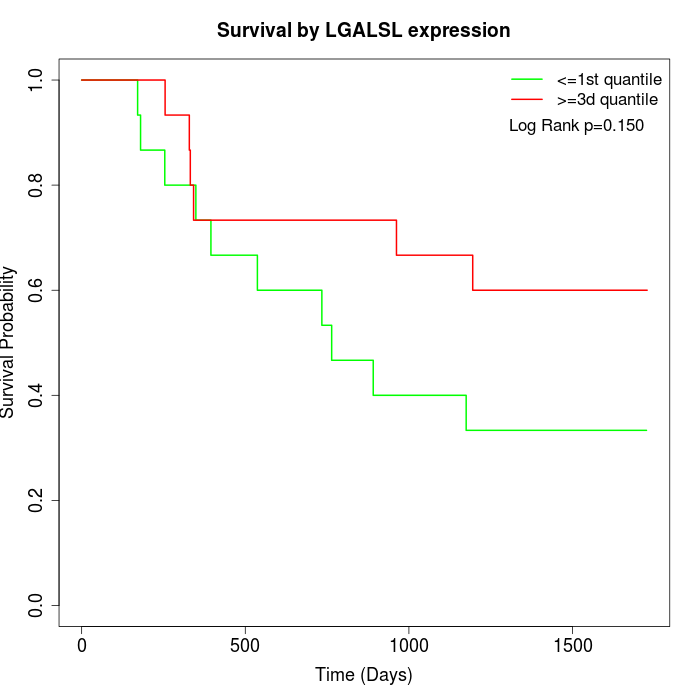
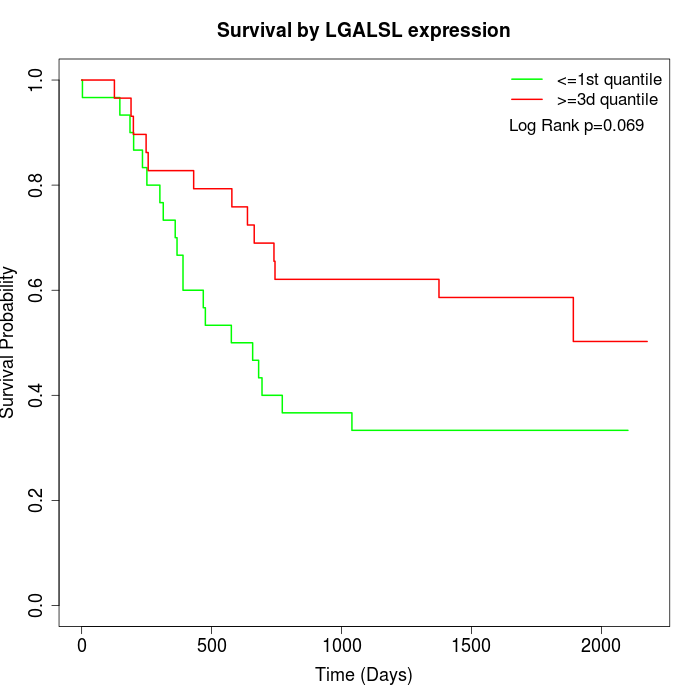
Survival by LGALSL expression:
Note: Click image to view full size file.
Copy number change of LGALSL:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | LGALSL | 29094 | 12 | 0 | 18 | |
| GSE20123 | LGALSL | 29094 | 12 | 0 | 18 | |
| GSE43470 | LGALSL | 29094 | 7 | 0 | 36 | |
| GSE46452 | LGALSL | 29094 | 2 | 3 | 54 | |
| GSE47630 | LGALSL | 29094 | 8 | 0 | 32 | |
| GSE54993 | LGALSL | 29094 | 0 | 6 | 64 | |
| GSE54994 | LGALSL | 29094 | 12 | 0 | 41 | |
| GSE60625 | LGALSL | 29094 | 0 | 3 | 8 | |
| GSE74703 | LGALSL | 29094 | 7 | 0 | 29 | |
| GSE74704 | LGALSL | 29094 | 10 | 0 | 10 | |
| TCGA | LGALSL | 29094 | 36 | 2 | 58 |
Total number of gains: 106; Total number of losses: 14; Total Number of normals: 368.
Somatic mutations of LGALSL:
Generating mutation plots.
Highly correlated genes for LGALSL:
Showing top 20/588 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| LGALSL | ANKRD13C | 0.807588 | 3 | 0 | 3 |
| LGALSL | RNF181 | 0.79271 | 3 | 0 | 3 |
| LGALSL | CUL2 | 0.768515 | 3 | 0 | 3 |
| LGALSL | ABCB6 | 0.763158 | 3 | 0 | 3 |
| LGALSL | CNOT4 | 0.752718 | 3 | 0 | 3 |
| LGALSL | MOB3C | 0.751749 | 3 | 0 | 3 |
| LGALSL | TMEM17 | 0.749185 | 3 | 0 | 3 |
| LGALSL | PPARGC1B | 0.735319 | 3 | 0 | 3 |
| LGALSL | RAB8A | 0.723634 | 3 | 0 | 3 |
| LGALSL | SPAST | 0.711194 | 3 | 0 | 3 |
| LGALSL | RBAK | 0.710981 | 3 | 0 | 3 |
| LGALSL | FECH | 0.707823 | 3 | 0 | 3 |
| LGALSL | EAF1 | 0.707666 | 4 | 0 | 3 |
| LGALSL | FOPNL | 0.706246 | 4 | 0 | 4 |
| LGALSL | OXNAD1 | 0.701967 | 3 | 0 | 3 |
| LGALSL | UBE2G2 | 0.69977 | 3 | 0 | 3 |
| LGALSL | ACAT1 | 0.698667 | 3 | 0 | 3 |
| LGALSL | SNX19 | 0.698284 | 3 | 0 | 3 |
| LGALSL | PTCD2 | 0.692355 | 3 | 0 | 3 |
| LGALSL | CHM | 0.690187 | 3 | 0 | 3 |
For details and further investigation, click here