| Full name: mitogen-activated protein kinase kinase kinase kinase 2 | Alias Symbol: GCK|BL44 | ||
| Type: protein-coding gene | Cytoband: 11q13.1 | ||
| Entrez ID: 5871 | HGNC ID: HGNC:6864 | Ensembl Gene: ENSG00000168067 | OMIM ID: 603166 |
| Drug and gene relationship at DGIdb | |||
MAP4K2 involved pathways:
| KEGG pathway | Description | View |
|---|---|---|
| hsa04010 | MAPK signaling pathway |
Expression of MAP4K2:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | MAP4K2 | 5871 | 204936_at | 0.9564 | 0.0265 | |
| GSE20347 | MAP4K2 | 5871 | 204936_at | 0.6458 | 0.0000 | |
| GSE23400 | MAP4K2 | 5871 | 204936_at | -0.0136 | 0.6967 | |
| GSE26886 | MAP4K2 | 5871 | 204936_at | 1.0263 | 0.0000 | |
| GSE29001 | MAP4K2 | 5871 | 204936_at | 0.1707 | 0.1723 | |
| GSE38129 | MAP4K2 | 5871 | 204936_at | 0.6891 | 0.0000 | |
| GSE45670 | MAP4K2 | 5871 | 204936_at | 0.4859 | 0.0012 | |
| GSE53622 | MAP4K2 | 5871 | 20476 | 0.9245 | 0.0000 | |
| GSE53624 | MAP4K2 | 5871 | 20476 | 1.2347 | 0.0000 | |
| GSE63941 | MAP4K2 | 5871 | 204936_at | 0.8759 | 0.0519 | |
| GSE77861 | MAP4K2 | 5871 | 204936_at | 0.1757 | 0.2931 | |
| GSE97050 | MAP4K2 | 5871 | A_24_P287075 | 0.5043 | 0.2396 | |
| SRP007169 | MAP4K2 | 5871 | RNAseq | 1.1823 | 0.0232 | |
| SRP008496 | MAP4K2 | 5871 | RNAseq | 1.1615 | 0.0002 | |
| SRP064894 | MAP4K2 | 5871 | RNAseq | 0.6043 | 0.0000 | |
| SRP133303 | MAP4K2 | 5871 | RNAseq | 0.4654 | 0.0196 | |
| SRP159526 | MAP4K2 | 5871 | RNAseq | 0.9856 | 0.0004 | |
| SRP193095 | MAP4K2 | 5871 | RNAseq | 0.7933 | 0.0000 | |
| SRP219564 | MAP4K2 | 5871 | RNAseq | 0.3207 | 0.3311 | |
| TCGA | MAP4K2 | 5871 | RNAseq | 0.1001 | 0.1384 |
Upregulated datasets: 4; Downregulated datasets: 0.
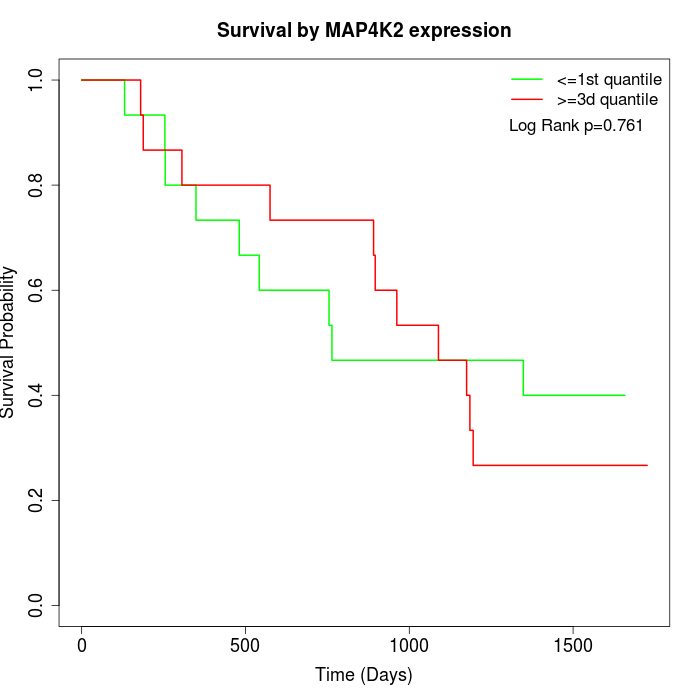
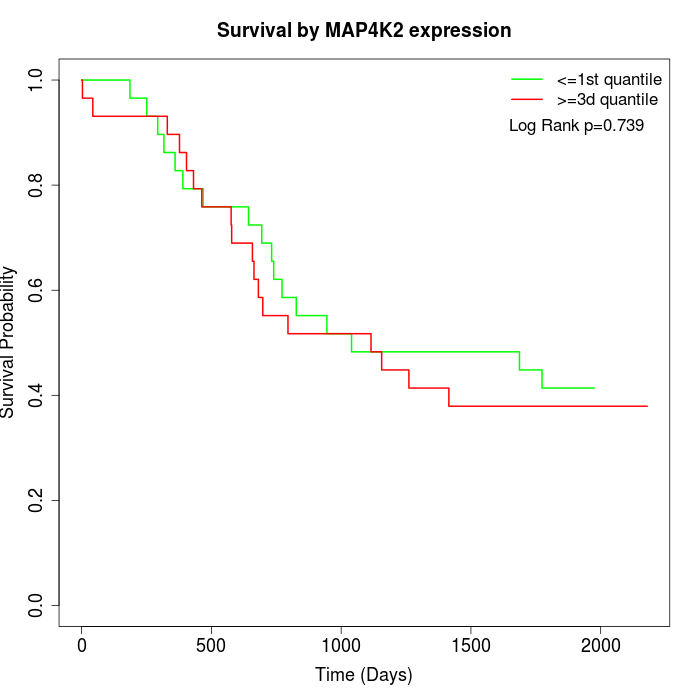
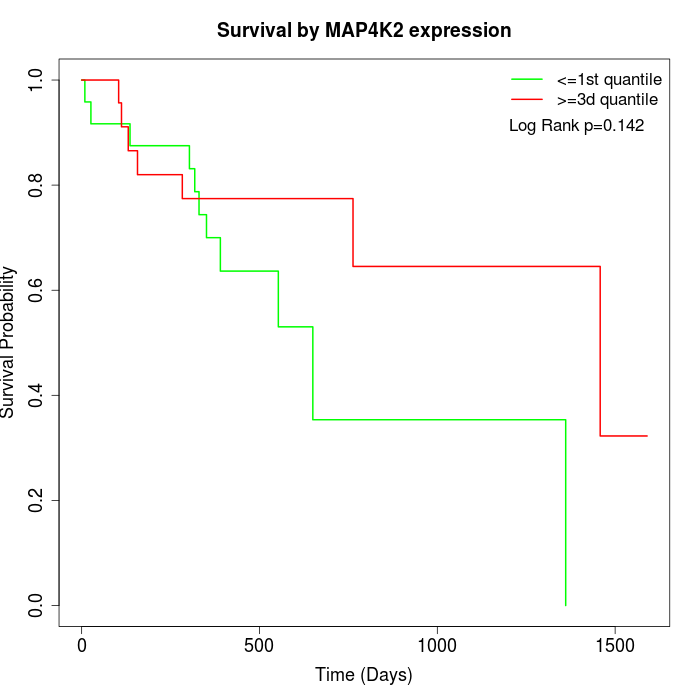
Survival by MAP4K2 expression:
Note: Click image to view full size file.
Copy number change of MAP4K2:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | MAP4K2 | 5871 | 7 | 6 | 17 | |
| GSE20123 | MAP4K2 | 5871 | 7 | 6 | 17 | |
| GSE43470 | MAP4K2 | 5871 | 1 | 3 | 39 | |
| GSE46452 | MAP4K2 | 5871 | 9 | 4 | 46 | |
| GSE47630 | MAP4K2 | 5871 | 5 | 6 | 29 | |
| GSE54993 | MAP4K2 | 5871 | 3 | 1 | 66 | |
| GSE54994 | MAP4K2 | 5871 | 6 | 5 | 42 | |
| GSE60625 | MAP4K2 | 5871 | 0 | 3 | 8 | |
| GSE74703 | MAP4K2 | 5871 | 1 | 1 | 34 | |
| GSE74704 | MAP4K2 | 5871 | 5 | 4 | 11 | |
| TCGA | MAP4K2 | 5871 | 20 | 8 | 68 |
Total number of gains: 64; Total number of losses: 47; Total Number of normals: 377.
Somatic mutations of MAP4K2:
Generating mutation plots.
Highly correlated genes for MAP4K2:
Showing top 20/1740 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| MAP4K2 | A4GALT | 0.815922 | 4 | 0 | 4 |
| MAP4K2 | ZNF358 | 0.794943 | 3 | 0 | 3 |
| MAP4K2 | ROMO1 | 0.771168 | 3 | 0 | 3 |
| MAP4K2 | ZNF707 | 0.758939 | 3 | 0 | 3 |
| MAP4K2 | IFFO2 | 0.754558 | 3 | 0 | 3 |
| MAP4K2 | DNMT1 | 0.750608 | 8 | 0 | 8 |
| MAP4K2 | DBF4 | 0.74933 | 6 | 0 | 6 |
| MAP4K2 | SLC35B2 | 0.74589 | 3 | 0 | 3 |
| MAP4K2 | THOC6 | 0.742108 | 6 | 0 | 6 |
| MAP4K2 | COMMD2 | 0.742059 | 3 | 0 | 3 |
| MAP4K2 | PYCR2 | 0.741044 | 3 | 0 | 3 |
| MAP4K2 | FIP1L1 | 0.740855 | 3 | 0 | 3 |
| MAP4K2 | INO80E | 0.738865 | 4 | 0 | 3 |
| MAP4K2 | CCDC86 | 0.737019 | 6 | 0 | 5 |
| MAP4K2 | RAB34 | 0.735684 | 3 | 0 | 3 |
| MAP4K2 | TAOK1 | 0.733891 | 3 | 0 | 3 |
| MAP4K2 | NACC1 | 0.73217 | 4 | 0 | 4 |
| MAP4K2 | MCM6 | 0.729559 | 8 | 0 | 8 |
| MAP4K2 | PLXNA1 | 0.728473 | 9 | 0 | 8 |
| MAP4K2 | FADS1 | 0.728261 | 3 | 0 | 3 |
For details and further investigation, click here