| Full name: MAM domain containing glycosylphosphatidylinositol anchor 2 | Alias Symbol: | ||
| Type: protein-coding gene | Cytoband: 14q21.3 | ||
| Entrez ID: 161357 | HGNC ID: HGNC:19835 | Ensembl Gene: ENSG00000139915 | OMIM ID: 611128 |
| Drug and gene relationship at DGIdb | |||
Expression of MDGA2:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | MDGA2 | 161357 | 1570114_at | 0.0453 | 0.8757 | |
| GSE26886 | MDGA2 | 161357 | 1570114_at | 0.1319 | 0.0987 | |
| GSE45670 | MDGA2 | 161357 | 1570114_at | 0.0933 | 0.1794 | |
| GSE53622 | MDGA2 | 161357 | 20522 | 0.1956 | 0.3278 | |
| GSE53624 | MDGA2 | 161357 | 20522 | 0.5954 | 0.0001 | |
| GSE63941 | MDGA2 | 161357 | 1570114_at | -0.0491 | 0.7365 | |
| GSE77861 | MDGA2 | 161357 | 1570114_at | -0.1148 | 0.2285 | |
| TCGA | MDGA2 | 161357 | RNAseq | -0.3999 | 0.7094 |
Upregulated datasets: 0; Downregulated datasets: 0.
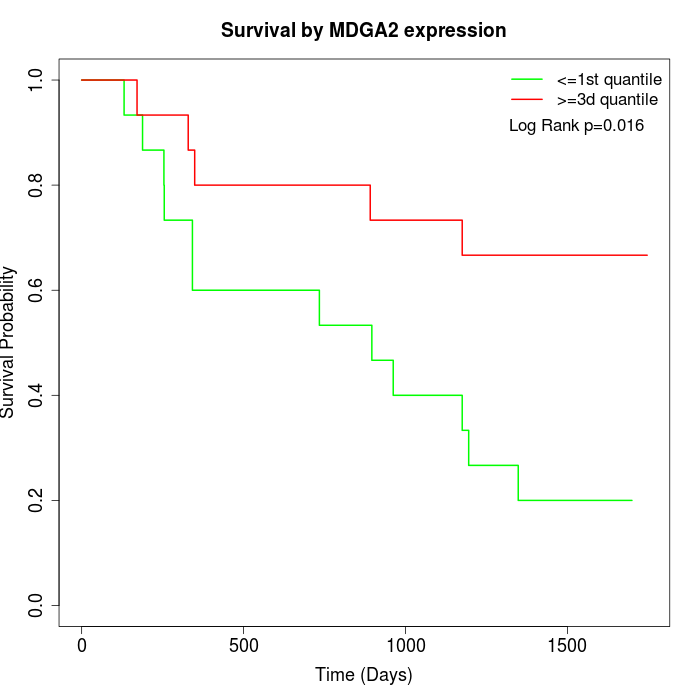
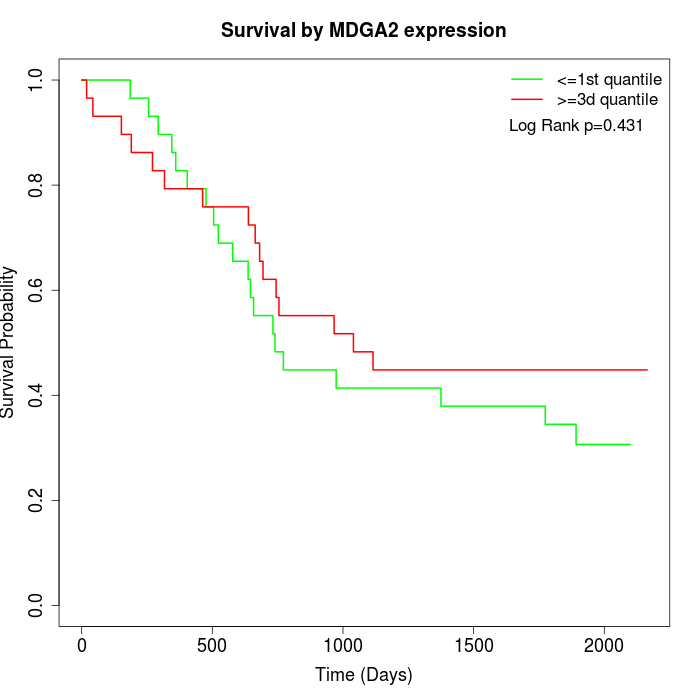
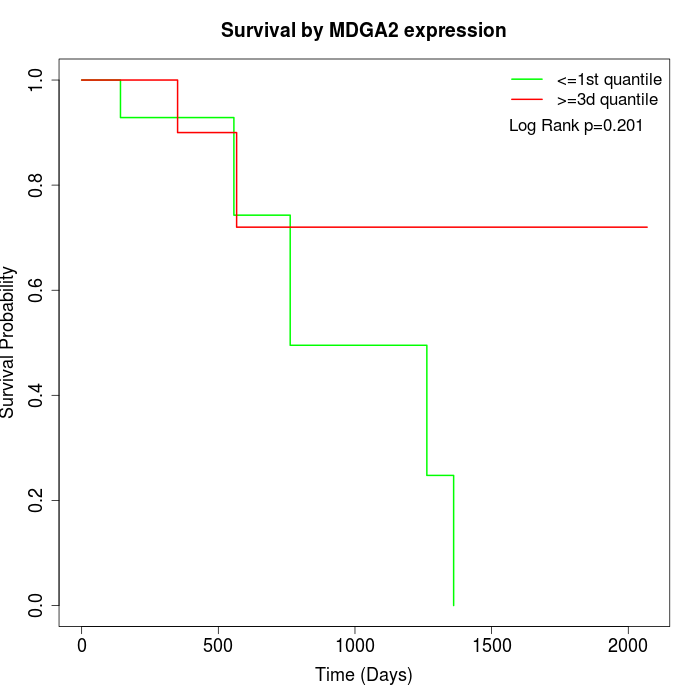
Survival by MDGA2 expression:
Note: Click image to view full size file.
Copy number change of MDGA2:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | MDGA2 | 161357 | 6 | 5 | 19 | |
| GSE20123 | MDGA2 | 161357 | 6 | 5 | 19 | |
| GSE43470 | MDGA2 | 161357 | 10 | 2 | 31 | |
| GSE46452 | MDGA2 | 161357 | 16 | 3 | 40 | |
| GSE47630 | MDGA2 | 161357 | 11 | 10 | 19 | |
| GSE54993 | MDGA2 | 161357 | 3 | 9 | 58 | |
| GSE54994 | MDGA2 | 161357 | 19 | 4 | 30 | |
| GSE60625 | MDGA2 | 161357 | 0 | 2 | 9 | |
| GSE74703 | MDGA2 | 161357 | 9 | 2 | 25 | |
| GSE74704 | MDGA2 | 161357 | 3 | 3 | 14 | |
| TCGA | MDGA2 | 161357 | 33 | 14 | 49 |
Total number of gains: 116; Total number of losses: 59; Total Number of normals: 313.
Somatic mutations of MDGA2:
Generating mutation plots.
Highly correlated genes for MDGA2:
Showing top 20/28 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| MDGA2 | DOC2A | 0.664351 | 3 | 0 | 3 |
| MDGA2 | C2orf81 | 0.652033 | 3 | 0 | 3 |
| MDGA2 | GYPA | 0.6288 | 3 | 0 | 3 |
| MDGA2 | POU3F1 | 0.61588 | 3 | 0 | 3 |
| MDGA2 | ROBO4 | 0.61245 | 3 | 0 | 3 |
| MDGA2 | SIRT6 | 0.603061 | 3 | 0 | 3 |
| MDGA2 | DSCAM | 0.588019 | 3 | 0 | 3 |
| MDGA2 | SRCAP | 0.582646 | 3 | 0 | 3 |
| MDGA2 | NKX1-1 | 0.571527 | 4 | 0 | 3 |
| MDGA2 | CLCF1 | 0.571487 | 3 | 0 | 3 |
| MDGA2 | CYP2A13 | 0.571377 | 3 | 0 | 3 |
| MDGA2 | GP9 | 0.56865 | 3 | 0 | 3 |
| MDGA2 | NCS1 | 0.564041 | 3 | 0 | 3 |
| MDGA2 | VSX1 | 0.559353 | 4 | 0 | 3 |
| MDGA2 | SV2B | 0.551763 | 3 | 0 | 3 |
| MDGA2 | ANKRD34C | 0.546748 | 4 | 0 | 3 |
| MDGA2 | P2RY4 | 0.540929 | 3 | 0 | 3 |
| MDGA2 | PCBP4 | 0.540433 | 4 | 0 | 3 |
| MDGA2 | KIF5A | 0.536642 | 3 | 0 | 3 |
| MDGA2 | ASB10 | 0.536294 | 3 | 0 | 3 |
For details and further investigation, click here