| Full name: meiosis regulator for oocyte development | Alias Symbol: FLJ20323|MIO|Sea4|Yulink | ||
| Type: protein-coding gene | Cytoband: 7p21.3 | ||
| Entrez ID: 54468 | HGNC ID: HGNC:21905 | Ensembl Gene: ENSG00000164654 | OMIM ID: 615359 |
Screen Evidence:
| |||
MIOS involved pathways:
| KEGG pathway | Description | View |
|---|---|---|
| hsa04150 | mTOR signaling pathway |
Expression of MIOS:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | MIOS | 54468 | 211724_x_at | -0.5230 | 0.3490 | |
| GSE20347 | MIOS | 54468 | 211724_x_at | -0.7152 | 0.0000 | |
| GSE23400 | MIOS | 54468 | 211724_x_at | -0.5144 | 0.0000 | |
| GSE26886 | MIOS | 54468 | 211724_x_at | -0.7094 | 0.0019 | |
| GSE29001 | MIOS | 54468 | 211724_x_at | -0.7801 | 0.0277 | |
| GSE38129 | MIOS | 54468 | 211724_x_at | -0.3967 | 0.0321 | |
| GSE45670 | MIOS | 54468 | 211724_x_at | -0.4332 | 0.0103 | |
| GSE53622 | MIOS | 54468 | 6503 | -0.5968 | 0.0000 | |
| GSE53624 | MIOS | 54468 | 6503 | -0.9336 | 0.0000 | |
| GSE63941 | MIOS | 54468 | 211724_x_at | 0.9903 | 0.0128 | |
| GSE77861 | MIOS | 54468 | 211724_x_at | -0.5590 | 0.0392 | |
| GSE97050 | MIOS | 54468 | A_23_P59602 | 0.3949 | 0.4468 | |
| SRP007169 | MIOS | 54468 | RNAseq | -1.6147 | 0.0000 | |
| SRP008496 | MIOS | 54468 | RNAseq | -1.1838 | 0.0000 | |
| SRP064894 | MIOS | 54468 | RNAseq | -1.0282 | 0.0000 | |
| SRP133303 | MIOS | 54468 | RNAseq | -0.6385 | 0.0002 | |
| SRP159526 | MIOS | 54468 | RNAseq | -1.1794 | 0.0000 | |
| SRP193095 | MIOS | 54468 | RNAseq | -0.9911 | 0.0000 | |
| SRP219564 | MIOS | 54468 | RNAseq | -0.9078 | 0.0872 | |
| TCGA | MIOS | 54468 | RNAseq | -0.1605 | 0.0035 |
Upregulated datasets: 0; Downregulated datasets: 4.
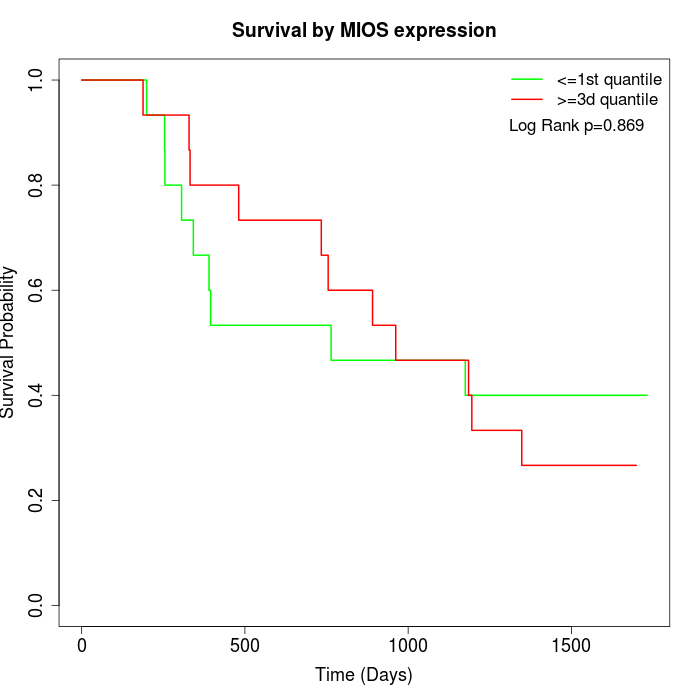
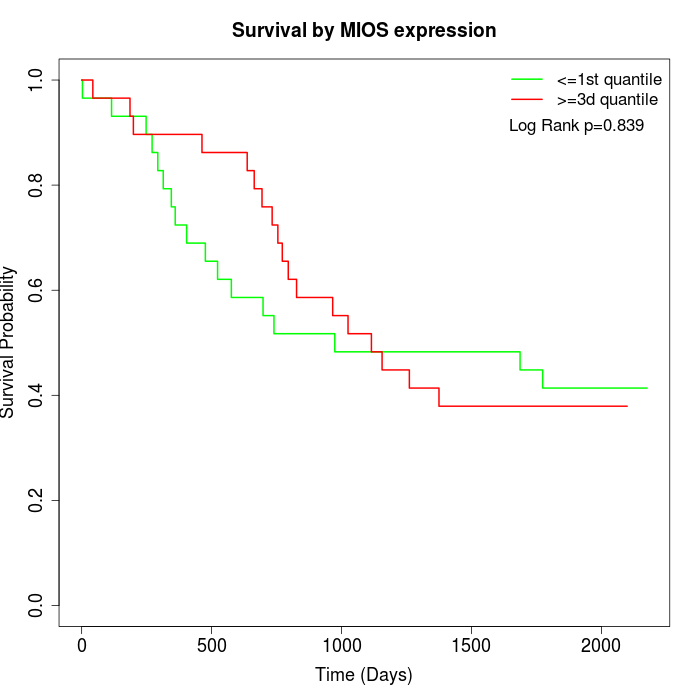
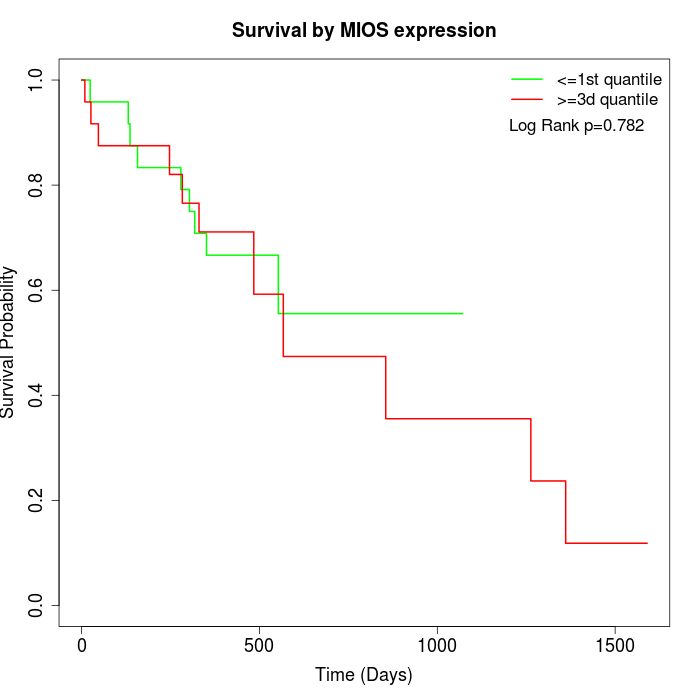
Survival by MIOS expression:
Note: Click image to view full size file.
Copy number change of MIOS:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | MIOS | 54468 | 15 | 2 | 13 | |
| GSE20123 | MIOS | 54468 | 15 | 2 | 13 | |
| GSE43470 | MIOS | 54468 | 8 | 0 | 35 | |
| GSE46452 | MIOS | 54468 | 11 | 1 | 47 | |
| GSE47630 | MIOS | 54468 | 9 | 1 | 30 | |
| GSE54993 | MIOS | 54468 | 0 | 6 | 64 | |
| GSE54994 | MIOS | 54468 | 21 | 1 | 31 | |
| GSE60625 | MIOS | 54468 | 0 | 0 | 11 | |
| GSE74703 | MIOS | 54468 | 8 | 0 | 28 | |
| GSE74704 | MIOS | 54468 | 9 | 1 | 10 | |
| TCGA | MIOS | 54468 | 56 | 4 | 36 |
Total number of gains: 152; Total number of losses: 18; Total Number of normals: 318.
Somatic mutations of MIOS:
Generating mutation plots.
Highly correlated genes for MIOS:
Showing top 20/1355 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| MIOS | ELOVL1 | 0.793337 | 3 | 0 | 3 |
| MIOS | CYSRT1 | 0.784858 | 6 | 0 | 6 |
| MIOS | TTC39B | 0.77949 | 3 | 0 | 3 |
| MIOS | RASSF5 | 0.779294 | 7 | 0 | 7 |
| MIOS | A2ML1 | 0.777389 | 6 | 0 | 6 |
| MIOS | S100A16 | 0.772971 | 7 | 0 | 7 |
| MIOS | PLD1 | 0.772747 | 11 | 0 | 11 |
| MIOS | ACPP | 0.764006 | 10 | 0 | 10 |
| MIOS | LTA4H | 0.760494 | 3 | 0 | 3 |
| MIOS | KRT78 | 0.759405 | 6 | 0 | 6 |
| MIOS | CAPN14 | 0.756381 | 6 | 0 | 6 |
| MIOS | ZNF185 | 0.755013 | 11 | 0 | 10 |
| MIOS | C6orf132 | 0.753964 | 6 | 0 | 6 |
| MIOS | SFTA2 | 0.746473 | 4 | 0 | 4 |
| MIOS | VSIG10L | 0.741294 | 6 | 0 | 6 |
| MIOS | RAB25 | 0.738248 | 12 | 0 | 12 |
| MIOS | CSTB | 0.737415 | 10 | 0 | 9 |
| MIOS | IL1RN | 0.736445 | 10 | 0 | 10 |
| MIOS | PRSS27 | 0.736396 | 7 | 0 | 7 |
| MIOS | BLNK | 0.735659 | 11 | 0 | 11 |
For details and further investigation, click here