| Full name: 2'-5'-oligoadenylate synthetase 1 | Alias Symbol: OIASI|IFI-4 | ||
| Type: protein-coding gene | Cytoband: 12q24.13 | ||
| Entrez ID: 4938 | HGNC ID: HGNC:8086 | Ensembl Gene: ENSG00000089127 | OMIM ID: 164350 |
OAS1 involved pathways:
| KEGG pathway | Description | View |
|---|---|---|
| hsa05164 | Influenza A | |
| hsa05168 | Herpes simplex infection |
Expression of OAS1:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | OAS1 | 4938 | 202869_at | -0.2758 | 0.8213 | |
| GSE20347 | OAS1 | 4938 | 202869_at | -0.1709 | 0.6339 | |
| GSE23400 | OAS1 | 4938 | 202869_at | 0.0521 | 0.6977 | |
| GSE26886 | OAS1 | 4938 | 202869_at | -1.1411 | 0.0154 | |
| GSE29001 | OAS1 | 4938 | 202869_at | 0.4587 | 0.2438 | |
| GSE38129 | OAS1 | 4938 | 202869_at | 0.0021 | 0.9956 | |
| GSE45670 | OAS1 | 4938 | 202869_at | 1.0737 | 0.0140 | |
| GSE53622 | OAS1 | 4938 | 136796 | -0.1449 | 0.4259 | |
| GSE53624 | OAS1 | 4938 | 136796 | -0.2296 | 0.0651 | |
| GSE63941 | OAS1 | 4938 | 202869_at | 1.4796 | 0.3201 | |
| GSE77861 | OAS1 | 4938 | 202869_at | -0.7335 | 0.1054 | |
| GSE97050 | OAS1 | 4938 | A_23_P64828 | 0.1089 | 0.9113 | |
| SRP007169 | OAS1 | 4938 | RNAseq | -0.2835 | 0.5823 | |
| SRP008496 | OAS1 | 4938 | RNAseq | -0.8987 | 0.0260 | |
| SRP064894 | OAS1 | 4938 | RNAseq | 0.2994 | 0.3986 | |
| SRP133303 | OAS1 | 4938 | RNAseq | -0.1400 | 0.6708 | |
| SRP159526 | OAS1 | 4938 | RNAseq | -1.7722 | 0.0345 | |
| SRP193095 | OAS1 | 4938 | RNAseq | -0.2646 | 0.2163 | |
| SRP219564 | OAS1 | 4938 | RNAseq | 0.1978 | 0.7947 | |
| TCGA | OAS1 | 4938 | RNAseq | 0.3728 | 0.0013 |
Upregulated datasets: 1; Downregulated datasets: 2.
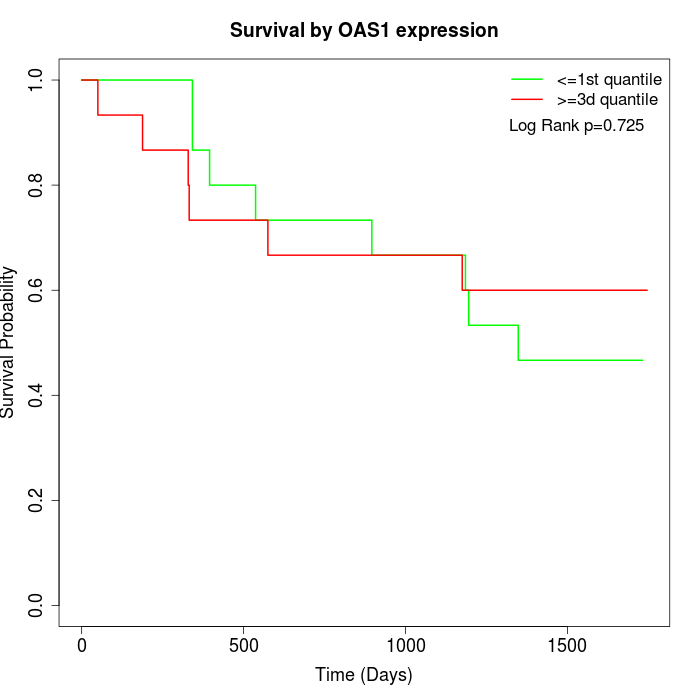
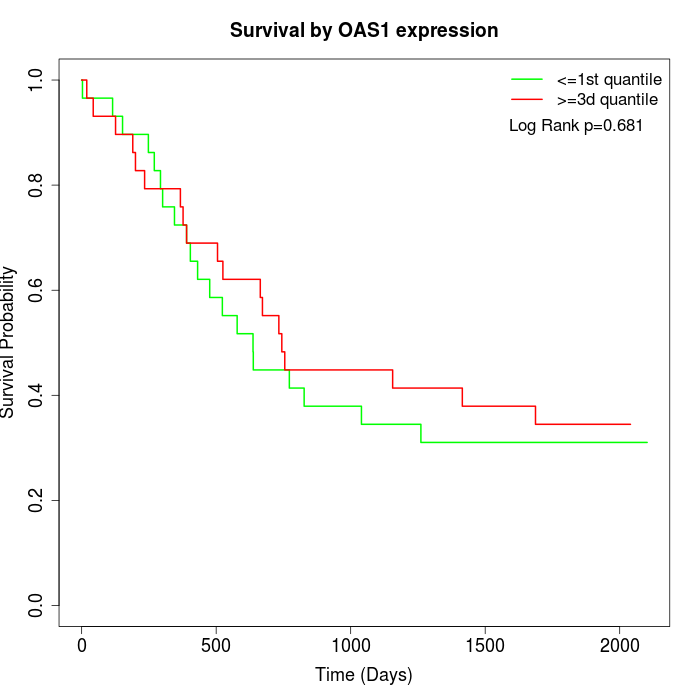
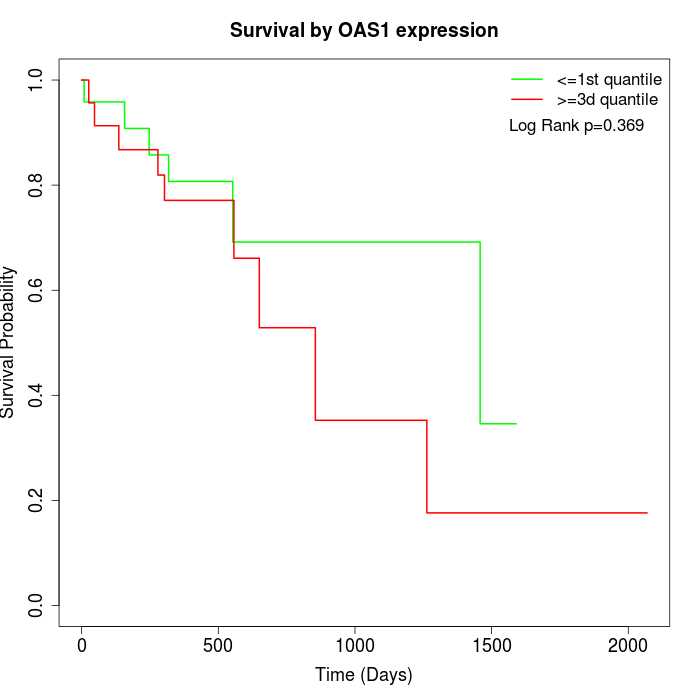
Survival by OAS1 expression:
Note: Click image to view full size file.
Copy number change of OAS1:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | OAS1 | 4938 | 4 | 3 | 23 | |
| GSE20123 | OAS1 | 4938 | 4 | 3 | 23 | |
| GSE43470 | OAS1 | 4938 | 2 | 0 | 41 | |
| GSE46452 | OAS1 | 4938 | 9 | 1 | 49 | |
| GSE47630 | OAS1 | 4938 | 9 | 3 | 28 | |
| GSE54993 | OAS1 | 4938 | 0 | 5 | 65 | |
| GSE54994 | OAS1 | 4938 | 4 | 2 | 47 | |
| GSE60625 | OAS1 | 4938 | 0 | 0 | 11 | |
| GSE74703 | OAS1 | 4938 | 2 | 0 | 34 | |
| GSE74704 | OAS1 | 4938 | 2 | 2 | 16 | |
| TCGA | OAS1 | 4938 | 22 | 11 | 63 |
Total number of gains: 58; Total number of losses: 30; Total Number of normals: 400.
Somatic mutations of OAS1:
Generating mutation plots.
Highly correlated genes for OAS1:
Showing top 20/234 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| OAS1 | GRPEL2 | 0.712155 | 3 | 0 | 3 |
| OAS1 | IDI1 | 0.707914 | 3 | 0 | 3 |
| OAS1 | HERC6 | 0.695378 | 13 | 0 | 12 |
| OAS1 | MX1 | 0.68464 | 12 | 0 | 11 |
| OAS1 | CNOT6L | 0.683808 | 3 | 0 | 3 |
| OAS1 | KLRC4 | 0.683752 | 3 | 0 | 3 |
| OAS1 | SPAG16 | 0.682085 | 3 | 0 | 3 |
| OAS1 | ACAA1 | 0.672658 | 3 | 0 | 3 |
| OAS1 | DCAF10 | 0.66467 | 3 | 0 | 3 |
| OAS1 | MRPS36 | 0.66089 | 3 | 0 | 3 |
| OAS1 | ATP6V1D | 0.660378 | 3 | 0 | 3 |
| OAS1 | PDCD6IP | 0.658217 | 3 | 0 | 3 |
| OAS1 | ZNF211 | 0.655327 | 4 | 0 | 3 |
| OAS1 | TJP1 | 0.650748 | 3 | 0 | 3 |
| OAS1 | FBXL5 | 0.64791 | 3 | 0 | 3 |
| OAS1 | SAMD9L | 0.64522 | 8 | 0 | 6 |
| OAS1 | EPS15 | 0.644457 | 3 | 0 | 3 |
| OAS1 | UBE2W | 0.643321 | 3 | 0 | 3 |
| OAS1 | DNAJC1 | 0.64076 | 4 | 0 | 4 |
| OAS1 | HIGD1A | 0.637437 | 3 | 0 | 3 |
For details and further investigation, click here