| Full name: 3-oxoacyl-ACP synthase, mitochondrial | Alias Symbol: KS|FLJ20604|FASN2D|CEM1 | ||
| Type: protein-coding gene | Cytoband: 3p24.2 | ||
| Entrez ID: 54995 | HGNC ID: HGNC:26063 | Ensembl Gene: ENSG00000151093 | OMIM ID: 610324 |
Screen Evidence:
| |||
Expression of OXSM:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | OXSM | 54995 | 219133_at | -0.1936 | 0.6321 | |
| GSE20347 | OXSM | 54995 | 219133_at | -0.3677 | 0.0100 | |
| GSE23400 | OXSM | 54995 | 219133_at | 0.1141 | 0.0978 | |
| GSE26886 | OXSM | 54995 | 219133_at | -0.7923 | 0.0001 | |
| GSE29001 | OXSM | 54995 | 219133_at | -0.3690 | 0.1806 | |
| GSE38129 | OXSM | 54995 | 219133_at | -0.2088 | 0.1197 | |
| GSE45670 | OXSM | 54995 | 219133_at | -0.0614 | 0.6549 | |
| GSE53622 | OXSM | 54995 | 61284 | -0.6493 | 0.0000 | |
| GSE53624 | OXSM | 54995 | 61284 | -0.4021 | 0.0000 | |
| GSE63941 | OXSM | 54995 | 219133_at | 0.2598 | 0.5173 | |
| GSE77861 | OXSM | 54995 | 219133_at | -0.3998 | 0.0287 | |
| GSE97050 | OXSM | 54995 | A_33_P3246083 | -0.2935 | 0.3948 | |
| SRP007169 | OXSM | 54995 | RNAseq | -1.1039 | 0.0052 | |
| SRP008496 | OXSM | 54995 | RNAseq | -1.0294 | 0.0010 | |
| SRP064894 | OXSM | 54995 | RNAseq | -0.2908 | 0.0337 | |
| SRP133303 | OXSM | 54995 | RNAseq | -0.3997 | 0.0510 | |
| SRP159526 | OXSM | 54995 | RNAseq | -0.7879 | 0.0001 | |
| SRP193095 | OXSM | 54995 | RNAseq | -0.7048 | 0.0000 | |
| SRP219564 | OXSM | 54995 | RNAseq | -0.7252 | 0.0994 | |
| TCGA | OXSM | 54995 | RNAseq | -0.2005 | 0.0061 |
Upregulated datasets: 0; Downregulated datasets: 2.
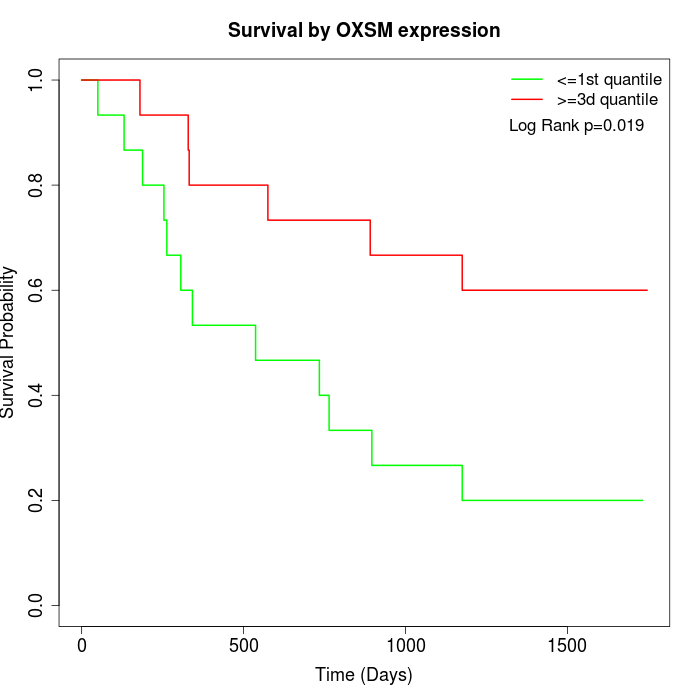
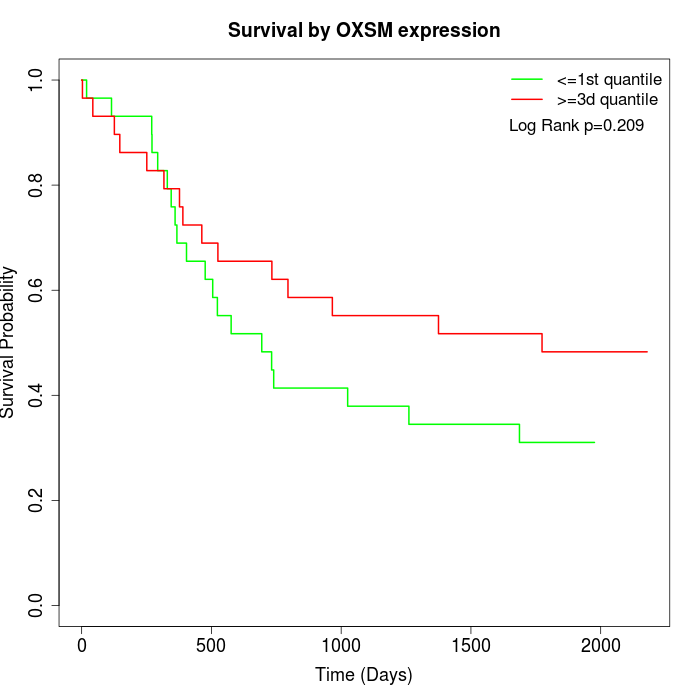
Survival by OXSM expression:
Note: Click image to view full size file.
Copy number change of OXSM:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | OXSM | 54995 | 0 | 18 | 12 | |
| GSE20123 | OXSM | 54995 | 0 | 18 | 12 | |
| GSE43470 | OXSM | 54995 | 0 | 20 | 23 | |
| GSE46452 | OXSM | 54995 | 2 | 17 | 40 | |
| GSE47630 | OXSM | 54995 | 1 | 24 | 15 | |
| GSE54993 | OXSM | 54995 | 6 | 2 | 62 | |
| GSE54994 | OXSM | 54995 | 1 | 32 | 20 | |
| GSE60625 | OXSM | 54995 | 5 | 0 | 6 | |
| GSE74703 | OXSM | 54995 | 0 | 17 | 19 | |
| GSE74704 | OXSM | 54995 | 0 | 12 | 8 | |
| TCGA | OXSM | 54995 | 1 | 68 | 27 |
Total number of gains: 16; Total number of losses: 228; Total Number of normals: 244.
Somatic mutations of OXSM:
Generating mutation plots.
Highly correlated genes for OXSM:
Showing top 20/1246 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| OXSM | FAM83D | 0.783617 | 3 | 0 | 3 |
| OXSM | CRK | 0.713751 | 3 | 0 | 3 |
| OXSM | MRPS11 | 0.707409 | 3 | 0 | 3 |
| OXSM | ZNF615 | 0.703679 | 3 | 0 | 3 |
| OXSM | IMPACT | 0.702399 | 4 | 0 | 4 |
| OXSM | RALA | 0.701518 | 7 | 0 | 6 |
| OXSM | MPP7 | 0.700993 | 5 | 0 | 5 |
| OXSM | SLC9A8 | 0.695362 | 3 | 0 | 3 |
| OXSM | TBC1D20 | 0.692591 | 3 | 0 | 3 |
| OXSM | ZNF823 | 0.689375 | 6 | 0 | 5 |
| OXSM | CYP4X1 | 0.687064 | 4 | 0 | 4 |
| OXSM | RNF144B | 0.682481 | 4 | 0 | 4 |
| OXSM | PCYOX1 | 0.678114 | 5 | 0 | 5 |
| OXSM | TAB3 | 0.677439 | 5 | 0 | 4 |
| OXSM | PITHD1 | 0.673674 | 5 | 0 | 4 |
| OXSM | GRPEL2 | 0.672546 | 5 | 0 | 5 |
| OXSM | PKIA | 0.671932 | 3 | 0 | 3 |
| OXSM | CCDC66 | 0.669852 | 3 | 0 | 3 |
| OXSM | FAM102A | 0.669475 | 4 | 0 | 3 |
| OXSM | BARD1 | 0.667588 | 5 | 0 | 4 |
For details and further investigation, click here