| Full name: RAS like proto-oncogene A | Alias Symbol: | ||
| Type: protein-coding gene | Cytoband: 7p14.1 | ||
| Entrez ID: 5898 | HGNC ID: HGNC:9839 | Ensembl Gene: ENSG00000006451 | OMIM ID: 179550 |
| Related drugs: VITAMIN E... [more] | |||
RALA involved pathways:
| KEGG pathway | Description | View |
|---|---|---|
| hsa04014 | Ras signaling pathway | |
| hsa04015 | Rap1 signaling pathway | |
| hsa04072 | Phospholipase D signaling pathway | |
| hsa05200 | Pathways in cancer | |
| hsa05212 | Pancreatic cancer |
Expression of RALA:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | RALA | 5898 | 224880_at | -0.7094 | 0.2189 | |
| GSE20347 | RALA | 5898 | 214435_x_at | -1.2734 | 0.0000 | |
| GSE23400 | RALA | 5898 | 214435_x_at | -0.8337 | 0.0000 | |
| GSE26886 | RALA | 5898 | 224880_at | -0.4668 | 0.0160 | |
| GSE29001 | RALA | 5898 | 214435_x_at | -1.1234 | 0.0047 | |
| GSE38129 | RALA | 5898 | 214435_x_at | -0.7035 | 0.0259 | |
| GSE45670 | RALA | 5898 | 224880_at | -0.4736 | 0.0068 | |
| GSE53622 | RALA | 5898 | 77323 | -0.6195 | 0.0000 | |
| GSE53624 | RALA | 5898 | 77323 | -0.7742 | 0.0000 | |
| GSE63941 | RALA | 5898 | 224880_at | -0.9382 | 0.0217 | |
| GSE77861 | RALA | 5898 | 224880_at | -0.7789 | 0.0052 | |
| GSE97050 | RALA | 5898 | A_24_P192262 | -0.4514 | 0.2988 | |
| SRP007169 | RALA | 5898 | RNAseq | -2.0008 | 0.0000 | |
| SRP008496 | RALA | 5898 | RNAseq | -1.5966 | 0.0000 | |
| SRP064894 | RALA | 5898 | RNAseq | -1.1393 | 0.0000 | |
| SRP133303 | RALA | 5898 | RNAseq | -0.5451 | 0.0052 | |
| SRP159526 | RALA | 5898 | RNAseq | -1.2906 | 0.0000 | |
| SRP193095 | RALA | 5898 | RNAseq | -1.3539 | 0.0000 | |
| SRP219564 | RALA | 5898 | RNAseq | -1.1320 | 0.0650 | |
| TCGA | RALA | 5898 | RNAseq | 0.0913 | 0.0935 |
Upregulated datasets: 0; Downregulated datasets: 7.
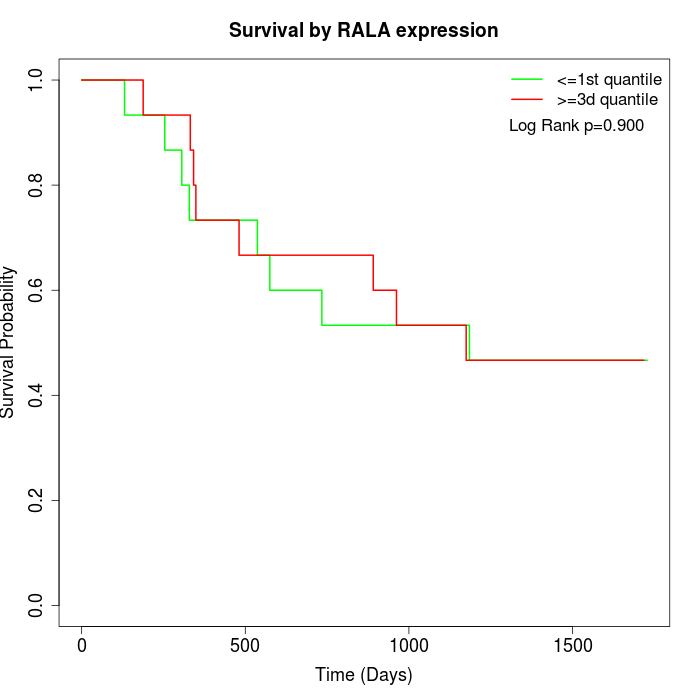
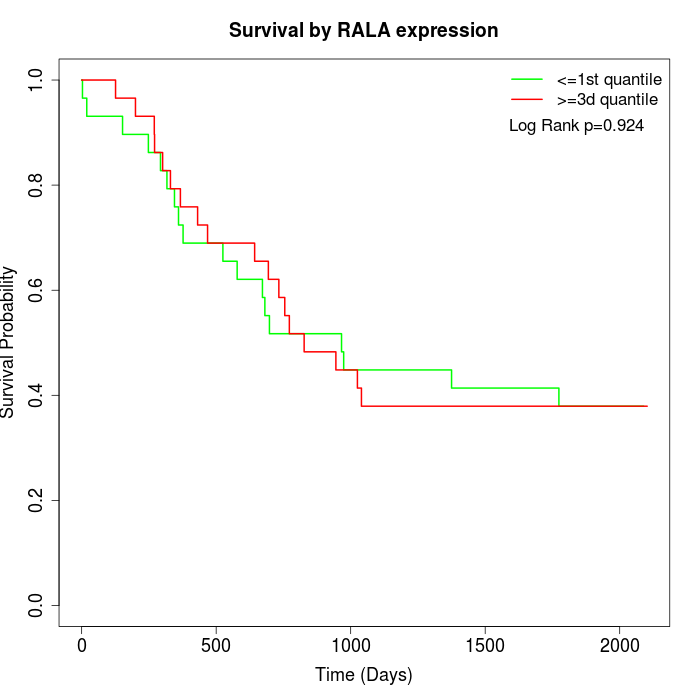
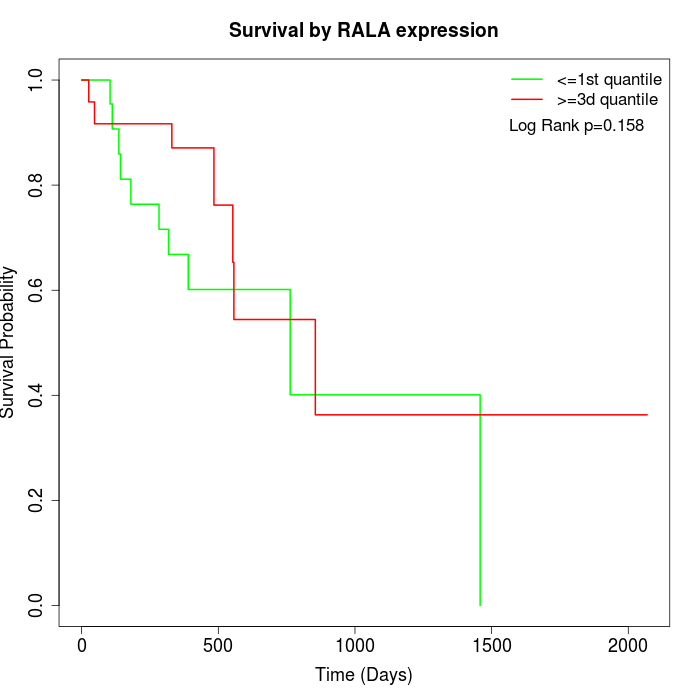
Survival by RALA expression:
Note: Click image to view full size file.
Copy number change of RALA:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | RALA | 5898 | 15 | 0 | 15 | |
| GSE20123 | RALA | 5898 | 14 | 0 | 16 | |
| GSE43470 | RALA | 5898 | 4 | 0 | 39 | |
| GSE46452 | RALA | 5898 | 11 | 1 | 47 | |
| GSE47630 | RALA | 5898 | 8 | 1 | 31 | |
| GSE54993 | RALA | 5898 | 0 | 7 | 63 | |
| GSE54994 | RALA | 5898 | 18 | 3 | 32 | |
| GSE60625 | RALA | 5898 | 0 | 0 | 11 | |
| GSE74703 | RALA | 5898 | 4 | 0 | 32 | |
| GSE74704 | RALA | 5898 | 8 | 0 | 12 | |
| TCGA | RALA | 5898 | 52 | 7 | 37 |
Total number of gains: 134; Total number of losses: 19; Total Number of normals: 335.
Somatic mutations of RALA:
Generating mutation plots.
Highly correlated genes for RALA:
Showing top 20/1369 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| RALA | PDCD6IP | 0.766399 | 11 | 0 | 10 |
| RALA | MNS1 | 0.762812 | 3 | 0 | 3 |
| RALA | ARIH1 | 0.74583 | 3 | 0 | 3 |
| RALA | C18orf25 | 0.744566 | 11 | 0 | 11 |
| RALA | CA13 | 0.741767 | 3 | 0 | 3 |
| RALA | ARF6 | 0.740002 | 11 | 0 | 11 |
| RALA | PIH1D2 | 0.738969 | 3 | 0 | 3 |
| RALA | SPTLC1 | 0.7375 | 11 | 0 | 11 |
| RALA | TJP1 | 0.733383 | 11 | 0 | 11 |
| RALA | VPS4B | 0.725172 | 11 | 0 | 11 |
| RALA | MCEE | 0.723008 | 5 | 0 | 4 |
| RALA | CCNYL1 | 0.722815 | 5 | 0 | 4 |
| RALA | ZNF823 | 0.722212 | 7 | 0 | 6 |
| RALA | RNF141 | 0.721842 | 12 | 0 | 11 |
| RALA | CES2 | 0.718659 | 8 | 0 | 8 |
| RALA | RANBP9 | 0.718593 | 11 | 0 | 11 |
| RALA | COMMD1 | 0.715111 | 3 | 0 | 3 |
| RALA | CCDC186 | 0.711241 | 10 | 0 | 10 |
| RALA | ECD | 0.70887 | 3 | 0 | 3 |
| RALA | SLK | 0.708767 | 12 | 0 | 11 |
For details and further investigation, click here